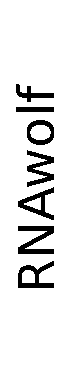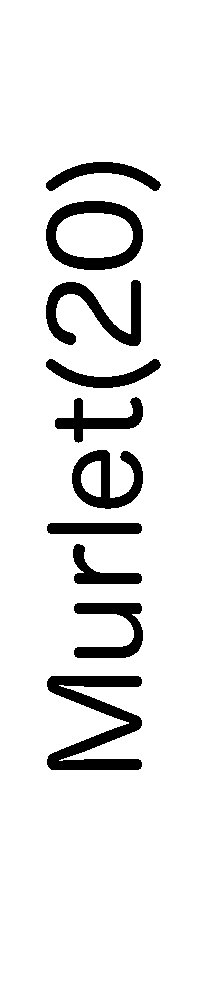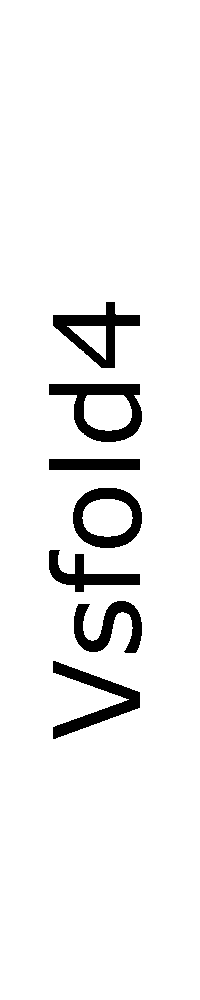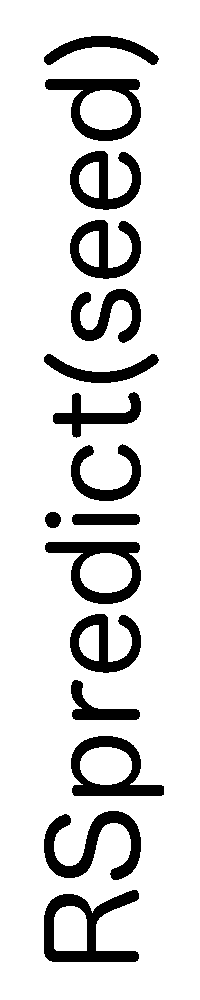| |
 |
 |
 |
 |
 |
 |
 |
 |
 |
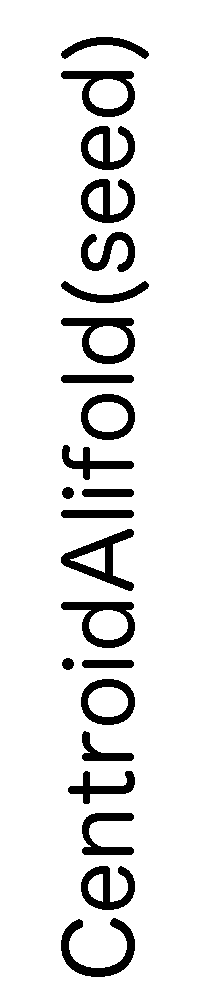 |
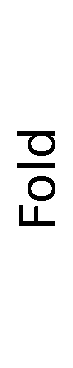 |
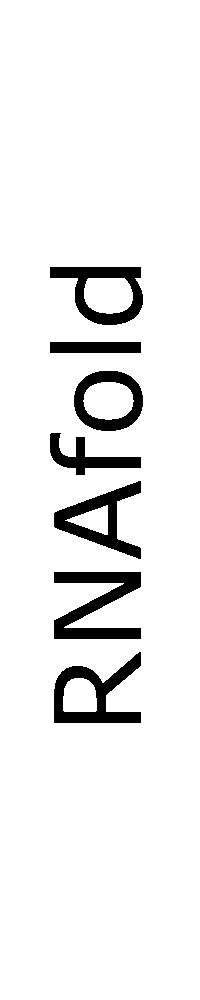 |
 |
 |
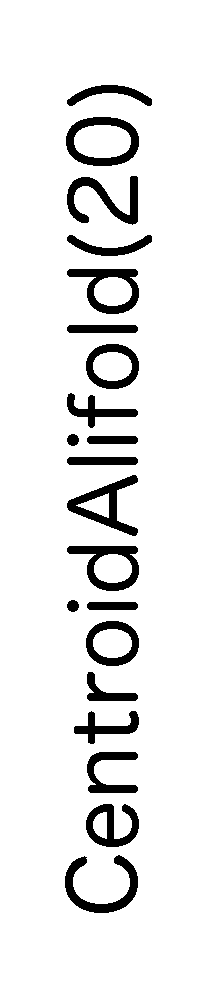 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
| ContextFold |
|
+
ContextFold vs IPknot
Matthews Correlation Coefficient
ContextFold:
0.680
IPknot:
0.548
Sensitivity
ContextFold:
0.644
IPknot:
0.475
Positive Predictive Value
ContextFold:
0.718
IPknot:
0.632
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PETfold_pre2.0(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
PETfold_pre2.0(seed):
0.529
Sensitivity
ContextFold:
0.644
PETfold_pre2.0(seed):
0.343
Positive Predictive Value
ContextFold:
0.718
PETfold_pre2.0(seed):
0.814
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Contrafold
Matthews Correlation Coefficient
ContextFold:
0.680
Contrafold:
0.508
Sensitivity
ContextFold:
0.644
Contrafold:
0.481
Positive Predictive Value
ContextFold:
0.718
Contrafold:
0.537
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidFold
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidFold:
0.470
Sensitivity
ContextFold:
0.644
CentroidFold:
0.369
Positive Predictive Value
ContextFold:
0.718
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Sfold
Matthews Correlation Coefficient
ContextFold:
0.680
Sfold:
0.469
Sensitivity
ContextFold:
0.644
Sfold:
0.412
Positive Predictive Value
ContextFold:
0.718
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MaxExpect
Matthews Correlation Coefficient
ContextFold:
0.680
MaxExpect:
0.465
Sensitivity
ContextFold:
0.644
MaxExpect:
0.443
Positive Predictive Value
ContextFold:
0.718
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs ProbKnot
Matthews Correlation Coefficient
ContextFold:
0.680
ProbKnot:
0.460
Sensitivity
ContextFold:
0.644
ProbKnot:
0.446
Positive Predictive Value
ContextFold:
0.718
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs UNAFold
Matthews Correlation Coefficient
ContextFold:
0.680
UNAFold:
0.445
Sensitivity
ContextFold:
0.644
UNAFold:
0.445
Positive Predictive Value
ContextFold:
0.718
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidAlifold(seed):
0.438
Sensitivity
ContextFold:
0.644
CentroidAlifold(seed):
0.213
Positive Predictive Value
ContextFold:
0.718
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Fold
Matthews Correlation Coefficient
ContextFold:
0.680
Fold:
0.437
Sensitivity
ContextFold:
0.644
Fold:
0.432
Positive Predictive Value
ContextFold:
0.718
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAfold
Matthews Correlation Coefficient
ContextFold:
0.680
RNAfold:
0.429
Sensitivity
ContextFold:
0.644
RNAfold:
0.422
Positive Predictive Value
ContextFold:
0.718
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidHomfold‑LAST:
0.428
Sensitivity
ContextFold:
0.644
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
ContextFold:
0.718
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
ContextFold:
0.680
PETfold_pre2.0(20):
0.413
Sensitivity
ContextFold:
0.644
PETfold_pre2.0(20):
0.211
Positive Predictive Value
ContextFold:
0.718
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidAlifold(20):
0.411
Sensitivity
ContextFold:
0.644
CentroidAlifold(20):
0.190
Positive Predictive Value
ContextFold:
0.718
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PknotsRG
Matthews Correlation Coefficient
ContextFold:
0.680
PknotsRG:
0.386
Sensitivity
ContextFold:
0.644
PknotsRG:
0.320
Positive Predictive Value
ContextFold:
0.718
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MXScarna(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
MXScarna(seed):
0.385
Sensitivity
ContextFold:
0.644
MXScarna(seed):
0.196
Positive Predictive Value
ContextFold:
0.718
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAalifold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
RNAalifold(20):
0.381
Sensitivity
ContextFold:
0.644
RNAalifold(20):
0.185
Positive Predictive Value
ContextFold:
0.718
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAalifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
RNAalifold(seed):
0.375
Sensitivity
ContextFold:
0.644
RNAalifold(seed):
0.169
Positive Predictive Value
ContextFold:
0.718
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Afold
Matthews Correlation Coefficient
ContextFold:
0.680
Afold:
0.358
Sensitivity
ContextFold:
0.644
Afold:
0.295
Positive Predictive Value
ContextFold:
0.718
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MXScarna(20)
Matthews Correlation Coefficient
ContextFold:
0.680
MXScarna(20):
0.359
Sensitivity
ContextFold:
0.644
MXScarna(20):
0.181
Positive Predictive Value
ContextFold:
0.718
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CRWrnafold
Matthews Correlation Coefficient
ContextFold:
0.680
CRWrnafold:
0.346
Sensitivity
ContextFold:
0.644
CRWrnafold:
0.275
Positive Predictive Value
ContextFold:
0.718
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAsubopt
Matthews Correlation Coefficient
ContextFold:
0.680
RNAsubopt:
0.336
Sensitivity
ContextFold:
0.644
RNAsubopt:
0.217
Positive Predictive Value
ContextFold:
0.718
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs McQFold
Matthews Correlation Coefficient
ContextFold:
0.680
McQFold:
0.314
Sensitivity
ContextFold:
0.644
McQFold:
0.205
Positive Predictive Value
ContextFold:
0.718
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs TurboFold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
TurboFold(20):
0.301
Sensitivity
ContextFold:
0.644
TurboFold(20):
0.108
Positive Predictive Value
ContextFold:
0.718
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PPfold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
PPfold(20):
0.298
Sensitivity
ContextFold:
0.644
PPfold(20):
0.101
Positive Predictive Value
ContextFold:
0.718
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAshapes
Matthews Correlation Coefficient
ContextFold:
0.680
RNAshapes:
0.295
Sensitivity
ContextFold:
0.644
RNAshapes:
0.180
Positive Predictive Value
ContextFold:
0.718
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Carnac(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Carnac(20):
0.288
Sensitivity
ContextFold:
0.644
Carnac(20):
0.093
Positive Predictive Value
ContextFold:
0.718
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNASLOpt
Matthews Correlation Coefficient
ContextFold:
0.680
RNASLOpt:
0.281
Sensitivity
ContextFold:
0.644
RNASLOpt:
0.158
Positive Predictive Value
ContextFold:
0.718
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RSpredict(20)
Matthews Correlation Coefficient
ContextFold:
0.680
RSpredict(20):
0.270
Sensitivity
ContextFold:
0.644
RSpredict(20):
0.124
Positive Predictive Value
ContextFold:
0.718
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Multilign(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Multilign(20):
0.251
Sensitivity
ContextFold:
0.644
Multilign(20):
0.083
Positive Predictive Value
ContextFold:
0.718
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAwolf
Matthews Correlation Coefficient
ContextFold:
0.680
RNAwolf:
0.251
Sensitivity
ContextFold:
0.644
RNAwolf:
0.191
Positive Predictive Value
ContextFold:
0.718
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Murlet(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Murlet(20):
0.230
Sensitivity
ContextFold:
0.644
Murlet(20):
0.067
Positive Predictive Value
ContextFold:
0.718
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Vsfold4
Matthews Correlation Coefficient
ContextFold:
0.680
Vsfold4:
0.217
Sensitivity
ContextFold:
0.644
Vsfold4:
0.125
Positive Predictive Value
ContextFold:
0.718
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Vsfold5
Matthews Correlation Coefficient
ContextFold:
0.680
Vsfold5:
0.211
Sensitivity
ContextFold:
0.644
Vsfold5:
0.125
Positive Predictive Value
ContextFold:
0.718
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RSpredict(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
RSpredict(seed):
0.212
Sensitivity
ContextFold:
0.644
RSpredict(seed):
0.075
Positive Predictive Value
ContextFold:
0.718
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CMfinder(20)
Matthews Correlation Coefficient
ContextFold:
0.680
CMfinder(20):
0.210
Sensitivity
ContextFold:
0.644
CMfinder(20):
0.055
Positive Predictive Value
ContextFold:
0.718
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs HotKnots
Matthews Correlation Coefficient
ContextFold:
0.680
HotKnots:
0.179
Sensitivity
ContextFold:
0.644
HotKnots:
0.051
Positive Predictive Value
ContextFold:
0.718
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNASampler(20)
Matthews Correlation Coefficient
ContextFold:
0.680
RNASampler(20):
0.176
Sensitivity
ContextFold:
0.644
RNASampler(20):
0.035
Positive Predictive Value
ContextFold:
0.718
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Cylofold
Matthews Correlation Coefficient
ContextFold:
0.680
Cylofold:
0.155
Sensitivity
ContextFold:
0.644
Cylofold:
0.040
Positive Predictive Value
ContextFold:
0.718
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Pknots
Matthews Correlation Coefficient
ContextFold:
0.680
Pknots:
0.143
Sensitivity
ContextFold:
0.644
Pknots:
0.033
Positive Predictive Value
ContextFold:
0.718
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Mastr(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Mastr(20):
0.134
Sensitivity
ContextFold:
0.644
Mastr(20):
0.023
Positive Predictive Value
ContextFold:
0.718
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Alterna
Matthews Correlation Coefficient
ContextFold:
0.680
Alterna:
0.129
Sensitivity
ContextFold:
0.644
Alterna:
0.030
Positive Predictive Value
ContextFold:
0.718
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MCFold
Matthews Correlation Coefficient
ContextFold:
0.680
MCFold:
0.125
Sensitivity
ContextFold:
0.644
MCFold:
0.036
Positive Predictive Value
ContextFold:
0.718
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RDfolder
Matthews Correlation Coefficient
ContextFold:
0.680
RDfolder:
0.122
Sensitivity
ContextFold:
0.644
RDfolder:
0.025
Positive Predictive Value
ContextFold:
0.718
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Murlet(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Murlet(seed):
0.079
Sensitivity
ContextFold:
0.644
Murlet(seed):
0.008
Positive Predictive Value
ContextFold:
0.718
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CMfinder(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
CMfinder(seed):
0.080
Sensitivity
ContextFold:
0.644
CMfinder(seed):
0.008
Positive Predictive Value
ContextFold:
0.718
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs TurboFold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
TurboFold(seed):
0.076
Sensitivity
ContextFold:
0.644
TurboFold(seed):
0.007
Positive Predictive Value
ContextFold:
0.718
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PPfold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
PPfold(seed):
0.069
Sensitivity
ContextFold:
0.644
PPfold(seed):
0.006
Positive Predictive Value
ContextFold:
0.718
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Carnac(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Carnac(seed):
0.066
Sensitivity
ContextFold:
0.644
Carnac(seed):
0.005
Positive Predictive Value
ContextFold:
0.718
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNASampler(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
RNASampler(seed):
0.063
Sensitivity
ContextFold:
0.644
RNASampler(seed):
0.005
Positive Predictive Value
ContextFold:
0.718
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs NanoFolder
Matthews Correlation Coefficient
ContextFold:
0.680
NanoFolder:
0.052
Sensitivity
ContextFold:
0.644
NanoFolder:
0.016
Positive Predictive Value
ContextFold:
0.718
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Mastr(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Mastr(seed):
0.051
Sensitivity
ContextFold:
0.644
Mastr(seed):
0.003
Positive Predictive Value
ContextFold:
0.718
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Multilign(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Multilign(seed):
0.043
Sensitivity
ContextFold:
0.644
Multilign(seed):
0.003
Positive Predictive Value
ContextFold:
0.718
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| IPknot |
1987
ContextFold vs IPknot
Matthews Correlation Coefficient
ContextFold:
0.680
IPknot:
0.548
Sensitivity
ContextFold:
0.644
IPknot:
0.475
Positive Predictive Value
ContextFold:
0.718
IPknot:
0.632
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
IPknot vs PETfold_pre2.0(seed)
Matthews Correlation Coefficient
IPknot:
0.548
PETfold_pre2.0(seed):
0.529
Sensitivity
IPknot:
0.475
PETfold_pre2.0(seed):
0.343
Positive Predictive Value
IPknot:
0.632
PETfold_pre2.0(seed):
0.814
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Contrafold
Matthews Correlation Coefficient
IPknot:
0.548
Contrafold:
0.508
Sensitivity
IPknot:
0.475
Contrafold:
0.481
Positive Predictive Value
IPknot:
0.632
Contrafold:
0.537
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidFold
Matthews Correlation Coefficient
IPknot:
0.548
CentroidFold:
0.470
Sensitivity
IPknot:
0.475
CentroidFold:
0.369
Positive Predictive Value
IPknot:
0.632
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Sfold
Matthews Correlation Coefficient
IPknot:
0.548
Sfold:
0.469
Sensitivity
IPknot:
0.475
Sfold:
0.412
Positive Predictive Value
IPknot:
0.632
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MaxExpect
Matthews Correlation Coefficient
IPknot:
0.548
MaxExpect:
0.465
Sensitivity
IPknot:
0.475
MaxExpect:
0.443
Positive Predictive Value
IPknot:
0.632
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs ProbKnot
Matthews Correlation Coefficient
IPknot:
0.548
ProbKnot:
0.460
Sensitivity
IPknot:
0.475
ProbKnot:
0.446
Positive Predictive Value
IPknot:
0.632
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs UNAFold
Matthews Correlation Coefficient
IPknot:
0.548
UNAFold:
0.445
Sensitivity
IPknot:
0.475
UNAFold:
0.445
Positive Predictive Value
IPknot:
0.632
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidAlifold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
CentroidAlifold(seed):
0.438
Sensitivity
IPknot:
0.475
CentroidAlifold(seed):
0.213
Positive Predictive Value
IPknot:
0.632
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Fold
Matthews Correlation Coefficient
IPknot:
0.548
Fold:
0.437
Sensitivity
IPknot:
0.475
Fold:
0.432
Positive Predictive Value
IPknot:
0.632
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAfold
Matthews Correlation Coefficient
IPknot:
0.548
RNAfold:
0.429
Sensitivity
IPknot:
0.475
RNAfold:
0.422
Positive Predictive Value
IPknot:
0.632
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
IPknot:
0.548
CentroidHomfold‑LAST:
0.428
Sensitivity
IPknot:
0.475
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
IPknot:
0.632
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
IPknot:
0.548
PETfold_pre2.0(20):
0.413
Sensitivity
IPknot:
0.475
PETfold_pre2.0(20):
0.211
Positive Predictive Value
IPknot:
0.632
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidAlifold(20)
Matthews Correlation Coefficient
IPknot:
0.548
CentroidAlifold(20):
0.411
Sensitivity
IPknot:
0.475
CentroidAlifold(20):
0.190
Positive Predictive Value
IPknot:
0.632
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PknotsRG
Matthews Correlation Coefficient
IPknot:
0.548
PknotsRG:
0.386
Sensitivity
IPknot:
0.475
PknotsRG:
0.320
Positive Predictive Value
IPknot:
0.632
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MXScarna(seed)
Matthews Correlation Coefficient
IPknot:
0.548
MXScarna(seed):
0.385
Sensitivity
IPknot:
0.475
MXScarna(seed):
0.196
Positive Predictive Value
IPknot:
0.632
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAalifold(20)
Matthews Correlation Coefficient
IPknot:
0.548
RNAalifold(20):
0.381
Sensitivity
IPknot:
0.475
RNAalifold(20):
0.185
Positive Predictive Value
IPknot:
0.632
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAalifold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
RNAalifold(seed):
0.375
Sensitivity
IPknot:
0.475
RNAalifold(seed):
0.169
Positive Predictive Value
IPknot:
0.632
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Afold
Matthews Correlation Coefficient
IPknot:
0.548
Afold:
0.358
Sensitivity
IPknot:
0.475
Afold:
0.295
Positive Predictive Value
IPknot:
0.632
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MXScarna(20)
Matthews Correlation Coefficient
IPknot:
0.548
MXScarna(20):
0.359
Sensitivity
IPknot:
0.475
MXScarna(20):
0.181
Positive Predictive Value
IPknot:
0.632
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CRWrnafold
Matthews Correlation Coefficient
IPknot:
0.548
CRWrnafold:
0.346
Sensitivity
IPknot:
0.475
CRWrnafold:
0.275
Positive Predictive Value
IPknot:
0.632
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAsubopt
Matthews Correlation Coefficient
IPknot:
0.548
RNAsubopt:
0.336
Sensitivity
IPknot:
0.475
RNAsubopt:
0.217
Positive Predictive Value
IPknot:
0.632
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs McQFold
Matthews Correlation Coefficient
IPknot:
0.548
McQFold:
0.314
Sensitivity
IPknot:
0.475
McQFold:
0.205
Positive Predictive Value
IPknot:
0.632
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs TurboFold(20)
Matthews Correlation Coefficient
IPknot:
0.548
TurboFold(20):
0.301
Sensitivity
IPknot:
0.475
TurboFold(20):
0.108
Positive Predictive Value
IPknot:
0.632
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PPfold(20)
Matthews Correlation Coefficient
IPknot:
0.548
PPfold(20):
0.298
Sensitivity
IPknot:
0.475
PPfold(20):
0.101
Positive Predictive Value
IPknot:
0.632
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAshapes
Matthews Correlation Coefficient
IPknot:
0.548
RNAshapes:
0.295
Sensitivity
IPknot:
0.475
RNAshapes:
0.180
Positive Predictive Value
IPknot:
0.632
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Carnac(20)
Matthews Correlation Coefficient
IPknot:
0.548
Carnac(20):
0.288
Sensitivity
IPknot:
0.475
Carnac(20):
0.093
Positive Predictive Value
IPknot:
0.632
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNASLOpt
Matthews Correlation Coefficient
IPknot:
0.548
RNASLOpt:
0.281
Sensitivity
IPknot:
0.475
RNASLOpt:
0.158
Positive Predictive Value
IPknot:
0.632
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RSpredict(20)
Matthews Correlation Coefficient
IPknot:
0.548
RSpredict(20):
0.270
Sensitivity
IPknot:
0.475
RSpredict(20):
0.124
Positive Predictive Value
IPknot:
0.632
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Multilign(20)
Matthews Correlation Coefficient
IPknot:
0.548
Multilign(20):
0.251
Sensitivity
IPknot:
0.475
Multilign(20):
0.083
Positive Predictive Value
IPknot:
0.632
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAwolf
Matthews Correlation Coefficient
IPknot:
0.548
RNAwolf:
0.251
Sensitivity
IPknot:
0.475
RNAwolf:
0.191
Positive Predictive Value
IPknot:
0.632
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Murlet(20)
Matthews Correlation Coefficient
IPknot:
0.548
Murlet(20):
0.230
Sensitivity
IPknot:
0.475
Murlet(20):
0.067
Positive Predictive Value
IPknot:
0.632
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Vsfold4
Matthews Correlation Coefficient
IPknot:
0.548
Vsfold4:
0.217
Sensitivity
IPknot:
0.475
Vsfold4:
0.125
Positive Predictive Value
IPknot:
0.632
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Vsfold5
Matthews Correlation Coefficient
IPknot:
0.548
Vsfold5:
0.211
Sensitivity
IPknot:
0.475
Vsfold5:
0.125
Positive Predictive Value
IPknot:
0.632
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RSpredict(seed)
Matthews Correlation Coefficient
IPknot:
0.548
RSpredict(seed):
0.212
Sensitivity
IPknot:
0.475
RSpredict(seed):
0.075
Positive Predictive Value
IPknot:
0.632
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CMfinder(20)
Matthews Correlation Coefficient
IPknot:
0.548
CMfinder(20):
0.210
Sensitivity
IPknot:
0.475
CMfinder(20):
0.055
Positive Predictive Value
IPknot:
0.632
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs HotKnots
Matthews Correlation Coefficient
IPknot:
0.548
HotKnots:
0.179
Sensitivity
IPknot:
0.475
HotKnots:
0.051
Positive Predictive Value
IPknot:
0.632
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNASampler(20)
Matthews Correlation Coefficient
IPknot:
0.548
RNASampler(20):
0.176
Sensitivity
IPknot:
0.475
RNASampler(20):
0.035
Positive Predictive Value
IPknot:
0.632
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Cylofold
Matthews Correlation Coefficient
IPknot:
0.548
Cylofold:
0.155
Sensitivity
IPknot:
0.475
Cylofold:
0.040
Positive Predictive Value
IPknot:
0.632
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Pknots
Matthews Correlation Coefficient
IPknot:
0.548
Pknots:
0.143
Sensitivity
IPknot:
0.475
Pknots:
0.033
Positive Predictive Value
IPknot:
0.632
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Mastr(20)
Matthews Correlation Coefficient
IPknot:
0.548
Mastr(20):
0.134
Sensitivity
IPknot:
0.475
Mastr(20):
0.023
Positive Predictive Value
IPknot:
0.632
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Alterna
Matthews Correlation Coefficient
IPknot:
0.548
Alterna:
0.129
Sensitivity
IPknot:
0.475
Alterna:
0.030
Positive Predictive Value
IPknot:
0.632
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MCFold
Matthews Correlation Coefficient
IPknot:
0.548
MCFold:
0.125
Sensitivity
IPknot:
0.475
MCFold:
0.036
Positive Predictive Value
IPknot:
0.632
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RDfolder
Matthews Correlation Coefficient
IPknot:
0.548
RDfolder:
0.122
Sensitivity
IPknot:
0.475
RDfolder:
0.025
Positive Predictive Value
IPknot:
0.632
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Murlet(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Murlet(seed):
0.079
Sensitivity
IPknot:
0.475
Murlet(seed):
0.008
Positive Predictive Value
IPknot:
0.632
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CMfinder(seed)
Matthews Correlation Coefficient
IPknot:
0.548
CMfinder(seed):
0.080
Sensitivity
IPknot:
0.475
CMfinder(seed):
0.008
Positive Predictive Value
IPknot:
0.632
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs TurboFold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
TurboFold(seed):
0.076
Sensitivity
IPknot:
0.475
TurboFold(seed):
0.007
Positive Predictive Value
IPknot:
0.632
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PPfold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
PPfold(seed):
0.069
Sensitivity
IPknot:
0.475
PPfold(seed):
0.006
Positive Predictive Value
IPknot:
0.632
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Carnac(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Carnac(seed):
0.066
Sensitivity
IPknot:
0.475
Carnac(seed):
0.005
Positive Predictive Value
IPknot:
0.632
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNASampler(seed)
Matthews Correlation Coefficient
IPknot:
0.548
RNASampler(seed):
0.063
Sensitivity
IPknot:
0.475
RNASampler(seed):
0.005
Positive Predictive Value
IPknot:
0.632
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs NanoFolder
Matthews Correlation Coefficient
IPknot:
0.548
NanoFolder:
0.052
Sensitivity
IPknot:
0.475
NanoFolder:
0.016
Positive Predictive Value
IPknot:
0.632
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Mastr(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Mastr(seed):
0.051
Sensitivity
IPknot:
0.475
Mastr(seed):
0.003
Positive Predictive Value
IPknot:
0.632
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Multilign(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Multilign(seed):
0.043
Sensitivity
IPknot:
0.475
Multilign(seed):
0.003
Positive Predictive Value
IPknot:
0.632
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PETfold_pre2.0(seed) |
1987
ContextFold vs PETfold_pre2.0(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
PETfold_pre2.0(seed):
0.529
Sensitivity
ContextFold:
0.644
PETfold_pre2.0(seed):
0.343
Positive Predictive Value
ContextFold:
0.718
PETfold_pre2.0(seed):
0.814
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs PETfold_pre2.0(seed)
Matthews Correlation Coefficient
IPknot:
0.548
PETfold_pre2.0(seed):
0.529
Sensitivity
IPknot:
0.475
PETfold_pre2.0(seed):
0.343
Positive Predictive Value
IPknot:
0.632
PETfold_pre2.0(seed):
0.814
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PETfold_pre2.0(seed) vs Contrafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Contrafold:
0.508
Sensitivity
PETfold_pre2.0(seed):
0.343
Contrafold:
0.481
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Contrafold:
0.537
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidFold:
0.470
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidFold:
0.369
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Sfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Sfold:
0.469
Sensitivity
PETfold_pre2.0(seed):
0.343
Sfold:
0.412
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MaxExpect
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MaxExpect:
0.465
Sensitivity
PETfold_pre2.0(seed):
0.343
MaxExpect:
0.443
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs ProbKnot
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
ProbKnot:
0.460
Sensitivity
PETfold_pre2.0(seed):
0.343
ProbKnot:
0.446
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs UNAFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
UNAFold:
0.445
Sensitivity
PETfold_pre2.0(seed):
0.343
UNAFold:
0.445
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidAlifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidAlifold(seed):
0.438
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidAlifold(seed):
0.213
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Fold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Fold:
0.437
Sensitivity
PETfold_pre2.0(seed):
0.343
Fold:
0.432
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAfold:
0.429
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAfold:
0.422
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidHomfold‑LAST:
0.428
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PETfold_pre2.0(20):
0.413
Sensitivity
PETfold_pre2.0(seed):
0.343
PETfold_pre2.0(20):
0.211
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidAlifold(20):
0.411
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidAlifold(20):
0.190
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PknotsRG:
0.386
Sensitivity
PETfold_pre2.0(seed):
0.343
PknotsRG:
0.320
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MXScarna(seed):
0.385
Sensitivity
PETfold_pre2.0(seed):
0.343
MXScarna(seed):
0.196
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAalifold(20):
0.381
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAalifold(20):
0.185
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAalifold(seed):
0.375
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAalifold(seed):
0.169
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Afold:
0.358
Sensitivity
PETfold_pre2.0(seed):
0.343
Afold:
0.295
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MXScarna(20):
0.359
Sensitivity
PETfold_pre2.0(seed):
0.343
MXScarna(20):
0.181
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CRWrnafold:
0.346
Sensitivity
PETfold_pre2.0(seed):
0.343
CRWrnafold:
0.275
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAsubopt:
0.336
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAsubopt:
0.217
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
McQFold:
0.314
Sensitivity
PETfold_pre2.0(seed):
0.343
McQFold:
0.205
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
TurboFold(20):
0.301
Sensitivity
PETfold_pre2.0(seed):
0.343
TurboFold(20):
0.108
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PPfold(20):
0.298
Sensitivity
PETfold_pre2.0(seed):
0.343
PPfold(20):
0.101
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAshapes:
0.295
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAshapes:
0.180
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Carnac(20):
0.288
Sensitivity
PETfold_pre2.0(seed):
0.343
Carnac(20):
0.093
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNASLOpt:
0.281
Sensitivity
PETfold_pre2.0(seed):
0.343
RNASLOpt:
0.158
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RSpredict(20):
0.270
Sensitivity
PETfold_pre2.0(seed):
0.343
RSpredict(20):
0.124
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Multilign(20):
0.251
Sensitivity
PETfold_pre2.0(seed):
0.343
Multilign(20):
0.083
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAwolf:
0.251
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAwolf:
0.191
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Murlet(20):
0.230
Sensitivity
PETfold_pre2.0(seed):
0.343
Murlet(20):
0.067
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Vsfold4:
0.217
Sensitivity
PETfold_pre2.0(seed):
0.343
Vsfold4:
0.125
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Vsfold5:
0.211
Sensitivity
PETfold_pre2.0(seed):
0.343
Vsfold5:
0.125
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RSpredict(seed):
0.212
Sensitivity
PETfold_pre2.0(seed):
0.343
RSpredict(seed):
0.075
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CMfinder(20):
0.210
Sensitivity
PETfold_pre2.0(seed):
0.343
CMfinder(20):
0.055
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
HotKnots:
0.179
Sensitivity
PETfold_pre2.0(seed):
0.343
HotKnots:
0.051
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNASampler(20):
0.176
Sensitivity
PETfold_pre2.0(seed):
0.343
RNASampler(20):
0.035
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Cylofold:
0.155
Sensitivity
PETfold_pre2.0(seed):
0.343
Cylofold:
0.040
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Pknots:
0.143
Sensitivity
PETfold_pre2.0(seed):
0.343
Pknots:
0.033
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Mastr(20):
0.134
Sensitivity
PETfold_pre2.0(seed):
0.343
Mastr(20):
0.023
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Alterna:
0.129
Sensitivity
PETfold_pre2.0(seed):
0.343
Alterna:
0.030
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MCFold:
0.125
Sensitivity
PETfold_pre2.0(seed):
0.343
MCFold:
0.036
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RDfolder:
0.122
Sensitivity
PETfold_pre2.0(seed):
0.343
RDfolder:
0.025
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Murlet(seed):
0.079
Sensitivity
PETfold_pre2.0(seed):
0.343
Murlet(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CMfinder(seed):
0.080
Sensitivity
PETfold_pre2.0(seed):
0.343
CMfinder(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
TurboFold(seed):
0.076
Sensitivity
PETfold_pre2.0(seed):
0.343
TurboFold(seed):
0.007
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PPfold(seed):
0.069
Sensitivity
PETfold_pre2.0(seed):
0.343
PPfold(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Carnac(seed):
0.066
Sensitivity
PETfold_pre2.0(seed):
0.343
Carnac(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNASampler(seed):
0.063
Sensitivity
PETfold_pre2.0(seed):
0.343
RNASampler(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
NanoFolder:
0.052
Sensitivity
PETfold_pre2.0(seed):
0.343
NanoFolder:
0.016
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Mastr(seed):
0.051
Sensitivity
PETfold_pre2.0(seed):
0.343
Mastr(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Multilign(seed):
0.043
Sensitivity
PETfold_pre2.0(seed):
0.343
Multilign(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Contrafold |
1987
ContextFold vs Contrafold
Matthews Correlation Coefficient
ContextFold:
0.680
Contrafold:
0.508
Sensitivity
ContextFold:
0.644
Contrafold:
0.481
Positive Predictive Value
ContextFold:
0.718
Contrafold:
0.537
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Contrafold
Matthews Correlation Coefficient
IPknot:
0.548
Contrafold:
0.508
Sensitivity
IPknot:
0.475
Contrafold:
0.481
Positive Predictive Value
IPknot:
0.632
Contrafold:
0.537
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Contrafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Contrafold:
0.508
Sensitivity
PETfold_pre2.0(seed):
0.343
Contrafold:
0.481
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Contrafold:
0.537
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Contrafold vs CentroidFold
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidFold:
0.470
Sensitivity
Contrafold:
0.481
CentroidFold:
0.369
Positive Predictive Value
Contrafold:
0.537
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Sfold
Matthews Correlation Coefficient
Contrafold:
0.508
Sfold:
0.469
Sensitivity
Contrafold:
0.481
Sfold:
0.412
Positive Predictive Value
Contrafold:
0.537
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MaxExpect
Matthews Correlation Coefficient
Contrafold:
0.508
MaxExpect:
0.465
Sensitivity
Contrafold:
0.481
MaxExpect:
0.443
Positive Predictive Value
Contrafold:
0.537
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs ProbKnot
Matthews Correlation Coefficient
Contrafold:
0.508
ProbKnot:
0.460
Sensitivity
Contrafold:
0.481
ProbKnot:
0.446
Positive Predictive Value
Contrafold:
0.537
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs UNAFold
Matthews Correlation Coefficient
Contrafold:
0.508
UNAFold:
0.445
Sensitivity
Contrafold:
0.481
UNAFold:
0.445
Positive Predictive Value
Contrafold:
0.537
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidAlifold(seed):
0.438
Sensitivity
Contrafold:
0.481
CentroidAlifold(seed):
0.213
Positive Predictive Value
Contrafold:
0.537
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Fold
Matthews Correlation Coefficient
Contrafold:
0.508
Fold:
0.437
Sensitivity
Contrafold:
0.481
Fold:
0.432
Positive Predictive Value
Contrafold:
0.537
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAfold
Matthews Correlation Coefficient
Contrafold:
0.508
RNAfold:
0.429
Sensitivity
Contrafold:
0.481
RNAfold:
0.422
Positive Predictive Value
Contrafold:
0.537
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidHomfold‑LAST:
0.428
Sensitivity
Contrafold:
0.481
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
Contrafold:
0.537
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
Contrafold:
0.508
PETfold_pre2.0(20):
0.413
Sensitivity
Contrafold:
0.481
PETfold_pre2.0(20):
0.211
Positive Predictive Value
Contrafold:
0.537
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CentroidAlifold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidAlifold(20):
0.411
Sensitivity
Contrafold:
0.481
CentroidAlifold(20):
0.190
Positive Predictive Value
Contrafold:
0.537
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PknotsRG
Matthews Correlation Coefficient
Contrafold:
0.508
PknotsRG:
0.386
Sensitivity
Contrafold:
0.481
PknotsRG:
0.320
Positive Predictive Value
Contrafold:
0.537
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MXScarna(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
MXScarna(seed):
0.385
Sensitivity
Contrafold:
0.481
MXScarna(seed):
0.196
Positive Predictive Value
Contrafold:
0.537
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAalifold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
RNAalifold(20):
0.381
Sensitivity
Contrafold:
0.481
RNAalifold(20):
0.185
Positive Predictive Value
Contrafold:
0.537
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAalifold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
RNAalifold(seed):
0.375
Sensitivity
Contrafold:
0.481
RNAalifold(seed):
0.169
Positive Predictive Value
Contrafold:
0.537
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Afold
Matthews Correlation Coefficient
Contrafold:
0.508
Afold:
0.358
Sensitivity
Contrafold:
0.481
Afold:
0.295
Positive Predictive Value
Contrafold:
0.537
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MXScarna(20)
Matthews Correlation Coefficient
Contrafold:
0.508
MXScarna(20):
0.359
Sensitivity
Contrafold:
0.481
MXScarna(20):
0.181
Positive Predictive Value
Contrafold:
0.537
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CRWrnafold
Matthews Correlation Coefficient
Contrafold:
0.508
CRWrnafold:
0.346
Sensitivity
Contrafold:
0.481
CRWrnafold:
0.275
Positive Predictive Value
Contrafold:
0.537
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAsubopt
Matthews Correlation Coefficient
Contrafold:
0.508
RNAsubopt:
0.336
Sensitivity
Contrafold:
0.481
RNAsubopt:
0.217
Positive Predictive Value
Contrafold:
0.537
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs McQFold
Matthews Correlation Coefficient
Contrafold:
0.508
McQFold:
0.314
Sensitivity
Contrafold:
0.481
McQFold:
0.205
Positive Predictive Value
Contrafold:
0.537
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs TurboFold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
TurboFold(20):
0.301
Sensitivity
Contrafold:
0.481
TurboFold(20):
0.108
Positive Predictive Value
Contrafold:
0.537
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PPfold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
PPfold(20):
0.298
Sensitivity
Contrafold:
0.481
PPfold(20):
0.101
Positive Predictive Value
Contrafold:
0.537
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAshapes
Matthews Correlation Coefficient
Contrafold:
0.508
RNAshapes:
0.295
Sensitivity
Contrafold:
0.481
RNAshapes:
0.180
Positive Predictive Value
Contrafold:
0.537
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Carnac(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Carnac(20):
0.288
Sensitivity
Contrafold:
0.481
Carnac(20):
0.093
Positive Predictive Value
Contrafold:
0.537
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNASLOpt
Matthews Correlation Coefficient
Contrafold:
0.508
RNASLOpt:
0.281
Sensitivity
Contrafold:
0.481
RNASLOpt:
0.158
Positive Predictive Value
Contrafold:
0.537
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RSpredict(20)
Matthews Correlation Coefficient
Contrafold:
0.508
RSpredict(20):
0.270
Sensitivity
Contrafold:
0.481
RSpredict(20):
0.124
Positive Predictive Value
Contrafold:
0.537
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Multilign(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Multilign(20):
0.251
Sensitivity
Contrafold:
0.481
Multilign(20):
0.083
Positive Predictive Value
Contrafold:
0.537
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAwolf
Matthews Correlation Coefficient
Contrafold:
0.508
RNAwolf:
0.251
Sensitivity
Contrafold:
0.481
RNAwolf:
0.191
Positive Predictive Value
Contrafold:
0.537
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Murlet(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Murlet(20):
0.230
Sensitivity
Contrafold:
0.481
Murlet(20):
0.067
Positive Predictive Value
Contrafold:
0.537
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Vsfold4
Matthews Correlation Coefficient
Contrafold:
0.508
Vsfold4:
0.217
Sensitivity
Contrafold:
0.481
Vsfold4:
0.125
Positive Predictive Value
Contrafold:
0.537
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Vsfold5
Matthews Correlation Coefficient
Contrafold:
0.508
Vsfold5:
0.211
Sensitivity
Contrafold:
0.481
Vsfold5:
0.125
Positive Predictive Value
Contrafold:
0.537
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RSpredict(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
RSpredict(seed):
0.212
Sensitivity
Contrafold:
0.481
RSpredict(seed):
0.075
Positive Predictive Value
Contrafold:
0.537
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CMfinder(20)
Matthews Correlation Coefficient
Contrafold:
0.508
CMfinder(20):
0.210
Sensitivity
Contrafold:
0.481
CMfinder(20):
0.055
Positive Predictive Value
Contrafold:
0.537
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs HotKnots
Matthews Correlation Coefficient
Contrafold:
0.508
HotKnots:
0.179
Sensitivity
Contrafold:
0.481
HotKnots:
0.051
Positive Predictive Value
Contrafold:
0.537
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNASampler(20)
Matthews Correlation Coefficient
Contrafold:
0.508
RNASampler(20):
0.176
Sensitivity
Contrafold:
0.481
RNASampler(20):
0.035
Positive Predictive Value
Contrafold:
0.537
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Cylofold
Matthews Correlation Coefficient
Contrafold:
0.508
Cylofold:
0.155
Sensitivity
Contrafold:
0.481
Cylofold:
0.040
Positive Predictive Value
Contrafold:
0.537
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Pknots
Matthews Correlation Coefficient
Contrafold:
0.508
Pknots:
0.143
Sensitivity
Contrafold:
0.481
Pknots:
0.033
Positive Predictive Value
Contrafold:
0.537
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Mastr(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Mastr(20):
0.134
Sensitivity
Contrafold:
0.481
Mastr(20):
0.023
Positive Predictive Value
Contrafold:
0.537
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Alterna
Matthews Correlation Coefficient
Contrafold:
0.508
Alterna:
0.129
Sensitivity
Contrafold:
0.481
Alterna:
0.030
Positive Predictive Value
Contrafold:
0.537
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MCFold
Matthews Correlation Coefficient
Contrafold:
0.508
MCFold:
0.125
Sensitivity
Contrafold:
0.481
MCFold:
0.036
Positive Predictive Value
Contrafold:
0.537
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RDfolder
Matthews Correlation Coefficient
Contrafold:
0.508
RDfolder:
0.122
Sensitivity
Contrafold:
0.481
RDfolder:
0.025
Positive Predictive Value
Contrafold:
0.537
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Murlet(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Murlet(seed):
0.079
Sensitivity
Contrafold:
0.481
Murlet(seed):
0.008
Positive Predictive Value
Contrafold:
0.537
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CMfinder(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
CMfinder(seed):
0.080
Sensitivity
Contrafold:
0.481
CMfinder(seed):
0.008
Positive Predictive Value
Contrafold:
0.537
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs TurboFold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
TurboFold(seed):
0.076
Sensitivity
Contrafold:
0.481
TurboFold(seed):
0.007
Positive Predictive Value
Contrafold:
0.537
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PPfold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
PPfold(seed):
0.069
Sensitivity
Contrafold:
0.481
PPfold(seed):
0.006
Positive Predictive Value
Contrafold:
0.537
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Carnac(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Carnac(seed):
0.066
Sensitivity
Contrafold:
0.481
Carnac(seed):
0.005
Positive Predictive Value
Contrafold:
0.537
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNASampler(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
RNASampler(seed):
0.063
Sensitivity
Contrafold:
0.481
RNASampler(seed):
0.005
Positive Predictive Value
Contrafold:
0.537
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs NanoFolder
Matthews Correlation Coefficient
Contrafold:
0.508
NanoFolder:
0.052
Sensitivity
Contrafold:
0.481
NanoFolder:
0.016
Positive Predictive Value
Contrafold:
0.537
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Mastr(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Mastr(seed):
0.051
Sensitivity
Contrafold:
0.481
Mastr(seed):
0.003
Positive Predictive Value
Contrafold:
0.537
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Multilign(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Multilign(seed):
0.043
Sensitivity
Contrafold:
0.481
Multilign(seed):
0.003
Positive Predictive Value
Contrafold:
0.537
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidFold |
1987
ContextFold vs CentroidFold
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidFold:
0.470
Sensitivity
ContextFold:
0.644
CentroidFold:
0.369
Positive Predictive Value
ContextFold:
0.718
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CentroidFold
Matthews Correlation Coefficient
IPknot:
0.548
CentroidFold:
0.470
Sensitivity
IPknot:
0.475
CentroidFold:
0.369
Positive Predictive Value
IPknot:
0.632
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CentroidFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidFold:
0.470
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidFold:
0.369
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CentroidFold
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidFold:
0.470
Sensitivity
Contrafold:
0.481
CentroidFold:
0.369
Positive Predictive Value
Contrafold:
0.537
CentroidFold:
0.600
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
CentroidFold vs Sfold
Matthews Correlation Coefficient
CentroidFold:
0.470
Sfold:
0.469
Sensitivity
CentroidFold:
0.369
Sfold:
0.412
Positive Predictive Value
CentroidFold:
0.600
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.000451186043411
|
+
CentroidFold vs MaxExpect
Matthews Correlation Coefficient
CentroidFold:
0.470
MaxExpect:
0.465
Sensitivity
CentroidFold:
0.369
MaxExpect:
0.443
Positive Predictive Value
CentroidFold:
0.600
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs ProbKnot
Matthews Correlation Coefficient
CentroidFold:
0.470
ProbKnot:
0.460
Sensitivity
CentroidFold:
0.369
ProbKnot:
0.446
Positive Predictive Value
CentroidFold:
0.600
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs UNAFold
Matthews Correlation Coefficient
CentroidFold:
0.470
UNAFold:
0.445
Sensitivity
CentroidFold:
0.369
UNAFold:
0.445
Positive Predictive Value
CentroidFold:
0.600
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
CentroidAlifold(seed):
0.438
Sensitivity
CentroidFold:
0.369
CentroidAlifold(seed):
0.213
Positive Predictive Value
CentroidFold:
0.600
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Fold
Matthews Correlation Coefficient
CentroidFold:
0.470
Fold:
0.437
Sensitivity
CentroidFold:
0.369
Fold:
0.432
Positive Predictive Value
CentroidFold:
0.600
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAfold
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAfold:
0.429
Sensitivity
CentroidFold:
0.369
RNAfold:
0.422
Positive Predictive Value
CentroidFold:
0.600
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidFold:
0.470
CentroidHomfold‑LAST:
0.428
Sensitivity
CentroidFold:
0.369
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
CentroidFold:
0.600
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
PETfold_pre2.0(20):
0.413
Sensitivity
CentroidFold:
0.369
PETfold_pre2.0(20):
0.211
Positive Predictive Value
CentroidFold:
0.600
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
CentroidAlifold(20):
0.411
Sensitivity
CentroidFold:
0.369
CentroidAlifold(20):
0.190
Positive Predictive Value
CentroidFold:
0.600
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PknotsRG
Matthews Correlation Coefficient
CentroidFold:
0.470
PknotsRG:
0.386
Sensitivity
CentroidFold:
0.369
PknotsRG:
0.320
Positive Predictive Value
CentroidFold:
0.600
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
MXScarna(seed):
0.385
Sensitivity
CentroidFold:
0.369
MXScarna(seed):
0.196
Positive Predictive Value
CentroidFold:
0.600
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAalifold(20):
0.381
Sensitivity
CentroidFold:
0.369
RNAalifold(20):
0.185
Positive Predictive Value
CentroidFold:
0.600
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAalifold(seed):
0.375
Sensitivity
CentroidFold:
0.369
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidFold:
0.600
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Afold
Matthews Correlation Coefficient
CentroidFold:
0.470
Afold:
0.358
Sensitivity
CentroidFold:
0.369
Afold:
0.295
Positive Predictive Value
CentroidFold:
0.600
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MXScarna(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
MXScarna(20):
0.359
Sensitivity
CentroidFold:
0.369
MXScarna(20):
0.181
Positive Predictive Value
CentroidFold:
0.600
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CRWrnafold
Matthews Correlation Coefficient
CentroidFold:
0.470
CRWrnafold:
0.346
Sensitivity
CentroidFold:
0.369
CRWrnafold:
0.275
Positive Predictive Value
CentroidFold:
0.600
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAsubopt
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAsubopt:
0.336
Sensitivity
CentroidFold:
0.369
RNAsubopt:
0.217
Positive Predictive Value
CentroidFold:
0.600
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs McQFold
Matthews Correlation Coefficient
CentroidFold:
0.470
McQFold:
0.314
Sensitivity
CentroidFold:
0.369
McQFold:
0.205
Positive Predictive Value
CentroidFold:
0.600
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs TurboFold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
TurboFold(20):
0.301
Sensitivity
CentroidFold:
0.369
TurboFold(20):
0.108
Positive Predictive Value
CentroidFold:
0.600
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PPfold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
PPfold(20):
0.298
Sensitivity
CentroidFold:
0.369
PPfold(20):
0.101
Positive Predictive Value
CentroidFold:
0.600
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAshapes
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAshapes:
0.295
Sensitivity
CentroidFold:
0.369
RNAshapes:
0.180
Positive Predictive Value
CentroidFold:
0.600
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Carnac(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Carnac(20):
0.288
Sensitivity
CentroidFold:
0.369
Carnac(20):
0.093
Positive Predictive Value
CentroidFold:
0.600
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNASLOpt
Matthews Correlation Coefficient
CentroidFold:
0.470
RNASLOpt:
0.281
Sensitivity
CentroidFold:
0.369
RNASLOpt:
0.158
Positive Predictive Value
CentroidFold:
0.600
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RSpredict(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
RSpredict(20):
0.270
Sensitivity
CentroidFold:
0.369
RSpredict(20):
0.124
Positive Predictive Value
CentroidFold:
0.600
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Multilign(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Multilign(20):
0.251
Sensitivity
CentroidFold:
0.369
Multilign(20):
0.083
Positive Predictive Value
CentroidFold:
0.600
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAwolf
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAwolf:
0.251
Sensitivity
CentroidFold:
0.369
RNAwolf:
0.191
Positive Predictive Value
CentroidFold:
0.600
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Murlet(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Murlet(20):
0.230
Sensitivity
CentroidFold:
0.369
Murlet(20):
0.067
Positive Predictive Value
CentroidFold:
0.600
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Vsfold4
Matthews Correlation Coefficient
CentroidFold:
0.470
Vsfold4:
0.217
Sensitivity
CentroidFold:
0.369
Vsfold4:
0.125
Positive Predictive Value
CentroidFold:
0.600
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Vsfold5
Matthews Correlation Coefficient
CentroidFold:
0.470
Vsfold5:
0.211
Sensitivity
CentroidFold:
0.369
Vsfold5:
0.125
Positive Predictive Value
CentroidFold:
0.600
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
RSpredict(seed):
0.212
Sensitivity
CentroidFold:
0.369
RSpredict(seed):
0.075
Positive Predictive Value
CentroidFold:
0.600
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CMfinder(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
CMfinder(20):
0.210
Sensitivity
CentroidFold:
0.369
CMfinder(20):
0.055
Positive Predictive Value
CentroidFold:
0.600
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs HotKnots
Matthews Correlation Coefficient
CentroidFold:
0.470
HotKnots:
0.179
Sensitivity
CentroidFold:
0.369
HotKnots:
0.051
Positive Predictive Value
CentroidFold:
0.600
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNASampler(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNASampler(20):
0.176
Sensitivity
CentroidFold:
0.369
RNASampler(20):
0.035
Positive Predictive Value
CentroidFold:
0.600
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Cylofold
Matthews Correlation Coefficient
CentroidFold:
0.470
Cylofold:
0.155
Sensitivity
CentroidFold:
0.369
Cylofold:
0.040
Positive Predictive Value
CentroidFold:
0.600
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Pknots
Matthews Correlation Coefficient
CentroidFold:
0.470
Pknots:
0.143
Sensitivity
CentroidFold:
0.369
Pknots:
0.033
Positive Predictive Value
CentroidFold:
0.600
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Mastr(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Mastr(20):
0.134
Sensitivity
CentroidFold:
0.369
Mastr(20):
0.023
Positive Predictive Value
CentroidFold:
0.600
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Alterna
Matthews Correlation Coefficient
CentroidFold:
0.470
Alterna:
0.129
Sensitivity
CentroidFold:
0.369
Alterna:
0.030
Positive Predictive Value
CentroidFold:
0.600
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MCFold
Matthews Correlation Coefficient
CentroidFold:
0.470
MCFold:
0.125
Sensitivity
CentroidFold:
0.369
MCFold:
0.036
Positive Predictive Value
CentroidFold:
0.600
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RDfolder
Matthews Correlation Coefficient
CentroidFold:
0.470
RDfolder:
0.122
Sensitivity
CentroidFold:
0.369
RDfolder:
0.025
Positive Predictive Value
CentroidFold:
0.600
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Murlet(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Murlet(seed):
0.079
Sensitivity
CentroidFold:
0.369
Murlet(seed):
0.008
Positive Predictive Value
CentroidFold:
0.600
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
CMfinder(seed):
0.080
Sensitivity
CentroidFold:
0.369
CMfinder(seed):
0.008
Positive Predictive Value
CentroidFold:
0.600
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
TurboFold(seed):
0.076
Sensitivity
CentroidFold:
0.369
TurboFold(seed):
0.007
Positive Predictive Value
CentroidFold:
0.600
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PPfold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
PPfold(seed):
0.069
Sensitivity
CentroidFold:
0.369
PPfold(seed):
0.006
Positive Predictive Value
CentroidFold:
0.600
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Carnac(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Carnac(seed):
0.066
Sensitivity
CentroidFold:
0.369
Carnac(seed):
0.005
Positive Predictive Value
CentroidFold:
0.600
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNASampler(seed):
0.063
Sensitivity
CentroidFold:
0.369
RNASampler(seed):
0.005
Positive Predictive Value
CentroidFold:
0.600
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs NanoFolder
Matthews Correlation Coefficient
CentroidFold:
0.470
NanoFolder:
0.052
Sensitivity
CentroidFold:
0.369
NanoFolder:
0.016
Positive Predictive Value
CentroidFold:
0.600
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Mastr(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Mastr(seed):
0.051
Sensitivity
CentroidFold:
0.369
Mastr(seed):
0.003
Positive Predictive Value
CentroidFold:
0.600
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Multilign(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Multilign(seed):
0.043
Sensitivity
CentroidFold:
0.369
Multilign(seed):
0.003
Positive Predictive Value
CentroidFold:
0.600
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Sfold |
1987
ContextFold vs Sfold
Matthews Correlation Coefficient
ContextFold:
0.680
Sfold:
0.469
Sensitivity
ContextFold:
0.644
Sfold:
0.412
Positive Predictive Value
ContextFold:
0.718
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Sfold
Matthews Correlation Coefficient
IPknot:
0.548
Sfold:
0.469
Sensitivity
IPknot:
0.475
Sfold:
0.412
Positive Predictive Value
IPknot:
0.632
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Sfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Sfold:
0.469
Sensitivity
PETfold_pre2.0(seed):
0.343
Sfold:
0.412
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Sfold
Matthews Correlation Coefficient
Contrafold:
0.508
Sfold:
0.469
Sensitivity
Contrafold:
0.481
Sfold:
0.412
Positive Predictive Value
Contrafold:
0.537
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Sfold
Matthews Correlation Coefficient
CentroidFold:
0.470
Sfold:
0.469
Sensitivity
CentroidFold:
0.369
Sfold:
0.412
Positive Predictive Value
CentroidFold:
0.600
Sfold:
0.535
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.000451186043411
|
|
+
Sfold vs MaxExpect
Matthews Correlation Coefficient
Sfold:
0.469
MaxExpect:
0.465
Sensitivity
Sfold:
0.412
MaxExpect:
0.443
Positive Predictive Value
Sfold:
0.535
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs ProbKnot
Matthews Correlation Coefficient
Sfold:
0.469
ProbKnot:
0.460
Sensitivity
Sfold:
0.412
ProbKnot:
0.446
Positive Predictive Value
Sfold:
0.535
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs UNAFold
Matthews Correlation Coefficient
Sfold:
0.469
UNAFold:
0.445
Sensitivity
Sfold:
0.412
UNAFold:
0.445
Positive Predictive Value
Sfold:
0.535
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
CentroidAlifold(seed):
0.438
Sensitivity
Sfold:
0.412
CentroidAlifold(seed):
0.213
Positive Predictive Value
Sfold:
0.535
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Fold
Matthews Correlation Coefficient
Sfold:
0.469
Fold:
0.437
Sensitivity
Sfold:
0.412
Fold:
0.432
Positive Predictive Value
Sfold:
0.535
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAfold
Matthews Correlation Coefficient
Sfold:
0.469
RNAfold:
0.429
Sensitivity
Sfold:
0.412
RNAfold:
0.422
Positive Predictive Value
Sfold:
0.535
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
Sfold:
0.469
CentroidHomfold‑LAST:
0.428
Sensitivity
Sfold:
0.412
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
Sfold:
0.535
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
Sfold:
0.469
PETfold_pre2.0(20):
0.413
Sensitivity
Sfold:
0.412
PETfold_pre2.0(20):
0.211
Positive Predictive Value
Sfold:
0.535
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CentroidAlifold(20)
Matthews Correlation Coefficient
Sfold:
0.469
CentroidAlifold(20):
0.411
Sensitivity
Sfold:
0.412
CentroidAlifold(20):
0.190
Positive Predictive Value
Sfold:
0.535
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PknotsRG
Matthews Correlation Coefficient
Sfold:
0.469
PknotsRG:
0.386
Sensitivity
Sfold:
0.412
PknotsRG:
0.320
Positive Predictive Value
Sfold:
0.535
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs MXScarna(seed)
Matthews Correlation Coefficient
Sfold:
0.469
MXScarna(seed):
0.385
Sensitivity
Sfold:
0.412
MXScarna(seed):
0.196
Positive Predictive Value
Sfold:
0.535
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAalifold(20)
Matthews Correlation Coefficient
Sfold:
0.469
RNAalifold(20):
0.381
Sensitivity
Sfold:
0.412
RNAalifold(20):
0.185
Positive Predictive Value
Sfold:
0.535
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAalifold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
RNAalifold(seed):
0.375
Sensitivity
Sfold:
0.412
RNAalifold(seed):
0.169
Positive Predictive Value
Sfold:
0.535
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Afold
Matthews Correlation Coefficient
Sfold:
0.469
Afold:
0.358
Sensitivity
Sfold:
0.412
Afold:
0.295
Positive Predictive Value
Sfold:
0.535
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs MXScarna(20)
Matthews Correlation Coefficient
Sfold:
0.469
MXScarna(20):
0.359
Sensitivity
Sfold:
0.412
MXScarna(20):
0.181
Positive Predictive Value
Sfold:
0.535
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CRWrnafold
Matthews Correlation Coefficient
Sfold:
0.469
CRWrnafold:
0.346
Sensitivity
Sfold:
0.412
CRWrnafold:
0.275
Positive Predictive Value
Sfold:
0.535
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAsubopt
Matthews Correlation Coefficient
Sfold:
0.469
RNAsubopt:
0.336
Sensitivity
Sfold:
0.412
RNAsubopt:
0.217
Positive Predictive Value
Sfold:
0.535
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs McQFold
Matthews Correlation Coefficient
Sfold:
0.469
McQFold:
0.314
Sensitivity
Sfold:
0.412
McQFold:
0.205
Positive Predictive Value
Sfold:
0.535
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs TurboFold(20)
Matthews Correlation Coefficient
Sfold:
0.469
TurboFold(20):
0.301
Sensitivity
Sfold:
0.412
TurboFold(20):
0.108
Positive Predictive Value
Sfold:
0.535
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PPfold(20)
Matthews Correlation Coefficient
Sfold:
0.469
PPfold(20):
0.298
Sensitivity
Sfold:
0.412
PPfold(20):
0.101
Positive Predictive Value
Sfold:
0.535
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAshapes
Matthews Correlation Coefficient
Sfold:
0.469
RNAshapes:
0.295
Sensitivity
Sfold:
0.412
RNAshapes:
0.180
Positive Predictive Value
Sfold:
0.535
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Carnac(20)
Matthews Correlation Coefficient
Sfold:
0.469
Carnac(20):
0.288
Sensitivity
Sfold:
0.412
Carnac(20):
0.093
Positive Predictive Value
Sfold:
0.535
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNASLOpt
Matthews Correlation Coefficient
Sfold:
0.469
RNASLOpt:
0.281
Sensitivity
Sfold:
0.412
RNASLOpt:
0.158
Positive Predictive Value
Sfold:
0.535
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RSpredict(20)
Matthews Correlation Coefficient
Sfold:
0.469
RSpredict(20):
0.270
Sensitivity
Sfold:
0.412
RSpredict(20):
0.124
Positive Predictive Value
Sfold:
0.535
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Multilign(20)
Matthews Correlation Coefficient
Sfold:
0.469
Multilign(20):
0.251
Sensitivity
Sfold:
0.412
Multilign(20):
0.083
Positive Predictive Value
Sfold:
0.535
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAwolf
Matthews Correlation Coefficient
Sfold:
0.469
RNAwolf:
0.251
Sensitivity
Sfold:
0.412
RNAwolf:
0.191
Positive Predictive Value
Sfold:
0.535
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Murlet(20)
Matthews Correlation Coefficient
Sfold:
0.469
Murlet(20):
0.230
Sensitivity
Sfold:
0.412
Murlet(20):
0.067
Positive Predictive Value
Sfold:
0.535
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Vsfold4
Matthews Correlation Coefficient
Sfold:
0.469
Vsfold4:
0.217
Sensitivity
Sfold:
0.412
Vsfold4:
0.125
Positive Predictive Value
Sfold:
0.535
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Vsfold5
Matthews Correlation Coefficient
Sfold:
0.469
Vsfold5:
0.211
Sensitivity
Sfold:
0.412
Vsfold5:
0.125
Positive Predictive Value
Sfold:
0.535
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RSpredict(seed)
Matthews Correlation Coefficient
Sfold:
0.469
RSpredict(seed):
0.212
Sensitivity
Sfold:
0.412
RSpredict(seed):
0.075
Positive Predictive Value
Sfold:
0.535
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CMfinder(20)
Matthews Correlation Coefficient
Sfold:
0.469
CMfinder(20):
0.210
Sensitivity
Sfold:
0.412
CMfinder(20):
0.055
Positive Predictive Value
Sfold:
0.535
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs HotKnots
Matthews Correlation Coefficient
Sfold:
0.469
HotKnots:
0.179
Sensitivity
Sfold:
0.412
HotKnots:
0.051
Positive Predictive Value
Sfold:
0.535
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNASampler(20)
Matthews Correlation Coefficient
Sfold:
0.469
RNASampler(20):
0.176
Sensitivity
Sfold:
0.412
RNASampler(20):
0.035
Positive Predictive Value
Sfold:
0.535
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Cylofold
Matthews Correlation Coefficient
Sfold:
0.469
Cylofold:
0.155
Sensitivity
Sfold:
0.412
Cylofold:
0.040
Positive Predictive Value
Sfold:
0.535
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Pknots
Matthews Correlation Coefficient
Sfold:
0.469
Pknots:
0.143
Sensitivity
Sfold:
0.412
Pknots:
0.033
Positive Predictive Value
Sfold:
0.535
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Mastr(20)
Matthews Correlation Coefficient
Sfold:
0.469
Mastr(20):
0.134
Sensitivity
Sfold:
0.412
Mastr(20):
0.023
Positive Predictive Value
Sfold:
0.535
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Alterna
Matthews Correlation Coefficient
Sfold:
0.469
Alterna:
0.129
Sensitivity
Sfold:
0.412
Alterna:
0.030
Positive Predictive Value
Sfold:
0.535
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs MCFold
Matthews Correlation Coefficient
Sfold:
0.469
MCFold:
0.125
Sensitivity
Sfold:
0.412
MCFold:
0.036
Positive Predictive Value
Sfold:
0.535
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RDfolder
Matthews Correlation Coefficient
Sfold:
0.469
RDfolder:
0.122
Sensitivity
Sfold:
0.412
RDfolder:
0.025
Positive Predictive Value
Sfold:
0.535
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Murlet(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Murlet(seed):
0.079
Sensitivity
Sfold:
0.412
Murlet(seed):
0.008
Positive Predictive Value
Sfold:
0.535
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CMfinder(seed)
Matthews Correlation Coefficient
Sfold:
0.469
CMfinder(seed):
0.080
Sensitivity
Sfold:
0.412
CMfinder(seed):
0.008
Positive Predictive Value
Sfold:
0.535
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs TurboFold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
TurboFold(seed):
0.076
Sensitivity
Sfold:
0.412
TurboFold(seed):
0.007
Positive Predictive Value
Sfold:
0.535
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PPfold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
PPfold(seed):
0.069
Sensitivity
Sfold:
0.412
PPfold(seed):
0.006
Positive Predictive Value
Sfold:
0.535
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Carnac(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Carnac(seed):
0.066
Sensitivity
Sfold:
0.412
Carnac(seed):
0.005
Positive Predictive Value
Sfold:
0.535
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNASampler(seed)
Matthews Correlation Coefficient
Sfold:
0.469
RNASampler(seed):
0.063
Sensitivity
Sfold:
0.412
RNASampler(seed):
0.005
Positive Predictive Value
Sfold:
0.535
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs NanoFolder
Matthews Correlation Coefficient
Sfold:
0.469
NanoFolder:
0.052
Sensitivity
Sfold:
0.412
NanoFolder:
0.016
Positive Predictive Value
Sfold:
0.535
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Mastr(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Mastr(seed):
0.051
Sensitivity
Sfold:
0.412
Mastr(seed):
0.003
Positive Predictive Value
Sfold:
0.535
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Multilign(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Multilign(seed):
0.043
Sensitivity
Sfold:
0.412
Multilign(seed):
0.003
Positive Predictive Value
Sfold:
0.535
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MaxExpect |
1987
ContextFold vs MaxExpect
Matthews Correlation Coefficient
ContextFold:
0.680
MaxExpect:
0.465
Sensitivity
ContextFold:
0.644
MaxExpect:
0.443
Positive Predictive Value
ContextFold:
0.718
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs MaxExpect
Matthews Correlation Coefficient
IPknot:
0.548
MaxExpect:
0.465
Sensitivity
IPknot:
0.475
MaxExpect:
0.443
Positive Predictive Value
IPknot:
0.632
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs MaxExpect
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MaxExpect:
0.465
Sensitivity
PETfold_pre2.0(seed):
0.343
MaxExpect:
0.443
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs MaxExpect
Matthews Correlation Coefficient
Contrafold:
0.508
MaxExpect:
0.465
Sensitivity
Contrafold:
0.481
MaxExpect:
0.443
Positive Predictive Value
Contrafold:
0.537
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs MaxExpect
Matthews Correlation Coefficient
CentroidFold:
0.470
MaxExpect:
0.465
Sensitivity
CentroidFold:
0.369
MaxExpect:
0.443
Positive Predictive Value
CentroidFold:
0.600
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs MaxExpect
Matthews Correlation Coefficient
Sfold:
0.469
MaxExpect:
0.465
Sensitivity
Sfold:
0.412
MaxExpect:
0.443
Positive Predictive Value
Sfold:
0.535
MaxExpect:
0.488
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
MaxExpect vs ProbKnot
Matthews Correlation Coefficient
MaxExpect:
0.465
ProbKnot:
0.460
Sensitivity
MaxExpect:
0.443
ProbKnot:
0.446
Positive Predictive Value
MaxExpect:
0.488
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs UNAFold
Matthews Correlation Coefficient
MaxExpect:
0.465
UNAFold:
0.445
Sensitivity
MaxExpect:
0.443
UNAFold:
0.445
Positive Predictive Value
MaxExpect:
0.488
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CentroidAlifold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
CentroidAlifold(seed):
0.438
Sensitivity
MaxExpect:
0.443
CentroidAlifold(seed):
0.213
Positive Predictive Value
MaxExpect:
0.488
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Fold
Matthews Correlation Coefficient
MaxExpect:
0.465
Fold:
0.437
Sensitivity
MaxExpect:
0.443
Fold:
0.432
Positive Predictive Value
MaxExpect:
0.488
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAfold
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAfold:
0.429
Sensitivity
MaxExpect:
0.443
RNAfold:
0.422
Positive Predictive Value
MaxExpect:
0.488
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
MaxExpect:
0.465
CentroidHomfold‑LAST:
0.428
Sensitivity
MaxExpect:
0.443
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
MaxExpect:
0.488
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
PETfold_pre2.0(20):
0.413
Sensitivity
MaxExpect:
0.443
PETfold_pre2.0(20):
0.211
Positive Predictive Value
MaxExpect:
0.488
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CentroidAlifold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
CentroidAlifold(20):
0.411
Sensitivity
MaxExpect:
0.443
CentroidAlifold(20):
0.190
Positive Predictive Value
MaxExpect:
0.488
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PknotsRG
Matthews Correlation Coefficient
MaxExpect:
0.465
PknotsRG:
0.386
Sensitivity
MaxExpect:
0.443
PknotsRG:
0.320
Positive Predictive Value
MaxExpect:
0.488
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs MXScarna(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
MXScarna(seed):
0.385
Sensitivity
MaxExpect:
0.443
MXScarna(seed):
0.196
Positive Predictive Value
MaxExpect:
0.488
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAalifold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAalifold(20):
0.381
Sensitivity
MaxExpect:
0.443
RNAalifold(20):
0.185
Positive Predictive Value
MaxExpect:
0.488
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAalifold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAalifold(seed):
0.375
Sensitivity
MaxExpect:
0.443
RNAalifold(seed):
0.169
Positive Predictive Value
MaxExpect:
0.488
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Afold
Matthews Correlation Coefficient
MaxExpect:
0.465
Afold:
0.358
Sensitivity
MaxExpect:
0.443
Afold:
0.295
Positive Predictive Value
MaxExpect:
0.488
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs MXScarna(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
MXScarna(20):
0.359
Sensitivity
MaxExpect:
0.443
MXScarna(20):
0.181
Positive Predictive Value
MaxExpect:
0.488
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CRWrnafold
Matthews Correlation Coefficient
MaxExpect:
0.465
CRWrnafold:
0.346
Sensitivity
MaxExpect:
0.443
CRWrnafold:
0.275
Positive Predictive Value
MaxExpect:
0.488
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAsubopt
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAsubopt:
0.336
Sensitivity
MaxExpect:
0.443
RNAsubopt:
0.217
Positive Predictive Value
MaxExpect:
0.488
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs McQFold
Matthews Correlation Coefficient
MaxExpect:
0.465
McQFold:
0.314
Sensitivity
MaxExpect:
0.443
McQFold:
0.205
Positive Predictive Value
MaxExpect:
0.488
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs TurboFold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
TurboFold(20):
0.301
Sensitivity
MaxExpect:
0.443
TurboFold(20):
0.108
Positive Predictive Value
MaxExpect:
0.488
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PPfold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
PPfold(20):
0.298
Sensitivity
MaxExpect:
0.443
PPfold(20):
0.101
Positive Predictive Value
MaxExpect:
0.488
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAshapes
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAshapes:
0.295
Sensitivity
MaxExpect:
0.443
RNAshapes:
0.180
Positive Predictive Value
MaxExpect:
0.488
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Carnac(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Carnac(20):
0.288
Sensitivity
MaxExpect:
0.443
Carnac(20):
0.093
Positive Predictive Value
MaxExpect:
0.488
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNASLOpt
Matthews Correlation Coefficient
MaxExpect:
0.465
RNASLOpt:
0.281
Sensitivity
MaxExpect:
0.443
RNASLOpt:
0.158
Positive Predictive Value
MaxExpect:
0.488
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RSpredict(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
RSpredict(20):
0.270
Sensitivity
MaxExpect:
0.443
RSpredict(20):
0.124
Positive Predictive Value
MaxExpect:
0.488
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Multilign(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Multilign(20):
0.251
Sensitivity
MaxExpect:
0.443
Multilign(20):
0.083
Positive Predictive Value
MaxExpect:
0.488
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAwolf
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAwolf:
0.251
Sensitivity
MaxExpect:
0.443
RNAwolf:
0.191
Positive Predictive Value
MaxExpect:
0.488
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Murlet(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Murlet(20):
0.230
Sensitivity
MaxExpect:
0.443
Murlet(20):
0.067
Positive Predictive Value
MaxExpect:
0.488
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Vsfold4
Matthews Correlation Coefficient
MaxExpect:
0.465
Vsfold4:
0.217
Sensitivity
MaxExpect:
0.443
Vsfold4:
0.125
Positive Predictive Value
MaxExpect:
0.488
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Vsfold5
Matthews Correlation Coefficient
MaxExpect:
0.465
Vsfold5:
0.211
Sensitivity
MaxExpect:
0.443
Vsfold5:
0.125
Positive Predictive Value
MaxExpect:
0.488
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RSpredict(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
RSpredict(seed):
0.212
Sensitivity
MaxExpect:
0.443
RSpredict(seed):
0.075
Positive Predictive Value
MaxExpect:
0.488
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CMfinder(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
CMfinder(20):
0.210
Sensitivity
MaxExpect:
0.443
CMfinder(20):
0.055
Positive Predictive Value
MaxExpect:
0.488
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs HotKnots
Matthews Correlation Coefficient
MaxExpect:
0.465
HotKnots:
0.179
Sensitivity
MaxExpect:
0.443
HotKnots:
0.051
Positive Predictive Value
MaxExpect:
0.488
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNASampler(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNASampler(20):
0.176
Sensitivity
MaxExpect:
0.443
RNASampler(20):
0.035
Positive Predictive Value
MaxExpect:
0.488
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Cylofold
Matthews Correlation Coefficient
MaxExpect:
0.465
Cylofold:
0.155
Sensitivity
MaxExpect:
0.443
Cylofold:
0.040
Positive Predictive Value
MaxExpect:
0.488
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Pknots
Matthews Correlation Coefficient
MaxExpect:
0.465
Pknots:
0.143
Sensitivity
MaxExpect:
0.443
Pknots:
0.033
Positive Predictive Value
MaxExpect:
0.488
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Mastr(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Mastr(20):
0.134
Sensitivity
MaxExpect:
0.443
Mastr(20):
0.023
Positive Predictive Value
MaxExpect:
0.488
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Alterna
Matthews Correlation Coefficient
MaxExpect:
0.465
Alterna:
0.129
Sensitivity
MaxExpect:
0.443
Alterna:
0.030
Positive Predictive Value
MaxExpect:
0.488
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs MCFold
Matthews Correlation Coefficient
MaxExpect:
0.465
MCFold:
0.125
Sensitivity
MaxExpect:
0.443
MCFold:
0.036
Positive Predictive Value
MaxExpect:
0.488
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RDfolder
Matthews Correlation Coefficient
MaxExpect:
0.465
RDfolder:
0.122
Sensitivity
MaxExpect:
0.443
RDfolder:
0.025
Positive Predictive Value
MaxExpect:
0.488
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Murlet(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Murlet(seed):
0.079
Sensitivity
MaxExpect:
0.443
Murlet(seed):
0.008
Positive Predictive Value
MaxExpect:
0.488
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CMfinder(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
CMfinder(seed):
0.080
Sensitivity
MaxExpect:
0.443
CMfinder(seed):
0.008
Positive Predictive Value
MaxExpect:
0.488
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs TurboFold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
TurboFold(seed):
0.076
Sensitivity
MaxExpect:
0.443
TurboFold(seed):
0.007
Positive Predictive Value
MaxExpect:
0.488
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PPfold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
PPfold(seed):
0.069
Sensitivity
MaxExpect:
0.443
PPfold(seed):
0.006
Positive Predictive Value
MaxExpect:
0.488
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Carnac(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Carnac(seed):
0.066
Sensitivity
MaxExpect:
0.443
Carnac(seed):
0.005
Positive Predictive Value
MaxExpect:
0.488
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNASampler(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNASampler(seed):
0.063
Sensitivity
MaxExpect:
0.443
RNASampler(seed):
0.005
Positive Predictive Value
MaxExpect:
0.488
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs NanoFolder
Matthews Correlation Coefficient
MaxExpect:
0.465
NanoFolder:
0.052
Sensitivity
MaxExpect:
0.443
NanoFolder:
0.016
Positive Predictive Value
MaxExpect:
0.488
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Mastr(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Mastr(seed):
0.051
Sensitivity
MaxExpect:
0.443
Mastr(seed):
0.003
Positive Predictive Value
MaxExpect:
0.488
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Multilign(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Multilign(seed):
0.043
Sensitivity
MaxExpect:
0.443
Multilign(seed):
0.003
Positive Predictive Value
MaxExpect:
0.488
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| ProbKnot |
1987
ContextFold vs ProbKnot
Matthews Correlation Coefficient
ContextFold:
0.680
ProbKnot:
0.460
Sensitivity
ContextFold:
0.644
ProbKnot:
0.446
Positive Predictive Value
ContextFold:
0.718
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs ProbKnot
Matthews Correlation Coefficient
IPknot:
0.548
ProbKnot:
0.460
Sensitivity
IPknot:
0.475
ProbKnot:
0.446
Positive Predictive Value
IPknot:
0.632
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs ProbKnot
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
ProbKnot:
0.460
Sensitivity
PETfold_pre2.0(seed):
0.343
ProbKnot:
0.446
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs ProbKnot
Matthews Correlation Coefficient
Contrafold:
0.508
ProbKnot:
0.460
Sensitivity
Contrafold:
0.481
ProbKnot:
0.446
Positive Predictive Value
Contrafold:
0.537
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs ProbKnot
Matthews Correlation Coefficient
CentroidFold:
0.470
ProbKnot:
0.460
Sensitivity
CentroidFold:
0.369
ProbKnot:
0.446
Positive Predictive Value
CentroidFold:
0.600
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs ProbKnot
Matthews Correlation Coefficient
Sfold:
0.469
ProbKnot:
0.460
Sensitivity
Sfold:
0.412
ProbKnot:
0.446
Positive Predictive Value
Sfold:
0.535
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs ProbKnot
Matthews Correlation Coefficient
MaxExpect:
0.465
ProbKnot:
0.460
Sensitivity
MaxExpect:
0.443
ProbKnot:
0.446
Positive Predictive Value
MaxExpect:
0.488
ProbKnot:
0.475
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
ProbKnot vs UNAFold
Matthews Correlation Coefficient
ProbKnot:
0.460
UNAFold:
0.445
Sensitivity
ProbKnot:
0.446
UNAFold:
0.445
Positive Predictive Value
ProbKnot:
0.475
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CentroidAlifold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
CentroidAlifold(seed):
0.438
Sensitivity
ProbKnot:
0.446
CentroidAlifold(seed):
0.213
Positive Predictive Value
ProbKnot:
0.475
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Fold
Matthews Correlation Coefficient
ProbKnot:
0.460
Fold:
0.437
Sensitivity
ProbKnot:
0.446
Fold:
0.432
Positive Predictive Value
ProbKnot:
0.475
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAfold
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAfold:
0.429
Sensitivity
ProbKnot:
0.446
RNAfold:
0.422
Positive Predictive Value
ProbKnot:
0.475
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
ProbKnot:
0.460
CentroidHomfold‑LAST:
0.428
Sensitivity
ProbKnot:
0.446
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
ProbKnot:
0.475
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
PETfold_pre2.0(20):
0.413
Sensitivity
ProbKnot:
0.446
PETfold_pre2.0(20):
0.211
Positive Predictive Value
ProbKnot:
0.475
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CentroidAlifold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
CentroidAlifold(20):
0.411
Sensitivity
ProbKnot:
0.446
CentroidAlifold(20):
0.190
Positive Predictive Value
ProbKnot:
0.475
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PknotsRG
Matthews Correlation Coefficient
ProbKnot:
0.460
PknotsRG:
0.386
Sensitivity
ProbKnot:
0.446
PknotsRG:
0.320
Positive Predictive Value
ProbKnot:
0.475
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs MXScarna(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
MXScarna(seed):
0.385
Sensitivity
ProbKnot:
0.446
MXScarna(seed):
0.196
Positive Predictive Value
ProbKnot:
0.475
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAalifold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAalifold(20):
0.381
Sensitivity
ProbKnot:
0.446
RNAalifold(20):
0.185
Positive Predictive Value
ProbKnot:
0.475
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAalifold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAalifold(seed):
0.375
Sensitivity
ProbKnot:
0.446
RNAalifold(seed):
0.169
Positive Predictive Value
ProbKnot:
0.475
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Afold
Matthews Correlation Coefficient
ProbKnot:
0.460
Afold:
0.358
Sensitivity
ProbKnot:
0.446
Afold:
0.295
Positive Predictive Value
ProbKnot:
0.475
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs MXScarna(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
MXScarna(20):
0.359
Sensitivity
ProbKnot:
0.446
MXScarna(20):
0.181
Positive Predictive Value
ProbKnot:
0.475
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CRWrnafold
Matthews Correlation Coefficient
ProbKnot:
0.460
CRWrnafold:
0.346
Sensitivity
ProbKnot:
0.446
CRWrnafold:
0.275
Positive Predictive Value
ProbKnot:
0.475
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAsubopt
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAsubopt:
0.336
Sensitivity
ProbKnot:
0.446
RNAsubopt:
0.217
Positive Predictive Value
ProbKnot:
0.475
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs McQFold
Matthews Correlation Coefficient
ProbKnot:
0.460
McQFold:
0.314
Sensitivity
ProbKnot:
0.446
McQFold:
0.205
Positive Predictive Value
ProbKnot:
0.475
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs TurboFold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
TurboFold(20):
0.301
Sensitivity
ProbKnot:
0.446
TurboFold(20):
0.108
Positive Predictive Value
ProbKnot:
0.475
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PPfold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
PPfold(20):
0.298
Sensitivity
ProbKnot:
0.446
PPfold(20):
0.101
Positive Predictive Value
ProbKnot:
0.475
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAshapes
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAshapes:
0.295
Sensitivity
ProbKnot:
0.446
RNAshapes:
0.180
Positive Predictive Value
ProbKnot:
0.475
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Carnac(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Carnac(20):
0.288
Sensitivity
ProbKnot:
0.446
Carnac(20):
0.093
Positive Predictive Value
ProbKnot:
0.475
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNASLOpt
Matthews Correlation Coefficient
ProbKnot:
0.460
RNASLOpt:
0.281
Sensitivity
ProbKnot:
0.446
RNASLOpt:
0.158
Positive Predictive Value
ProbKnot:
0.475
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RSpredict(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
RSpredict(20):
0.270
Sensitivity
ProbKnot:
0.446
RSpredict(20):
0.124
Positive Predictive Value
ProbKnot:
0.475
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Multilign(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Multilign(20):
0.251
Sensitivity
ProbKnot:
0.446
Multilign(20):
0.083
Positive Predictive Value
ProbKnot:
0.475
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAwolf
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAwolf:
0.251
Sensitivity
ProbKnot:
0.446
RNAwolf:
0.191
Positive Predictive Value
ProbKnot:
0.475
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Murlet(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Murlet(20):
0.230
Sensitivity
ProbKnot:
0.446
Murlet(20):
0.067
Positive Predictive Value
ProbKnot:
0.475
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Vsfold4
Matthews Correlation Coefficient
ProbKnot:
0.460
Vsfold4:
0.217
Sensitivity
ProbKnot:
0.446
Vsfold4:
0.125
Positive Predictive Value
ProbKnot:
0.475
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Vsfold5
Matthews Correlation Coefficient
ProbKnot:
0.460
Vsfold5:
0.211
Sensitivity
ProbKnot:
0.446
Vsfold5:
0.125
Positive Predictive Value
ProbKnot:
0.475
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RSpredict(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
RSpredict(seed):
0.212
Sensitivity
ProbKnot:
0.446
RSpredict(seed):
0.075
Positive Predictive Value
ProbKnot:
0.475
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CMfinder(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
CMfinder(20):
0.210
Sensitivity
ProbKnot:
0.446
CMfinder(20):
0.055
Positive Predictive Value
ProbKnot:
0.475
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs HotKnots
Matthews Correlation Coefficient
ProbKnot:
0.460
HotKnots:
0.179
Sensitivity
ProbKnot:
0.446
HotKnots:
0.051
Positive Predictive Value
ProbKnot:
0.475
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNASampler(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNASampler(20):
0.176
Sensitivity
ProbKnot:
0.446
RNASampler(20):
0.035
Positive Predictive Value
ProbKnot:
0.475
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Cylofold
Matthews Correlation Coefficient
ProbKnot:
0.460
Cylofold:
0.155
Sensitivity
ProbKnot:
0.446
Cylofold:
0.040
Positive Predictive Value
ProbKnot:
0.475
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Pknots
Matthews Correlation Coefficient
ProbKnot:
0.460
Pknots:
0.143
Sensitivity
ProbKnot:
0.446
Pknots:
0.033
Positive Predictive Value
ProbKnot:
0.475
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Mastr(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Mastr(20):
0.134
Sensitivity
ProbKnot:
0.446
Mastr(20):
0.023
Positive Predictive Value
ProbKnot:
0.475
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Alterna
Matthews Correlation Coefficient
ProbKnot:
0.460
Alterna:
0.129
Sensitivity
ProbKnot:
0.446
Alterna:
0.030
Positive Predictive Value
ProbKnot:
0.475
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs MCFold
Matthews Correlation Coefficient
ProbKnot:
0.460
MCFold:
0.125
Sensitivity
ProbKnot:
0.446
MCFold:
0.036
Positive Predictive Value
ProbKnot:
0.475
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RDfolder
Matthews Correlation Coefficient
ProbKnot:
0.460
RDfolder:
0.122
Sensitivity
ProbKnot:
0.446
RDfolder:
0.025
Positive Predictive Value
ProbKnot:
0.475
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Murlet(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Murlet(seed):
0.079
Sensitivity
ProbKnot:
0.446
Murlet(seed):
0.008
Positive Predictive Value
ProbKnot:
0.475
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CMfinder(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
CMfinder(seed):
0.080
Sensitivity
ProbKnot:
0.446
CMfinder(seed):
0.008
Positive Predictive Value
ProbKnot:
0.475
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs TurboFold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
TurboFold(seed):
0.076
Sensitivity
ProbKnot:
0.446
TurboFold(seed):
0.007
Positive Predictive Value
ProbKnot:
0.475
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PPfold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
PPfold(seed):
0.069
Sensitivity
ProbKnot:
0.446
PPfold(seed):
0.006
Positive Predictive Value
ProbKnot:
0.475
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Carnac(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Carnac(seed):
0.066
Sensitivity
ProbKnot:
0.446
Carnac(seed):
0.005
Positive Predictive Value
ProbKnot:
0.475
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNASampler(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNASampler(seed):
0.063
Sensitivity
ProbKnot:
0.446
RNASampler(seed):
0.005
Positive Predictive Value
ProbKnot:
0.475
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs NanoFolder
Matthews Correlation Coefficient
ProbKnot:
0.460
NanoFolder:
0.052
Sensitivity
ProbKnot:
0.446
NanoFolder:
0.016
Positive Predictive Value
ProbKnot:
0.475
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Mastr(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Mastr(seed):
0.051
Sensitivity
ProbKnot:
0.446
Mastr(seed):
0.003
Positive Predictive Value
ProbKnot:
0.475
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Multilign(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Multilign(seed):
0.043
Sensitivity
ProbKnot:
0.446
Multilign(seed):
0.003
Positive Predictive Value
ProbKnot:
0.475
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| UNAFold |
1987
ContextFold vs UNAFold
Matthews Correlation Coefficient
ContextFold:
0.680
UNAFold:
0.445
Sensitivity
ContextFold:
0.644
UNAFold:
0.445
Positive Predictive Value
ContextFold:
0.718
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs UNAFold
Matthews Correlation Coefficient
IPknot:
0.548
UNAFold:
0.445
Sensitivity
IPknot:
0.475
UNAFold:
0.445
Positive Predictive Value
IPknot:
0.632
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs UNAFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
UNAFold:
0.445
Sensitivity
PETfold_pre2.0(seed):
0.343
UNAFold:
0.445
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs UNAFold
Matthews Correlation Coefficient
Contrafold:
0.508
UNAFold:
0.445
Sensitivity
Contrafold:
0.481
UNAFold:
0.445
Positive Predictive Value
Contrafold:
0.537
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs UNAFold
Matthews Correlation Coefficient
CentroidFold:
0.470
UNAFold:
0.445
Sensitivity
CentroidFold:
0.369
UNAFold:
0.445
Positive Predictive Value
CentroidFold:
0.600
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs UNAFold
Matthews Correlation Coefficient
Sfold:
0.469
UNAFold:
0.445
Sensitivity
Sfold:
0.412
UNAFold:
0.445
Positive Predictive Value
Sfold:
0.535
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs UNAFold
Matthews Correlation Coefficient
MaxExpect:
0.465
UNAFold:
0.445
Sensitivity
MaxExpect:
0.443
UNAFold:
0.445
Positive Predictive Value
MaxExpect:
0.488
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs UNAFold
Matthews Correlation Coefficient
ProbKnot:
0.460
UNAFold:
0.445
Sensitivity
ProbKnot:
0.446
UNAFold:
0.445
Positive Predictive Value
ProbKnot:
0.475
UNAFold:
0.446
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
UNAFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
CentroidAlifold(seed):
0.438
Sensitivity
UNAFold:
0.445
CentroidAlifold(seed):
0.213
Positive Predictive Value
UNAFold:
0.446
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Fold
Matthews Correlation Coefficient
UNAFold:
0.445
Fold:
0.437
Sensitivity
UNAFold:
0.445
Fold:
0.432
Positive Predictive Value
UNAFold:
0.446
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAfold
Matthews Correlation Coefficient
UNAFold:
0.445
RNAfold:
0.429
Sensitivity
UNAFold:
0.445
RNAfold:
0.422
Positive Predictive Value
UNAFold:
0.446
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
UNAFold:
0.445
CentroidHomfold‑LAST:
0.428
Sensitivity
UNAFold:
0.445
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
UNAFold:
0.446
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
UNAFold:
0.445
PETfold_pre2.0(20):
0.413
Sensitivity
UNAFold:
0.445
PETfold_pre2.0(20):
0.211
Positive Predictive Value
UNAFold:
0.446
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
CentroidAlifold(20):
0.411
Sensitivity
UNAFold:
0.445
CentroidAlifold(20):
0.190
Positive Predictive Value
UNAFold:
0.446
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PknotsRG
Matthews Correlation Coefficient
UNAFold:
0.445
PknotsRG:
0.386
Sensitivity
UNAFold:
0.445
PknotsRG:
0.320
Positive Predictive Value
UNAFold:
0.446
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs MXScarna(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
MXScarna(seed):
0.385
Sensitivity
UNAFold:
0.445
MXScarna(seed):
0.196
Positive Predictive Value
UNAFold:
0.446
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAalifold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
RNAalifold(20):
0.381
Sensitivity
UNAFold:
0.445
RNAalifold(20):
0.185
Positive Predictive Value
UNAFold:
0.446
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAalifold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
RNAalifold(seed):
0.375
Sensitivity
UNAFold:
0.445
RNAalifold(seed):
0.169
Positive Predictive Value
UNAFold:
0.446
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Afold
Matthews Correlation Coefficient
UNAFold:
0.445
Afold:
0.358
Sensitivity
UNAFold:
0.445
Afold:
0.295
Positive Predictive Value
UNAFold:
0.446
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs MXScarna(20)
Matthews Correlation Coefficient
UNAFold:
0.445
MXScarna(20):
0.359
Sensitivity
UNAFold:
0.445
MXScarna(20):
0.181
Positive Predictive Value
UNAFold:
0.446
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CRWrnafold
Matthews Correlation Coefficient
UNAFold:
0.445
CRWrnafold:
0.346
Sensitivity
UNAFold:
0.445
CRWrnafold:
0.275
Positive Predictive Value
UNAFold:
0.446
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAsubopt
Matthews Correlation Coefficient
UNAFold:
0.445
RNAsubopt:
0.336
Sensitivity
UNAFold:
0.445
RNAsubopt:
0.217
Positive Predictive Value
UNAFold:
0.446
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs McQFold
Matthews Correlation Coefficient
UNAFold:
0.445
McQFold:
0.314
Sensitivity
UNAFold:
0.445
McQFold:
0.205
Positive Predictive Value
UNAFold:
0.446
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs TurboFold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
TurboFold(20):
0.301
Sensitivity
UNAFold:
0.445
TurboFold(20):
0.108
Positive Predictive Value
UNAFold:
0.446
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PPfold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
PPfold(20):
0.298
Sensitivity
UNAFold:
0.445
PPfold(20):
0.101
Positive Predictive Value
UNAFold:
0.446
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAshapes
Matthews Correlation Coefficient
UNAFold:
0.445
RNAshapes:
0.295
Sensitivity
UNAFold:
0.445
RNAshapes:
0.180
Positive Predictive Value
UNAFold:
0.446
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Carnac(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Carnac(20):
0.288
Sensitivity
UNAFold:
0.445
Carnac(20):
0.093
Positive Predictive Value
UNAFold:
0.446
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNASLOpt
Matthews Correlation Coefficient
UNAFold:
0.445
RNASLOpt:
0.281
Sensitivity
UNAFold:
0.445
RNASLOpt:
0.158
Positive Predictive Value
UNAFold:
0.446
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RSpredict(20)
Matthews Correlation Coefficient
UNAFold:
0.445
RSpredict(20):
0.270
Sensitivity
UNAFold:
0.445
RSpredict(20):
0.124
Positive Predictive Value
UNAFold:
0.446
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Multilign(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Multilign(20):
0.251
Sensitivity
UNAFold:
0.445
Multilign(20):
0.083
Positive Predictive Value
UNAFold:
0.446
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAwolf
Matthews Correlation Coefficient
UNAFold:
0.445
RNAwolf:
0.251
Sensitivity
UNAFold:
0.445
RNAwolf:
0.191
Positive Predictive Value
UNAFold:
0.446
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Murlet(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Murlet(20):
0.230
Sensitivity
UNAFold:
0.445
Murlet(20):
0.067
Positive Predictive Value
UNAFold:
0.446
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Vsfold4
Matthews Correlation Coefficient
UNAFold:
0.445
Vsfold4:
0.217
Sensitivity
UNAFold:
0.445
Vsfold4:
0.125
Positive Predictive Value
UNAFold:
0.446
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Vsfold5
Matthews Correlation Coefficient
UNAFold:
0.445
Vsfold5:
0.211
Sensitivity
UNAFold:
0.445
Vsfold5:
0.125
Positive Predictive Value
UNAFold:
0.446
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RSpredict(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
RSpredict(seed):
0.212
Sensitivity
UNAFold:
0.445
RSpredict(seed):
0.075
Positive Predictive Value
UNAFold:
0.446
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CMfinder(20)
Matthews Correlation Coefficient
UNAFold:
0.445
CMfinder(20):
0.210
Sensitivity
UNAFold:
0.445
CMfinder(20):
0.055
Positive Predictive Value
UNAFold:
0.446
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs HotKnots
Matthews Correlation Coefficient
UNAFold:
0.445
HotKnots:
0.179
Sensitivity
UNAFold:
0.445
HotKnots:
0.051
Positive Predictive Value
UNAFold:
0.446
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNASampler(20)
Matthews Correlation Coefficient
UNAFold:
0.445
RNASampler(20):
0.176
Sensitivity
UNAFold:
0.445
RNASampler(20):
0.035
Positive Predictive Value
UNAFold:
0.446
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Cylofold
Matthews Correlation Coefficient
UNAFold:
0.445
Cylofold:
0.155
Sensitivity
UNAFold:
0.445
Cylofold:
0.040
Positive Predictive Value
UNAFold:
0.446
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Pknots
Matthews Correlation Coefficient
UNAFold:
0.445
Pknots:
0.143
Sensitivity
UNAFold:
0.445
Pknots:
0.033
Positive Predictive Value
UNAFold:
0.446
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Mastr(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Mastr(20):
0.134
Sensitivity
UNAFold:
0.445
Mastr(20):
0.023
Positive Predictive Value
UNAFold:
0.446
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Alterna
Matthews Correlation Coefficient
UNAFold:
0.445
Alterna:
0.129
Sensitivity
UNAFold:
0.445
Alterna:
0.030
Positive Predictive Value
UNAFold:
0.446
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs MCFold
Matthews Correlation Coefficient
UNAFold:
0.445
MCFold:
0.125
Sensitivity
UNAFold:
0.445
MCFold:
0.036
Positive Predictive Value
UNAFold:
0.446
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RDfolder
Matthews Correlation Coefficient
UNAFold:
0.445
RDfolder:
0.122
Sensitivity
UNAFold:
0.445
RDfolder:
0.025
Positive Predictive Value
UNAFold:
0.446
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Murlet(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Murlet(seed):
0.079
Sensitivity
UNAFold:
0.445
Murlet(seed):
0.008
Positive Predictive Value
UNAFold:
0.446
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CMfinder(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
CMfinder(seed):
0.080
Sensitivity
UNAFold:
0.445
CMfinder(seed):
0.008
Positive Predictive Value
UNAFold:
0.446
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs TurboFold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
TurboFold(seed):
0.076
Sensitivity
UNAFold:
0.445
TurboFold(seed):
0.007
Positive Predictive Value
UNAFold:
0.446
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PPfold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
PPfold(seed):
0.069
Sensitivity
UNAFold:
0.445
PPfold(seed):
0.006
Positive Predictive Value
UNAFold:
0.446
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Carnac(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Carnac(seed):
0.066
Sensitivity
UNAFold:
0.445
Carnac(seed):
0.005
Positive Predictive Value
UNAFold:
0.446
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNASampler(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
RNASampler(seed):
0.063
Sensitivity
UNAFold:
0.445
RNASampler(seed):
0.005
Positive Predictive Value
UNAFold:
0.446
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs NanoFolder
Matthews Correlation Coefficient
UNAFold:
0.445
NanoFolder:
0.052
Sensitivity
UNAFold:
0.445
NanoFolder:
0.016
Positive Predictive Value
UNAFold:
0.446
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Mastr(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Mastr(seed):
0.051
Sensitivity
UNAFold:
0.445
Mastr(seed):
0.003
Positive Predictive Value
UNAFold:
0.446
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Multilign(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Multilign(seed):
0.043
Sensitivity
UNAFold:
0.445
Multilign(seed):
0.003
Positive Predictive Value
UNAFold:
0.446
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidAlifold(seed) |
1987
ContextFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidAlifold(seed):
0.438
Sensitivity
ContextFold:
0.644
CentroidAlifold(seed):
0.213
Positive Predictive Value
ContextFold:
0.718
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CentroidAlifold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
CentroidAlifold(seed):
0.438
Sensitivity
IPknot:
0.475
CentroidAlifold(seed):
0.213
Positive Predictive Value
IPknot:
0.632
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CentroidAlifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidAlifold(seed):
0.438
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidAlifold(seed):
0.213
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidAlifold(seed):
0.438
Sensitivity
Contrafold:
0.481
CentroidAlifold(seed):
0.213
Positive Predictive Value
Contrafold:
0.537
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
CentroidAlifold(seed):
0.438
Sensitivity
CentroidFold:
0.369
CentroidAlifold(seed):
0.213
Positive Predictive Value
CentroidFold:
0.600
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
CentroidAlifold(seed):
0.438
Sensitivity
Sfold:
0.412
CentroidAlifold(seed):
0.213
Positive Predictive Value
Sfold:
0.535
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs CentroidAlifold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
CentroidAlifold(seed):
0.438
Sensitivity
MaxExpect:
0.443
CentroidAlifold(seed):
0.213
Positive Predictive Value
MaxExpect:
0.488
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs CentroidAlifold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
CentroidAlifold(seed):
0.438
Sensitivity
ProbKnot:
0.446
CentroidAlifold(seed):
0.213
Positive Predictive Value
ProbKnot:
0.475
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
CentroidAlifold(seed):
0.438
Sensitivity
UNAFold:
0.445
CentroidAlifold(seed):
0.213
Positive Predictive Value
UNAFold:
0.446
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
CentroidAlifold(seed) vs Fold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Fold:
0.437
Sensitivity
CentroidAlifold(seed):
0.213
Fold:
0.432
Positive Predictive Value
CentroidAlifold(seed):
0.901
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.88245777838
|
+
CentroidAlifold(seed) vs RNAfold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAfold:
0.429
Sensitivity
CentroidAlifold(seed):
0.213
RNAfold:
0.422
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
CentroidAlifold(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CentroidHomfold‑LAST:
0.428
Sensitivity
CentroidAlifold(seed):
0.213
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 6.53888633552e-08
|
+
CentroidAlifold(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PETfold_pre2.0(20):
0.413
Sensitivity
CentroidAlifold(seed):
0.213
PETfold_pre2.0(20):
0.211
Positive Predictive Value
CentroidAlifold(seed):
0.901
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CentroidAlifold(20):
0.411
Sensitivity
CentroidAlifold(seed):
0.213
CentroidAlifold(20):
0.190
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PknotsRG:
0.386
Sensitivity
CentroidAlifold(seed):
0.213
PknotsRG:
0.320
Positive Predictive Value
CentroidAlifold(seed):
0.901
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
MXScarna(seed):
0.385
Sensitivity
CentroidAlifold(seed):
0.213
MXScarna(seed):
0.196
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAalifold(20):
0.381
Sensitivity
CentroidAlifold(seed):
0.213
RNAalifold(20):
0.185
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAalifold(seed):
0.375
Sensitivity
CentroidAlifold(seed):
0.213
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Afold:
0.358
Sensitivity
CentroidAlifold(seed):
0.213
Afold:
0.295
Positive Predictive Value
CentroidAlifold(seed):
0.901
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
MXScarna(20):
0.359
Sensitivity
CentroidAlifold(seed):
0.213
MXScarna(20):
0.181
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CRWrnafold:
0.346
Sensitivity
CentroidAlifold(seed):
0.213
CRWrnafold:
0.275
Positive Predictive Value
CentroidAlifold(seed):
0.901
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAsubopt:
0.336
Sensitivity
CentroidAlifold(seed):
0.213
RNAsubopt:
0.217
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
McQFold:
0.314
Sensitivity
CentroidAlifold(seed):
0.213
McQFold:
0.205
Positive Predictive Value
CentroidAlifold(seed):
0.901
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
TurboFold(20):
0.301
Sensitivity
CentroidAlifold(seed):
0.213
TurboFold(20):
0.108
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PPfold(20):
0.298
Sensitivity
CentroidAlifold(seed):
0.213
PPfold(20):
0.101
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAshapes:
0.295
Sensitivity
CentroidAlifold(seed):
0.213
RNAshapes:
0.180
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Carnac(20):
0.288
Sensitivity
CentroidAlifold(seed):
0.213
Carnac(20):
0.093
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNASLOpt:
0.281
Sensitivity
CentroidAlifold(seed):
0.213
RNASLOpt:
0.158
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RSpredict(20):
0.270
Sensitivity
CentroidAlifold(seed):
0.213
RSpredict(20):
0.124
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Multilign(20):
0.251
Sensitivity
CentroidAlifold(seed):
0.213
Multilign(20):
0.083
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAwolf:
0.251
Sensitivity
CentroidAlifold(seed):
0.213
RNAwolf:
0.191
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Murlet(20):
0.230
Sensitivity
CentroidAlifold(seed):
0.213
Murlet(20):
0.067
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Vsfold4:
0.217
Sensitivity
CentroidAlifold(seed):
0.213
Vsfold4:
0.125
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Vsfold5:
0.211
Sensitivity
CentroidAlifold(seed):
0.213
Vsfold5:
0.125
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RSpredict(seed):
0.212
Sensitivity
CentroidAlifold(seed):
0.213
RSpredict(seed):
0.075
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CMfinder(20):
0.210
Sensitivity
CentroidAlifold(seed):
0.213
CMfinder(20):
0.055
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
HotKnots:
0.179
Sensitivity
CentroidAlifold(seed):
0.213
HotKnots:
0.051
Positive Predictive Value
CentroidAlifold(seed):
0.901
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNASampler(20):
0.176
Sensitivity
CentroidAlifold(seed):
0.213
RNASampler(20):
0.035
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Cylofold:
0.155
Sensitivity
CentroidAlifold(seed):
0.213
Cylofold:
0.040
Positive Predictive Value
CentroidAlifold(seed):
0.901
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Pknots:
0.143
Sensitivity
CentroidAlifold(seed):
0.213
Pknots:
0.033
Positive Predictive Value
CentroidAlifold(seed):
0.901
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Mastr(20):
0.134
Sensitivity
CentroidAlifold(seed):
0.213
Mastr(20):
0.023
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Alterna:
0.129
Sensitivity
CentroidAlifold(seed):
0.213
Alterna:
0.030
Positive Predictive Value
CentroidAlifold(seed):
0.901
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
MCFold:
0.125
Sensitivity
CentroidAlifold(seed):
0.213
MCFold:
0.036
Positive Predictive Value
CentroidAlifold(seed):
0.901
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RDfolder:
0.122
Sensitivity
CentroidAlifold(seed):
0.213
RDfolder:
0.025
Positive Predictive Value
CentroidAlifold(seed):
0.901
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Murlet(seed):
0.079
Sensitivity
CentroidAlifold(seed):
0.213
Murlet(seed):
0.008
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CMfinder(seed):
0.080
Sensitivity
CentroidAlifold(seed):
0.213
CMfinder(seed):
0.008
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
TurboFold(seed):
0.076
Sensitivity
CentroidAlifold(seed):
0.213
TurboFold(seed):
0.007
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PPfold(seed):
0.069
Sensitivity
CentroidAlifold(seed):
0.213
PPfold(seed):
0.006
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Carnac(seed):
0.066
Sensitivity
CentroidAlifold(seed):
0.213
Carnac(seed):
0.005
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNASampler(seed):
0.063
Sensitivity
CentroidAlifold(seed):
0.213
RNASampler(seed):
0.005
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
NanoFolder:
0.052
Sensitivity
CentroidAlifold(seed):
0.213
NanoFolder:
0.016
Positive Predictive Value
CentroidAlifold(seed):
0.901
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Mastr(seed):
0.051
Sensitivity
CentroidAlifold(seed):
0.213
Mastr(seed):
0.003
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Multilign(seed):
0.043
Sensitivity
CentroidAlifold(seed):
0.213
Multilign(seed):
0.003
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Fold |
1987
ContextFold vs Fold
Matthews Correlation Coefficient
ContextFold:
0.680
Fold:
0.437
Sensitivity
ContextFold:
0.644
Fold:
0.432
Positive Predictive Value
ContextFold:
0.718
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Fold
Matthews Correlation Coefficient
IPknot:
0.548
Fold:
0.437
Sensitivity
IPknot:
0.475
Fold:
0.432
Positive Predictive Value
IPknot:
0.632
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Fold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Fold:
0.437
Sensitivity
PETfold_pre2.0(seed):
0.343
Fold:
0.432
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Fold
Matthews Correlation Coefficient
Contrafold:
0.508
Fold:
0.437
Sensitivity
Contrafold:
0.481
Fold:
0.432
Positive Predictive Value
Contrafold:
0.537
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Fold
Matthews Correlation Coefficient
CentroidFold:
0.470
Fold:
0.437
Sensitivity
CentroidFold:
0.369
Fold:
0.432
Positive Predictive Value
CentroidFold:
0.600
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Fold
Matthews Correlation Coefficient
Sfold:
0.469
Fold:
0.437
Sensitivity
Sfold:
0.412
Fold:
0.432
Positive Predictive Value
Sfold:
0.535
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Fold
Matthews Correlation Coefficient
MaxExpect:
0.465
Fold:
0.437
Sensitivity
MaxExpect:
0.443
Fold:
0.432
Positive Predictive Value
MaxExpect:
0.488
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Fold
Matthews Correlation Coefficient
ProbKnot:
0.460
Fold:
0.437
Sensitivity
ProbKnot:
0.446
Fold:
0.432
Positive Predictive Value
ProbKnot:
0.475
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Fold
Matthews Correlation Coefficient
UNAFold:
0.445
Fold:
0.437
Sensitivity
UNAFold:
0.445
Fold:
0.432
Positive Predictive Value
UNAFold:
0.446
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Fold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Fold:
0.437
Sensitivity
CentroidAlifold(seed):
0.213
Fold:
0.432
Positive Predictive Value
CentroidAlifold(seed):
0.901
Fold:
0.444
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.88245777838
|
|
+
Fold vs RNAfold
Matthews Correlation Coefficient
Fold:
0.437
RNAfold:
0.429
Sensitivity
Fold:
0.432
RNAfold:
0.422
Positive Predictive Value
Fold:
0.444
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
Fold:
0.437
CentroidHomfold‑LAST:
0.428
Sensitivity
Fold:
0.432
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
Fold:
0.444
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
Fold:
0.437
PETfold_pre2.0(20):
0.413
Sensitivity
Fold:
0.432
PETfold_pre2.0(20):
0.211
Positive Predictive Value
Fold:
0.444
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CentroidAlifold(20)
Matthews Correlation Coefficient
Fold:
0.437
CentroidAlifold(20):
0.411
Sensitivity
Fold:
0.432
CentroidAlifold(20):
0.190
Positive Predictive Value
Fold:
0.444
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PknotsRG
Matthews Correlation Coefficient
Fold:
0.437
PknotsRG:
0.386
Sensitivity
Fold:
0.432
PknotsRG:
0.320
Positive Predictive Value
Fold:
0.444
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs MXScarna(seed)
Matthews Correlation Coefficient
Fold:
0.437
MXScarna(seed):
0.385
Sensitivity
Fold:
0.432
MXScarna(seed):
0.196
Positive Predictive Value
Fold:
0.444
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAalifold(20)
Matthews Correlation Coefficient
Fold:
0.437
RNAalifold(20):
0.381
Sensitivity
Fold:
0.432
RNAalifold(20):
0.185
Positive Predictive Value
Fold:
0.444
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAalifold(seed)
Matthews Correlation Coefficient
Fold:
0.437
RNAalifold(seed):
0.375
Sensitivity
Fold:
0.432
RNAalifold(seed):
0.169
Positive Predictive Value
Fold:
0.444
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Afold
Matthews Correlation Coefficient
Fold:
0.437
Afold:
0.358
Sensitivity
Fold:
0.432
Afold:
0.295
Positive Predictive Value
Fold:
0.444
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs MXScarna(20)
Matthews Correlation Coefficient
Fold:
0.437
MXScarna(20):
0.359
Sensitivity
Fold:
0.432
MXScarna(20):
0.181
Positive Predictive Value
Fold:
0.444
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CRWrnafold
Matthews Correlation Coefficient
Fold:
0.437
CRWrnafold:
0.346
Sensitivity
Fold:
0.432
CRWrnafold:
0.275
Positive Predictive Value
Fold:
0.444
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAsubopt
Matthews Correlation Coefficient
Fold:
0.437
RNAsubopt:
0.336
Sensitivity
Fold:
0.432
RNAsubopt:
0.217
Positive Predictive Value
Fold:
0.444
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs McQFold
Matthews Correlation Coefficient
Fold:
0.437
McQFold:
0.314
Sensitivity
Fold:
0.432
McQFold:
0.205
Positive Predictive Value
Fold:
0.444
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs TurboFold(20)
Matthews Correlation Coefficient
Fold:
0.437
TurboFold(20):
0.301
Sensitivity
Fold:
0.432
TurboFold(20):
0.108
Positive Predictive Value
Fold:
0.444
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PPfold(20)
Matthews Correlation Coefficient
Fold:
0.437
PPfold(20):
0.298
Sensitivity
Fold:
0.432
PPfold(20):
0.101
Positive Predictive Value
Fold:
0.444
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAshapes
Matthews Correlation Coefficient
Fold:
0.437
RNAshapes:
0.295
Sensitivity
Fold:
0.432
RNAshapes:
0.180
Positive Predictive Value
Fold:
0.444
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Carnac(20)
Matthews Correlation Coefficient
Fold:
0.437
Carnac(20):
0.288
Sensitivity
Fold:
0.432
Carnac(20):
0.093
Positive Predictive Value
Fold:
0.444
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNASLOpt
Matthews Correlation Coefficient
Fold:
0.437
RNASLOpt:
0.281
Sensitivity
Fold:
0.432
RNASLOpt:
0.158
Positive Predictive Value
Fold:
0.444
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RSpredict(20)
Matthews Correlation Coefficient
Fold:
0.437
RSpredict(20):
0.270
Sensitivity
Fold:
0.432
RSpredict(20):
0.124
Positive Predictive Value
Fold:
0.444
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Multilign(20)
Matthews Correlation Coefficient
Fold:
0.437
Multilign(20):
0.251
Sensitivity
Fold:
0.432
Multilign(20):
0.083
Positive Predictive Value
Fold:
0.444
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAwolf
Matthews Correlation Coefficient
Fold:
0.437
RNAwolf:
0.251
Sensitivity
Fold:
0.432
RNAwolf:
0.191
Positive Predictive Value
Fold:
0.444
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Murlet(20)
Matthews Correlation Coefficient
Fold:
0.437
Murlet(20):
0.230
Sensitivity
Fold:
0.432
Murlet(20):
0.067
Positive Predictive Value
Fold:
0.444
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Vsfold4
Matthews Correlation Coefficient
Fold:
0.437
Vsfold4:
0.217
Sensitivity
Fold:
0.432
Vsfold4:
0.125
Positive Predictive Value
Fold:
0.444
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Vsfold5
Matthews Correlation Coefficient
Fold:
0.437
Vsfold5:
0.211
Sensitivity
Fold:
0.432
Vsfold5:
0.125
Positive Predictive Value
Fold:
0.444
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RSpredict(seed)
Matthews Correlation Coefficient
Fold:
0.437
RSpredict(seed):
0.212
Sensitivity
Fold:
0.432
RSpredict(seed):
0.075
Positive Predictive Value
Fold:
0.444
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CMfinder(20)
Matthews Correlation Coefficient
Fold:
0.437
CMfinder(20):
0.210
Sensitivity
Fold:
0.432
CMfinder(20):
0.055
Positive Predictive Value
Fold:
0.444
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs HotKnots
Matthews Correlation Coefficient
Fold:
0.437
HotKnots:
0.179
Sensitivity
Fold:
0.432
HotKnots:
0.051
Positive Predictive Value
Fold:
0.444
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNASampler(20)
Matthews Correlation Coefficient
Fold:
0.437
RNASampler(20):
0.176
Sensitivity
Fold:
0.432
RNASampler(20):
0.035
Positive Predictive Value
Fold:
0.444
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Cylofold
Matthews Correlation Coefficient
Fold:
0.437
Cylofold:
0.155
Sensitivity
Fold:
0.432
Cylofold:
0.040
Positive Predictive Value
Fold:
0.444
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Pknots
Matthews Correlation Coefficient
Fold:
0.437
Pknots:
0.143
Sensitivity
Fold:
0.432
Pknots:
0.033
Positive Predictive Value
Fold:
0.444
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Mastr(20)
Matthews Correlation Coefficient
Fold:
0.437
Mastr(20):
0.134
Sensitivity
Fold:
0.432
Mastr(20):
0.023
Positive Predictive Value
Fold:
0.444
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Alterna
Matthews Correlation Coefficient
Fold:
0.437
Alterna:
0.129
Sensitivity
Fold:
0.432
Alterna:
0.030
Positive Predictive Value
Fold:
0.444
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs MCFold
Matthews Correlation Coefficient
Fold:
0.437
MCFold:
0.125
Sensitivity
Fold:
0.432
MCFold:
0.036
Positive Predictive Value
Fold:
0.444
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RDfolder
Matthews Correlation Coefficient
Fold:
0.437
RDfolder:
0.122
Sensitivity
Fold:
0.432
RDfolder:
0.025
Positive Predictive Value
Fold:
0.444
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Murlet(seed)
Matthews Correlation Coefficient
Fold:
0.437
Murlet(seed):
0.079
Sensitivity
Fold:
0.432
Murlet(seed):
0.008
Positive Predictive Value
Fold:
0.444
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CMfinder(seed)
Matthews Correlation Coefficient
Fold:
0.437
CMfinder(seed):
0.080
Sensitivity
Fold:
0.432
CMfinder(seed):
0.008
Positive Predictive Value
Fold:
0.444
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs TurboFold(seed)
Matthews Correlation Coefficient
Fold:
0.437
TurboFold(seed):
0.076
Sensitivity
Fold:
0.432
TurboFold(seed):
0.007
Positive Predictive Value
Fold:
0.444
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PPfold(seed)
Matthews Correlation Coefficient
Fold:
0.437
PPfold(seed):
0.069
Sensitivity
Fold:
0.432
PPfold(seed):
0.006
Positive Predictive Value
Fold:
0.444
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Carnac(seed)
Matthews Correlation Coefficient
Fold:
0.437
Carnac(seed):
0.066
Sensitivity
Fold:
0.432
Carnac(seed):
0.005
Positive Predictive Value
Fold:
0.444
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNASampler(seed)
Matthews Correlation Coefficient
Fold:
0.437
RNASampler(seed):
0.063
Sensitivity
Fold:
0.432
RNASampler(seed):
0.005
Positive Predictive Value
Fold:
0.444
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs NanoFolder
Matthews Correlation Coefficient
Fold:
0.437
NanoFolder:
0.052
Sensitivity
Fold:
0.432
NanoFolder:
0.016
Positive Predictive Value
Fold:
0.444
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Mastr(seed)
Matthews Correlation Coefficient
Fold:
0.437
Mastr(seed):
0.051
Sensitivity
Fold:
0.432
Mastr(seed):
0.003
Positive Predictive Value
Fold:
0.444
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Multilign(seed)
Matthews Correlation Coefficient
Fold:
0.437
Multilign(seed):
0.043
Sensitivity
Fold:
0.432
Multilign(seed):
0.003
Positive Predictive Value
Fold:
0.444
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAfold |
1987
ContextFold vs RNAfold
Matthews Correlation Coefficient
ContextFold:
0.680
RNAfold:
0.429
Sensitivity
ContextFold:
0.644
RNAfold:
0.422
Positive Predictive Value
ContextFold:
0.718
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNAfold
Matthews Correlation Coefficient
IPknot:
0.548
RNAfold:
0.429
Sensitivity
IPknot:
0.475
RNAfold:
0.422
Positive Predictive Value
IPknot:
0.632
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNAfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAfold:
0.429
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAfold:
0.422
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNAfold
Matthews Correlation Coefficient
Contrafold:
0.508
RNAfold:
0.429
Sensitivity
Contrafold:
0.481
RNAfold:
0.422
Positive Predictive Value
Contrafold:
0.537
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNAfold
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAfold:
0.429
Sensitivity
CentroidFold:
0.369
RNAfold:
0.422
Positive Predictive Value
CentroidFold:
0.600
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNAfold
Matthews Correlation Coefficient
Sfold:
0.469
RNAfold:
0.429
Sensitivity
Sfold:
0.412
RNAfold:
0.422
Positive Predictive Value
Sfold:
0.535
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNAfold
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAfold:
0.429
Sensitivity
MaxExpect:
0.443
RNAfold:
0.422
Positive Predictive Value
MaxExpect:
0.488
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNAfold
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAfold:
0.429
Sensitivity
ProbKnot:
0.446
RNAfold:
0.422
Positive Predictive Value
ProbKnot:
0.475
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNAfold
Matthews Correlation Coefficient
UNAFold:
0.445
RNAfold:
0.429
Sensitivity
UNAFold:
0.445
RNAfold:
0.422
Positive Predictive Value
UNAFold:
0.446
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNAfold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAfold:
0.429
Sensitivity
CentroidAlifold(seed):
0.213
RNAfold:
0.422
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
1987
Fold vs RNAfold
Matthews Correlation Coefficient
Fold:
0.437
RNAfold:
0.429
Sensitivity
Fold:
0.432
RNAfold:
0.422
Positive Predictive Value
Fold:
0.444
RNAfold:
0.437
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAfold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
RNAfold:
0.429
CentroidHomfold‑LAST:
0.428
Sensitivity
RNAfold:
0.422
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
RNAfold:
0.437
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.000639955457467
|
+
RNAfold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
RNAfold:
0.429
PETfold_pre2.0(20):
0.413
Sensitivity
RNAfold:
0.422
PETfold_pre2.0(20):
0.211
Positive Predictive Value
RNAfold:
0.437
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CentroidAlifold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
CentroidAlifold(20):
0.411
Sensitivity
RNAfold:
0.422
CentroidAlifold(20):
0.190
Positive Predictive Value
RNAfold:
0.437
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs PknotsRG
Matthews Correlation Coefficient
RNAfold:
0.429
PknotsRG:
0.386
Sensitivity
RNAfold:
0.422
PknotsRG:
0.320
Positive Predictive Value
RNAfold:
0.437
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs MXScarna(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
MXScarna(seed):
0.385
Sensitivity
RNAfold:
0.422
MXScarna(seed):
0.196
Positive Predictive Value
RNAfold:
0.437
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAalifold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
RNAalifold(20):
0.381
Sensitivity
RNAfold:
0.422
RNAalifold(20):
0.185
Positive Predictive Value
RNAfold:
0.437
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAalifold(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
RNAalifold(seed):
0.375
Sensitivity
RNAfold:
0.422
RNAalifold(seed):
0.169
Positive Predictive Value
RNAfold:
0.437
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Afold
Matthews Correlation Coefficient
RNAfold:
0.429
Afold:
0.358
Sensitivity
RNAfold:
0.422
Afold:
0.295
Positive Predictive Value
RNAfold:
0.437
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs MXScarna(20)
Matthews Correlation Coefficient
RNAfold:
0.429
MXScarna(20):
0.359
Sensitivity
RNAfold:
0.422
MXScarna(20):
0.181
Positive Predictive Value
RNAfold:
0.437
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CRWrnafold
Matthews Correlation Coefficient
RNAfold:
0.429
CRWrnafold:
0.346
Sensitivity
RNAfold:
0.422
CRWrnafold:
0.275
Positive Predictive Value
RNAfold:
0.437
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAsubopt
Matthews Correlation Coefficient
RNAfold:
0.429
RNAsubopt:
0.336
Sensitivity
RNAfold:
0.422
RNAsubopt:
0.217
Positive Predictive Value
RNAfold:
0.437
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs McQFold
Matthews Correlation Coefficient
RNAfold:
0.429
McQFold:
0.314
Sensitivity
RNAfold:
0.422
McQFold:
0.205
Positive Predictive Value
RNAfold:
0.437
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs TurboFold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
TurboFold(20):
0.301
Sensitivity
RNAfold:
0.422
TurboFold(20):
0.108
Positive Predictive Value
RNAfold:
0.437
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs PPfold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
PPfold(20):
0.298
Sensitivity
RNAfold:
0.422
PPfold(20):
0.101
Positive Predictive Value
RNAfold:
0.437
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAshapes
Matthews Correlation Coefficient
RNAfold:
0.429
RNAshapes:
0.295
Sensitivity
RNAfold:
0.422
RNAshapes:
0.180
Positive Predictive Value
RNAfold:
0.437
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Carnac(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Carnac(20):
0.288
Sensitivity
RNAfold:
0.422
Carnac(20):
0.093
Positive Predictive Value
RNAfold:
0.437
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNASLOpt
Matthews Correlation Coefficient
RNAfold:
0.429
RNASLOpt:
0.281
Sensitivity
RNAfold:
0.422
RNASLOpt:
0.158
Positive Predictive Value
RNAfold:
0.437
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RSpredict(20)
Matthews Correlation Coefficient
RNAfold:
0.429
RSpredict(20):
0.270
Sensitivity
RNAfold:
0.422
RSpredict(20):
0.124
Positive Predictive Value
RNAfold:
0.437
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Multilign(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Multilign(20):
0.251
Sensitivity
RNAfold:
0.422
Multilign(20):
0.083
Positive Predictive Value
RNAfold:
0.437
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAwolf
Matthews Correlation Coefficient
RNAfold:
0.429
RNAwolf:
0.251
Sensitivity
RNAfold:
0.422
RNAwolf:
0.191
Positive Predictive Value
RNAfold:
0.437
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Murlet(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Murlet(20):
0.230
Sensitivity
RNAfold:
0.422
Murlet(20):
0.067
Positive Predictive Value
RNAfold:
0.437
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Vsfold4
Matthews Correlation Coefficient
RNAfold:
0.429
Vsfold4:
0.217
Sensitivity
RNAfold:
0.422
Vsfold4:
0.125
Positive Predictive Value
RNAfold:
0.437
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Vsfold5
Matthews Correlation Coefficient
RNAfold:
0.429
Vsfold5:
0.211
Sensitivity
RNAfold:
0.422
Vsfold5:
0.125
Positive Predictive Value
RNAfold:
0.437
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RSpredict(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
RSpredict(seed):
0.212
Sensitivity
RNAfold:
0.422
RSpredict(seed):
0.075
Positive Predictive Value
RNAfold:
0.437
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CMfinder(20)
Matthews Correlation Coefficient
RNAfold:
0.429
CMfinder(20):
0.210
Sensitivity
RNAfold:
0.422
CMfinder(20):
0.055
Positive Predictive Value
RNAfold:
0.437
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs HotKnots
Matthews Correlation Coefficient
RNAfold:
0.429
HotKnots:
0.179
Sensitivity
RNAfold:
0.422
HotKnots:
0.051
Positive Predictive Value
RNAfold:
0.437
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNASampler(20)
Matthews Correlation Coefficient
RNAfold:
0.429
RNASampler(20):
0.176
Sensitivity
RNAfold:
0.422
RNASampler(20):
0.035
Positive Predictive Value
RNAfold:
0.437
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Cylofold
Matthews Correlation Coefficient
RNAfold:
0.429
Cylofold:
0.155
Sensitivity
RNAfold:
0.422
Cylofold:
0.040
Positive Predictive Value
RNAfold:
0.437
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Pknots
Matthews Correlation Coefficient
RNAfold:
0.429
Pknots:
0.143
Sensitivity
RNAfold:
0.422
Pknots:
0.033
Positive Predictive Value
RNAfold:
0.437
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Mastr(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Mastr(20):
0.134
Sensitivity
RNAfold:
0.422
Mastr(20):
0.023
Positive Predictive Value
RNAfold:
0.437
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Alterna
Matthews Correlation Coefficient
RNAfold:
0.429
Alterna:
0.129
Sensitivity
RNAfold:
0.422
Alterna:
0.030
Positive Predictive Value
RNAfold:
0.437
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs MCFold
Matthews Correlation Coefficient
RNAfold:
0.429
MCFold:
0.125
Sensitivity
RNAfold:
0.422
MCFold:
0.036
Positive Predictive Value
RNAfold:
0.437
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RDfolder
Matthews Correlation Coefficient
RNAfold:
0.429
RDfolder:
0.122
Sensitivity
RNAfold:
0.422
RDfolder:
0.025
Positive Predictive Value
RNAfold:
0.437
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Murlet(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Murlet(seed):
0.079
Sensitivity
RNAfold:
0.422
Murlet(seed):
0.008
Positive Predictive Value
RNAfold:
0.437
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CMfinder(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
CMfinder(seed):
0.080
Sensitivity
RNAfold:
0.422
CMfinder(seed):
0.008
Positive Predictive Value
RNAfold:
0.437
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs TurboFold(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
TurboFold(seed):
0.076
Sensitivity
RNAfold:
0.422
TurboFold(seed):
0.007
Positive Predictive Value
RNAfold:
0.437
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs PPfold(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
PPfold(seed):
0.069
Sensitivity
RNAfold:
0.422
PPfold(seed):
0.006
Positive Predictive Value
RNAfold:
0.437
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Carnac(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Carnac(seed):
0.066
Sensitivity
RNAfold:
0.422
Carnac(seed):
0.005
Positive Predictive Value
RNAfold:
0.437
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNASampler(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
RNASampler(seed):
0.063
Sensitivity
RNAfold:
0.422
RNASampler(seed):
0.005
Positive Predictive Value
RNAfold:
0.437
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs NanoFolder
Matthews Correlation Coefficient
RNAfold:
0.429
NanoFolder:
0.052
Sensitivity
RNAfold:
0.422
NanoFolder:
0.016
Positive Predictive Value
RNAfold:
0.437
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Mastr(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Mastr(seed):
0.051
Sensitivity
RNAfold:
0.422
Mastr(seed):
0.003
Positive Predictive Value
RNAfold:
0.437
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Multilign(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Multilign(seed):
0.043
Sensitivity
RNAfold:
0.422
Multilign(seed):
0.003
Positive Predictive Value
RNAfold:
0.437
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidHomfold‑LAST |
1987
ContextFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidHomfold‑LAST:
0.428
Sensitivity
ContextFold:
0.644
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
ContextFold:
0.718
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
IPknot:
0.548
CentroidHomfold‑LAST:
0.428
Sensitivity
IPknot:
0.475
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
IPknot:
0.632
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidHomfold‑LAST:
0.428
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidHomfold‑LAST:
0.428
Sensitivity
Contrafold:
0.481
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
Contrafold:
0.537
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidFold:
0.470
CentroidHomfold‑LAST:
0.428
Sensitivity
CentroidFold:
0.369
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
CentroidFold:
0.600
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
Sfold:
0.469
CentroidHomfold‑LAST:
0.428
Sensitivity
Sfold:
0.412
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
Sfold:
0.535
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
MaxExpect:
0.465
CentroidHomfold‑LAST:
0.428
Sensitivity
MaxExpect:
0.443
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
MaxExpect:
0.488
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
ProbKnot:
0.460
CentroidHomfold‑LAST:
0.428
Sensitivity
ProbKnot:
0.446
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
ProbKnot:
0.475
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
UNAFold:
0.445
CentroidHomfold‑LAST:
0.428
Sensitivity
UNAFold:
0.445
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
UNAFold:
0.446
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CentroidHomfold‑LAST:
0.428
Sensitivity
CentroidAlifold(seed):
0.213
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 6.53888633552e-08
|
1987
Fold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
Fold:
0.437
CentroidHomfold‑LAST:
0.428
Sensitivity
Fold:
0.432
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
Fold:
0.444
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
RNAfold:
0.429
CentroidHomfold‑LAST:
0.428
Sensitivity
RNAfold:
0.422
CentroidHomfold‑LAST:
0.216
Positive Predictive Value
RNAfold:
0.437
CentroidHomfold‑LAST:
0.850
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.000639955457467
|
|
+
CentroidHomfold‑LAST vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PETfold_pre2.0(20):
0.413
Sensitivity
CentroidHomfold‑LAST:
0.216
PETfold_pre2.0(20):
0.211
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CentroidAlifold(20):
0.411
Sensitivity
CentroidHomfold‑LAST:
0.216
CentroidAlifold(20):
0.190
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs PknotsRG
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PknotsRG:
0.386
Sensitivity
CentroidHomfold‑LAST:
0.216
PknotsRG:
0.320
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
MXScarna(seed):
0.385
Sensitivity
CentroidHomfold‑LAST:
0.216
MXScarna(seed):
0.196
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAalifold(20):
0.381
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAalifold(20):
0.185
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAalifold(seed):
0.375
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Afold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Afold:
0.358
Sensitivity
CentroidHomfold‑LAST:
0.216
Afold:
0.295
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs MXScarna(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
MXScarna(20):
0.359
Sensitivity
CentroidHomfold‑LAST:
0.216
MXScarna(20):
0.181
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CRWrnafold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CRWrnafold:
0.346
Sensitivity
CentroidHomfold‑LAST:
0.216
CRWrnafold:
0.275
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAsubopt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAsubopt:
0.336
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAsubopt:
0.217
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs McQFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
McQFold:
0.314
Sensitivity
CentroidHomfold‑LAST:
0.216
McQFold:
0.205
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs TurboFold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
TurboFold(20):
0.301
Sensitivity
CentroidHomfold‑LAST:
0.216
TurboFold(20):
0.108
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs PPfold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PPfold(20):
0.298
Sensitivity
CentroidHomfold‑LAST:
0.216
PPfold(20):
0.101
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAshapes
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAshapes:
0.295
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAshapes:
0.180
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Carnac(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Carnac(20):
0.288
Sensitivity
CentroidHomfold‑LAST:
0.216
Carnac(20):
0.093
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNASLOpt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNASLOpt:
0.281
Sensitivity
CentroidHomfold‑LAST:
0.216
RNASLOpt:
0.158
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RSpredict(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RSpredict(20):
0.270
Sensitivity
CentroidHomfold‑LAST:
0.216
RSpredict(20):
0.124
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Multilign(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Multilign(20):
0.251
Sensitivity
CentroidHomfold‑LAST:
0.216
Multilign(20):
0.083
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAwolf
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAwolf:
0.251
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAwolf:
0.191
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Murlet(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Murlet(20):
0.230
Sensitivity
CentroidHomfold‑LAST:
0.216
Murlet(20):
0.067
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Vsfold4
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Vsfold4:
0.217
Sensitivity
CentroidHomfold‑LAST:
0.216
Vsfold4:
0.125
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Vsfold5
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Vsfold5:
0.211
Sensitivity
CentroidHomfold‑LAST:
0.216
Vsfold5:
0.125
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RSpredict(seed):
0.212
Sensitivity
CentroidHomfold‑LAST:
0.216
RSpredict(seed):
0.075
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CMfinder(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CMfinder(20):
0.210
Sensitivity
CentroidHomfold‑LAST:
0.216
CMfinder(20):
0.055
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs HotKnots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
HotKnots:
0.179
Sensitivity
CentroidHomfold‑LAST:
0.216
HotKnots:
0.051
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNASampler(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNASampler(20):
0.176
Sensitivity
CentroidHomfold‑LAST:
0.216
RNASampler(20):
0.035
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Cylofold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Cylofold:
0.155
Sensitivity
CentroidHomfold‑LAST:
0.216
Cylofold:
0.040
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Pknots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Pknots:
0.143
Sensitivity
CentroidHomfold‑LAST:
0.216
Pknots:
0.033
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Mastr(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Mastr(20):
0.134
Sensitivity
CentroidHomfold‑LAST:
0.216
Mastr(20):
0.023
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Alterna
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Alterna:
0.129
Sensitivity
CentroidHomfold‑LAST:
0.216
Alterna:
0.030
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs MCFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
MCFold:
0.125
Sensitivity
CentroidHomfold‑LAST:
0.216
MCFold:
0.036
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RDfolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RDfolder:
0.122
Sensitivity
CentroidHomfold‑LAST:
0.216
RDfolder:
0.025
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Murlet(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Murlet(seed):
0.079
Sensitivity
CentroidHomfold‑LAST:
0.216
Murlet(seed):
0.008
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CMfinder(seed):
0.080
Sensitivity
CentroidHomfold‑LAST:
0.216
CMfinder(seed):
0.008
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
TurboFold(seed):
0.076
Sensitivity
CentroidHomfold‑LAST:
0.216
TurboFold(seed):
0.007
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs PPfold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PPfold(seed):
0.069
Sensitivity
CentroidHomfold‑LAST:
0.216
PPfold(seed):
0.006
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Carnac(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Carnac(seed):
0.066
Sensitivity
CentroidHomfold‑LAST:
0.216
Carnac(seed):
0.005
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNASampler(seed):
0.063
Sensitivity
CentroidHomfold‑LAST:
0.216
RNASampler(seed):
0.005
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs NanoFolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
NanoFolder:
0.052
Sensitivity
CentroidHomfold‑LAST:
0.216
NanoFolder:
0.016
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Mastr(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Mastr(seed):
0.051
Sensitivity
CentroidHomfold‑LAST:
0.216
Mastr(seed):
0.003
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Multilign(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Multilign(seed):
0.043
Sensitivity
CentroidHomfold‑LAST:
0.216
Multilign(seed):
0.003
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PETfold_pre2.0(20) |
1987
ContextFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
ContextFold:
0.680
PETfold_pre2.0(20):
0.413
Sensitivity
ContextFold:
0.644
PETfold_pre2.0(20):
0.211
Positive Predictive Value
ContextFold:
0.718
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
IPknot:
0.548
PETfold_pre2.0(20):
0.413
Sensitivity
IPknot:
0.475
PETfold_pre2.0(20):
0.211
Positive Predictive Value
IPknot:
0.632
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PETfold_pre2.0(20):
0.413
Sensitivity
PETfold_pre2.0(seed):
0.343
PETfold_pre2.0(20):
0.211
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
Contrafold:
0.508
PETfold_pre2.0(20):
0.413
Sensitivity
Contrafold:
0.481
PETfold_pre2.0(20):
0.211
Positive Predictive Value
Contrafold:
0.537
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
PETfold_pre2.0(20):
0.413
Sensitivity
CentroidFold:
0.369
PETfold_pre2.0(20):
0.211
Positive Predictive Value
CentroidFold:
0.600
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
Sfold:
0.469
PETfold_pre2.0(20):
0.413
Sensitivity
Sfold:
0.412
PETfold_pre2.0(20):
0.211
Positive Predictive Value
Sfold:
0.535
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
PETfold_pre2.0(20):
0.413
Sensitivity
MaxExpect:
0.443
PETfold_pre2.0(20):
0.211
Positive Predictive Value
MaxExpect:
0.488
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
PETfold_pre2.0(20):
0.413
Sensitivity
ProbKnot:
0.446
PETfold_pre2.0(20):
0.211
Positive Predictive Value
ProbKnot:
0.475
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
UNAFold:
0.445
PETfold_pre2.0(20):
0.413
Sensitivity
UNAFold:
0.445
PETfold_pre2.0(20):
0.211
Positive Predictive Value
UNAFold:
0.446
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PETfold_pre2.0(20):
0.413
Sensitivity
CentroidAlifold(seed):
0.213
PETfold_pre2.0(20):
0.211
Positive Predictive Value
CentroidAlifold(seed):
0.901
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
Fold:
0.437
PETfold_pre2.0(20):
0.413
Sensitivity
Fold:
0.432
PETfold_pre2.0(20):
0.211
Positive Predictive Value
Fold:
0.444
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
RNAfold:
0.429
PETfold_pre2.0(20):
0.413
Sensitivity
RNAfold:
0.422
PETfold_pre2.0(20):
0.211
Positive Predictive Value
RNAfold:
0.437
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PETfold_pre2.0(20):
0.413
Sensitivity
CentroidHomfold‑LAST:
0.216
PETfold_pre2.0(20):
0.211
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PETfold_pre2.0(20) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CentroidAlifold(20):
0.411
Sensitivity
PETfold_pre2.0(20):
0.211
CentroidAlifold(20):
0.190
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
PknotsRG:
0.386
Sensitivity
PETfold_pre2.0(20):
0.211
PknotsRG:
0.320
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
MXScarna(seed):
0.385
Sensitivity
PETfold_pre2.0(20):
0.211
MXScarna(seed):
0.196
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAalifold(20):
0.381
Sensitivity
PETfold_pre2.0(20):
0.211
RNAalifold(20):
0.185
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAalifold(seed):
0.375
Sensitivity
PETfold_pre2.0(20):
0.211
RNAalifold(seed):
0.169
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Afold:
0.358
Sensitivity
PETfold_pre2.0(20):
0.211
Afold:
0.295
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
MXScarna(20):
0.359
Sensitivity
PETfold_pre2.0(20):
0.211
MXScarna(20):
0.181
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CRWrnafold:
0.346
Sensitivity
PETfold_pre2.0(20):
0.211
CRWrnafold:
0.275
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAsubopt:
0.336
Sensitivity
PETfold_pre2.0(20):
0.211
RNAsubopt:
0.217
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
McQFold:
0.314
Sensitivity
PETfold_pre2.0(20):
0.211
McQFold:
0.205
Positive Predictive Value
PETfold_pre2.0(20):
0.807
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
TurboFold(20):
0.301
Sensitivity
PETfold_pre2.0(20):
0.211
TurboFold(20):
0.108
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
PPfold(20):
0.298
Sensitivity
PETfold_pre2.0(20):
0.211
PPfold(20):
0.101
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAshapes:
0.295
Sensitivity
PETfold_pre2.0(20):
0.211
RNAshapes:
0.180
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Carnac(20):
0.288
Sensitivity
PETfold_pre2.0(20):
0.211
Carnac(20):
0.093
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNASLOpt:
0.281
Sensitivity
PETfold_pre2.0(20):
0.211
RNASLOpt:
0.158
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RSpredict(20):
0.270
Sensitivity
PETfold_pre2.0(20):
0.211
RSpredict(20):
0.124
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Multilign(20):
0.251
Sensitivity
PETfold_pre2.0(20):
0.211
Multilign(20):
0.083
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAwolf:
0.251
Sensitivity
PETfold_pre2.0(20):
0.211
RNAwolf:
0.191
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Murlet(20):
0.230
Sensitivity
PETfold_pre2.0(20):
0.211
Murlet(20):
0.067
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Vsfold4:
0.217
Sensitivity
PETfold_pre2.0(20):
0.211
Vsfold4:
0.125
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Vsfold5:
0.211
Sensitivity
PETfold_pre2.0(20):
0.211
Vsfold5:
0.125
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RSpredict(seed):
0.212
Sensitivity
PETfold_pre2.0(20):
0.211
RSpredict(seed):
0.075
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CMfinder(20):
0.210
Sensitivity
PETfold_pre2.0(20):
0.211
CMfinder(20):
0.055
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
HotKnots:
0.179
Sensitivity
PETfold_pre2.0(20):
0.211
HotKnots:
0.051
Positive Predictive Value
PETfold_pre2.0(20):
0.807
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNASampler(20):
0.176
Sensitivity
PETfold_pre2.0(20):
0.211
RNASampler(20):
0.035
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Cylofold:
0.155
Sensitivity
PETfold_pre2.0(20):
0.211
Cylofold:
0.040
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Pknots:
0.143
Sensitivity
PETfold_pre2.0(20):
0.211
Pknots:
0.033
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Mastr(20):
0.134
Sensitivity
PETfold_pre2.0(20):
0.211
Mastr(20):
0.023
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Alterna:
0.129
Sensitivity
PETfold_pre2.0(20):
0.211
Alterna:
0.030
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
MCFold:
0.125
Sensitivity
PETfold_pre2.0(20):
0.211
MCFold:
0.036
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RDfolder:
0.122
Sensitivity
PETfold_pre2.0(20):
0.211
RDfolder:
0.025
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Murlet(seed):
0.079
Sensitivity
PETfold_pre2.0(20):
0.211
Murlet(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CMfinder(seed):
0.080
Sensitivity
PETfold_pre2.0(20):
0.211
CMfinder(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
TurboFold(seed):
0.076
Sensitivity
PETfold_pre2.0(20):
0.211
TurboFold(seed):
0.007
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
PPfold(seed):
0.069
Sensitivity
PETfold_pre2.0(20):
0.211
PPfold(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Carnac(seed):
0.066
Sensitivity
PETfold_pre2.0(20):
0.211
Carnac(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNASampler(seed):
0.063
Sensitivity
PETfold_pre2.0(20):
0.211
RNASampler(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
NanoFolder:
0.052
Sensitivity
PETfold_pre2.0(20):
0.211
NanoFolder:
0.016
Positive Predictive Value
PETfold_pre2.0(20):
0.807
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Mastr(seed):
0.051
Sensitivity
PETfold_pre2.0(20):
0.211
Mastr(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Multilign(seed):
0.043
Sensitivity
PETfold_pre2.0(20):
0.211
Multilign(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidAlifold(20) |
1987
ContextFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
CentroidAlifold(20):
0.411
Sensitivity
ContextFold:
0.644
CentroidAlifold(20):
0.190
Positive Predictive Value
ContextFold:
0.718
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CentroidAlifold(20)
Matthews Correlation Coefficient
IPknot:
0.548
CentroidAlifold(20):
0.411
Sensitivity
IPknot:
0.475
CentroidAlifold(20):
0.190
Positive Predictive Value
IPknot:
0.632
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CentroidAlifold(20):
0.411
Sensitivity
PETfold_pre2.0(seed):
0.343
CentroidAlifold(20):
0.190
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CentroidAlifold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
CentroidAlifold(20):
0.411
Sensitivity
Contrafold:
0.481
CentroidAlifold(20):
0.190
Positive Predictive Value
Contrafold:
0.537
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
CentroidAlifold(20):
0.411
Sensitivity
CentroidFold:
0.369
CentroidAlifold(20):
0.190
Positive Predictive Value
CentroidFold:
0.600
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs CentroidAlifold(20)
Matthews Correlation Coefficient
Sfold:
0.469
CentroidAlifold(20):
0.411
Sensitivity
Sfold:
0.412
CentroidAlifold(20):
0.190
Positive Predictive Value
Sfold:
0.535
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs CentroidAlifold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
CentroidAlifold(20):
0.411
Sensitivity
MaxExpect:
0.443
CentroidAlifold(20):
0.190
Positive Predictive Value
MaxExpect:
0.488
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs CentroidAlifold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
CentroidAlifold(20):
0.411
Sensitivity
ProbKnot:
0.446
CentroidAlifold(20):
0.190
Positive Predictive Value
ProbKnot:
0.475
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
CentroidAlifold(20):
0.411
Sensitivity
UNAFold:
0.445
CentroidAlifold(20):
0.190
Positive Predictive Value
UNAFold:
0.446
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CentroidAlifold(20):
0.411
Sensitivity
CentroidAlifold(seed):
0.213
CentroidAlifold(20):
0.190
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs CentroidAlifold(20)
Matthews Correlation Coefficient
Fold:
0.437
CentroidAlifold(20):
0.411
Sensitivity
Fold:
0.432
CentroidAlifold(20):
0.190
Positive Predictive Value
Fold:
0.444
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs CentroidAlifold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
CentroidAlifold(20):
0.411
Sensitivity
RNAfold:
0.422
CentroidAlifold(20):
0.190
Positive Predictive Value
RNAfold:
0.437
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CentroidAlifold(20):
0.411
Sensitivity
CentroidHomfold‑LAST:
0.216
CentroidAlifold(20):
0.190
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CentroidAlifold(20):
0.411
Sensitivity
PETfold_pre2.0(20):
0.211
CentroidAlifold(20):
0.190
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
CentroidAlifold(20) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
PknotsRG:
0.386
Sensitivity
CentroidAlifold(20):
0.190
PknotsRG:
0.320
Positive Predictive Value
CentroidAlifold(20):
0.891
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
MXScarna(seed):
0.385
Sensitivity
CentroidAlifold(20):
0.190
MXScarna(seed):
0.196
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAalifold(20):
0.381
Sensitivity
CentroidAlifold(20):
0.190
RNAalifold(20):
0.185
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAalifold(seed):
0.375
Sensitivity
CentroidAlifold(20):
0.190
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Afold:
0.358
Sensitivity
CentroidAlifold(20):
0.190
Afold:
0.295
Positive Predictive Value
CentroidAlifold(20):
0.891
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
MXScarna(20):
0.359
Sensitivity
CentroidAlifold(20):
0.190
MXScarna(20):
0.181
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
CRWrnafold:
0.346
Sensitivity
CentroidAlifold(20):
0.190
CRWrnafold:
0.275
Positive Predictive Value
CentroidAlifold(20):
0.891
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAsubopt:
0.336
Sensitivity
CentroidAlifold(20):
0.190
RNAsubopt:
0.217
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
McQFold:
0.314
Sensitivity
CentroidAlifold(20):
0.190
McQFold:
0.205
Positive Predictive Value
CentroidAlifold(20):
0.891
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
TurboFold(20):
0.301
Sensitivity
CentroidAlifold(20):
0.190
TurboFold(20):
0.108
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
PPfold(20):
0.298
Sensitivity
CentroidAlifold(20):
0.190
PPfold(20):
0.101
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAshapes:
0.295
Sensitivity
CentroidAlifold(20):
0.190
RNAshapes:
0.180
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Carnac(20):
0.288
Sensitivity
CentroidAlifold(20):
0.190
Carnac(20):
0.093
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNASLOpt:
0.281
Sensitivity
CentroidAlifold(20):
0.190
RNASLOpt:
0.158
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RSpredict(20):
0.270
Sensitivity
CentroidAlifold(20):
0.190
RSpredict(20):
0.124
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Multilign(20):
0.251
Sensitivity
CentroidAlifold(20):
0.190
Multilign(20):
0.083
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAwolf:
0.251
Sensitivity
CentroidAlifold(20):
0.190
RNAwolf:
0.191
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Murlet(20):
0.230
Sensitivity
CentroidAlifold(20):
0.190
Murlet(20):
0.067
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Vsfold4:
0.217
Sensitivity
CentroidAlifold(20):
0.190
Vsfold4:
0.125
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Vsfold5:
0.211
Sensitivity
CentroidAlifold(20):
0.190
Vsfold5:
0.125
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RSpredict(seed):
0.212
Sensitivity
CentroidAlifold(20):
0.190
RSpredict(seed):
0.075
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
CMfinder(20):
0.210
Sensitivity
CentroidAlifold(20):
0.190
CMfinder(20):
0.055
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
HotKnots:
0.179
Sensitivity
CentroidAlifold(20):
0.190
HotKnots:
0.051
Positive Predictive Value
CentroidAlifold(20):
0.891
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNASampler(20):
0.176
Sensitivity
CentroidAlifold(20):
0.190
RNASampler(20):
0.035
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Cylofold:
0.155
Sensitivity
CentroidAlifold(20):
0.190
Cylofold:
0.040
Positive Predictive Value
CentroidAlifold(20):
0.891
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Pknots:
0.143
Sensitivity
CentroidAlifold(20):
0.190
Pknots:
0.033
Positive Predictive Value
CentroidAlifold(20):
0.891
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Mastr(20):
0.134
Sensitivity
CentroidAlifold(20):
0.190
Mastr(20):
0.023
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Alterna:
0.129
Sensitivity
CentroidAlifold(20):
0.190
Alterna:
0.030
Positive Predictive Value
CentroidAlifold(20):
0.891
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
MCFold:
0.125
Sensitivity
CentroidAlifold(20):
0.190
MCFold:
0.036
Positive Predictive Value
CentroidAlifold(20):
0.891
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RDfolder:
0.122
Sensitivity
CentroidAlifold(20):
0.190
RDfolder:
0.025
Positive Predictive Value
CentroidAlifold(20):
0.891
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Murlet(seed):
0.079
Sensitivity
CentroidAlifold(20):
0.190
Murlet(seed):
0.008
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
CMfinder(seed):
0.080
Sensitivity
CentroidAlifold(20):
0.190
CMfinder(seed):
0.008
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
TurboFold(seed):
0.076
Sensitivity
CentroidAlifold(20):
0.190
TurboFold(seed):
0.007
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
PPfold(seed):
0.069
Sensitivity
CentroidAlifold(20):
0.190
PPfold(seed):
0.006
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Carnac(seed):
0.066
Sensitivity
CentroidAlifold(20):
0.190
Carnac(seed):
0.005
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNASampler(seed):
0.063
Sensitivity
CentroidAlifold(20):
0.190
RNASampler(seed):
0.005
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
NanoFolder:
0.052
Sensitivity
CentroidAlifold(20):
0.190
NanoFolder:
0.016
Positive Predictive Value
CentroidAlifold(20):
0.891
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Mastr(seed):
0.051
Sensitivity
CentroidAlifold(20):
0.190
Mastr(seed):
0.003
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Multilign(seed):
0.043
Sensitivity
CentroidAlifold(20):
0.190
Multilign(seed):
0.003
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PknotsRG |
1987
ContextFold vs PknotsRG
Matthews Correlation Coefficient
ContextFold:
0.680
PknotsRG:
0.386
Sensitivity
ContextFold:
0.644
PknotsRG:
0.320
Positive Predictive Value
ContextFold:
0.718
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs PknotsRG
Matthews Correlation Coefficient
IPknot:
0.548
PknotsRG:
0.386
Sensitivity
IPknot:
0.475
PknotsRG:
0.320
Positive Predictive Value
IPknot:
0.632
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PknotsRG:
0.386
Sensitivity
PETfold_pre2.0(seed):
0.343
PknotsRG:
0.320
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs PknotsRG
Matthews Correlation Coefficient
Contrafold:
0.508
PknotsRG:
0.386
Sensitivity
Contrafold:
0.481
PknotsRG:
0.320
Positive Predictive Value
Contrafold:
0.537
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs PknotsRG
Matthews Correlation Coefficient
CentroidFold:
0.470
PknotsRG:
0.386
Sensitivity
CentroidFold:
0.369
PknotsRG:
0.320
Positive Predictive Value
CentroidFold:
0.600
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs PknotsRG
Matthews Correlation Coefficient
Sfold:
0.469
PknotsRG:
0.386
Sensitivity
Sfold:
0.412
PknotsRG:
0.320
Positive Predictive Value
Sfold:
0.535
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs PknotsRG
Matthews Correlation Coefficient
MaxExpect:
0.465
PknotsRG:
0.386
Sensitivity
MaxExpect:
0.443
PknotsRG:
0.320
Positive Predictive Value
MaxExpect:
0.488
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs PknotsRG
Matthews Correlation Coefficient
ProbKnot:
0.460
PknotsRG:
0.386
Sensitivity
ProbKnot:
0.446
PknotsRG:
0.320
Positive Predictive Value
ProbKnot:
0.475
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs PknotsRG
Matthews Correlation Coefficient
UNAFold:
0.445
PknotsRG:
0.386
Sensitivity
UNAFold:
0.445
PknotsRG:
0.320
Positive Predictive Value
UNAFold:
0.446
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PknotsRG:
0.386
Sensitivity
CentroidAlifold(seed):
0.213
PknotsRG:
0.320
Positive Predictive Value
CentroidAlifold(seed):
0.901
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs PknotsRG
Matthews Correlation Coefficient
Fold:
0.437
PknotsRG:
0.386
Sensitivity
Fold:
0.432
PknotsRG:
0.320
Positive Predictive Value
Fold:
0.444
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs PknotsRG
Matthews Correlation Coefficient
RNAfold:
0.429
PknotsRG:
0.386
Sensitivity
RNAfold:
0.422
PknotsRG:
0.320
Positive Predictive Value
RNAfold:
0.437
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs PknotsRG
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PknotsRG:
0.386
Sensitivity
CentroidHomfold‑LAST:
0.216
PknotsRG:
0.320
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
PknotsRG:
0.386
Sensitivity
PETfold_pre2.0(20):
0.211
PknotsRG:
0.320
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
PknotsRG:
0.386
Sensitivity
CentroidAlifold(20):
0.190
PknotsRG:
0.320
Positive Predictive Value
CentroidAlifold(20):
0.891
PknotsRG:
0.465
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
PknotsRG vs MXScarna(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
MXScarna(seed):
0.385
Sensitivity
PknotsRG:
0.320
MXScarna(seed):
0.196
Positive Predictive Value
PknotsRG:
0.465
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.978553375983
|
+
PknotsRG vs RNAalifold(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAalifold(20):
0.381
Sensitivity
PknotsRG:
0.320
RNAalifold(20):
0.185
Positive Predictive Value
PknotsRG:
0.465
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 8.1815824276e-08
|
+
PknotsRG vs RNAalifold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAalifold(seed):
0.375
Sensitivity
PknotsRG:
0.320
RNAalifold(seed):
0.169
Positive Predictive Value
PknotsRG:
0.465
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Afold
Matthews Correlation Coefficient
PknotsRG:
0.386
Afold:
0.358
Sensitivity
PknotsRG:
0.320
Afold:
0.295
Positive Predictive Value
PknotsRG:
0.465
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs MXScarna(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
MXScarna(20):
0.359
Sensitivity
PknotsRG:
0.320
MXScarna(20):
0.181
Positive Predictive Value
PknotsRG:
0.465
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs CRWrnafold
Matthews Correlation Coefficient
PknotsRG:
0.386
CRWrnafold:
0.346
Sensitivity
PknotsRG:
0.320
CRWrnafold:
0.275
Positive Predictive Value
PknotsRG:
0.465
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNAsubopt
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAsubopt:
0.336
Sensitivity
PknotsRG:
0.320
RNAsubopt:
0.217
Positive Predictive Value
PknotsRG:
0.465
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs McQFold
Matthews Correlation Coefficient
PknotsRG:
0.386
McQFold:
0.314
Sensitivity
PknotsRG:
0.320
McQFold:
0.205
Positive Predictive Value
PknotsRG:
0.465
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs TurboFold(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
TurboFold(20):
0.301
Sensitivity
PknotsRG:
0.320
TurboFold(20):
0.108
Positive Predictive Value
PknotsRG:
0.465
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs PPfold(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
PPfold(20):
0.298
Sensitivity
PknotsRG:
0.320
PPfold(20):
0.101
Positive Predictive Value
PknotsRG:
0.465
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNAshapes
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAshapes:
0.295
Sensitivity
PknotsRG:
0.320
RNAshapes:
0.180
Positive Predictive Value
PknotsRG:
0.465
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Carnac(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Carnac(20):
0.288
Sensitivity
PknotsRG:
0.320
Carnac(20):
0.093
Positive Predictive Value
PknotsRG:
0.465
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNASLOpt
Matthews Correlation Coefficient
PknotsRG:
0.386
RNASLOpt:
0.281
Sensitivity
PknotsRG:
0.320
RNASLOpt:
0.158
Positive Predictive Value
PknotsRG:
0.465
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RSpredict(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
RSpredict(20):
0.270
Sensitivity
PknotsRG:
0.320
RSpredict(20):
0.124
Positive Predictive Value
PknotsRG:
0.465
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Multilign(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Multilign(20):
0.251
Sensitivity
PknotsRG:
0.320
Multilign(20):
0.083
Positive Predictive Value
PknotsRG:
0.465
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNAwolf
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAwolf:
0.251
Sensitivity
PknotsRG:
0.320
RNAwolf:
0.191
Positive Predictive Value
PknotsRG:
0.465
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Murlet(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Murlet(20):
0.230
Sensitivity
PknotsRG:
0.320
Murlet(20):
0.067
Positive Predictive Value
PknotsRG:
0.465
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Vsfold4
Matthews Correlation Coefficient
PknotsRG:
0.386
Vsfold4:
0.217
Sensitivity
PknotsRG:
0.320
Vsfold4:
0.125
Positive Predictive Value
PknotsRG:
0.465
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Vsfold5
Matthews Correlation Coefficient
PknotsRG:
0.386
Vsfold5:
0.211
Sensitivity
PknotsRG:
0.320
Vsfold5:
0.125
Positive Predictive Value
PknotsRG:
0.465
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RSpredict(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
RSpredict(seed):
0.212
Sensitivity
PknotsRG:
0.320
RSpredict(seed):
0.075
Positive Predictive Value
PknotsRG:
0.465
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs CMfinder(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
CMfinder(20):
0.210
Sensitivity
PknotsRG:
0.320
CMfinder(20):
0.055
Positive Predictive Value
PknotsRG:
0.465
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs HotKnots
Matthews Correlation Coefficient
PknotsRG:
0.386
HotKnots:
0.179
Sensitivity
PknotsRG:
0.320
HotKnots:
0.051
Positive Predictive Value
PknotsRG:
0.465
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNASampler(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNASampler(20):
0.176
Sensitivity
PknotsRG:
0.320
RNASampler(20):
0.035
Positive Predictive Value
PknotsRG:
0.465
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Cylofold
Matthews Correlation Coefficient
PknotsRG:
0.386
Cylofold:
0.155
Sensitivity
PknotsRG:
0.320
Cylofold:
0.040
Positive Predictive Value
PknotsRG:
0.465
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Pknots
Matthews Correlation Coefficient
PknotsRG:
0.386
Pknots:
0.143
Sensitivity
PknotsRG:
0.320
Pknots:
0.033
Positive Predictive Value
PknotsRG:
0.465
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Mastr(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Mastr(20):
0.134
Sensitivity
PknotsRG:
0.320
Mastr(20):
0.023
Positive Predictive Value
PknotsRG:
0.465
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Alterna
Matthews Correlation Coefficient
PknotsRG:
0.386
Alterna:
0.129
Sensitivity
PknotsRG:
0.320
Alterna:
0.030
Positive Predictive Value
PknotsRG:
0.465
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs MCFold
Matthews Correlation Coefficient
PknotsRG:
0.386
MCFold:
0.125
Sensitivity
PknotsRG:
0.320
MCFold:
0.036
Positive Predictive Value
PknotsRG:
0.465
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RDfolder
Matthews Correlation Coefficient
PknotsRG:
0.386
RDfolder:
0.122
Sensitivity
PknotsRG:
0.320
RDfolder:
0.025
Positive Predictive Value
PknotsRG:
0.465
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Murlet(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Murlet(seed):
0.079
Sensitivity
PknotsRG:
0.320
Murlet(seed):
0.008
Positive Predictive Value
PknotsRG:
0.465
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs CMfinder(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
CMfinder(seed):
0.080
Sensitivity
PknotsRG:
0.320
CMfinder(seed):
0.008
Positive Predictive Value
PknotsRG:
0.465
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs TurboFold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
TurboFold(seed):
0.076
Sensitivity
PknotsRG:
0.320
TurboFold(seed):
0.007
Positive Predictive Value
PknotsRG:
0.465
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs PPfold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
PPfold(seed):
0.069
Sensitivity
PknotsRG:
0.320
PPfold(seed):
0.006
Positive Predictive Value
PknotsRG:
0.465
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Carnac(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Carnac(seed):
0.066
Sensitivity
PknotsRG:
0.320
Carnac(seed):
0.005
Positive Predictive Value
PknotsRG:
0.465
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNASampler(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNASampler(seed):
0.063
Sensitivity
PknotsRG:
0.320
RNASampler(seed):
0.005
Positive Predictive Value
PknotsRG:
0.465
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs NanoFolder
Matthews Correlation Coefficient
PknotsRG:
0.386
NanoFolder:
0.052
Sensitivity
PknotsRG:
0.320
NanoFolder:
0.016
Positive Predictive Value
PknotsRG:
0.465
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Mastr(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Mastr(seed):
0.051
Sensitivity
PknotsRG:
0.320
Mastr(seed):
0.003
Positive Predictive Value
PknotsRG:
0.465
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Multilign(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Multilign(seed):
0.043
Sensitivity
PknotsRG:
0.320
Multilign(seed):
0.003
Positive Predictive Value
PknotsRG:
0.465
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MXScarna(seed) |
1987
ContextFold vs MXScarna(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
MXScarna(seed):
0.385
Sensitivity
ContextFold:
0.644
MXScarna(seed):
0.196
Positive Predictive Value
ContextFold:
0.718
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs MXScarna(seed)
Matthews Correlation Coefficient
IPknot:
0.548
MXScarna(seed):
0.385
Sensitivity
IPknot:
0.475
MXScarna(seed):
0.196
Positive Predictive Value
IPknot:
0.632
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MXScarna(seed):
0.385
Sensitivity
PETfold_pre2.0(seed):
0.343
MXScarna(seed):
0.196
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs MXScarna(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
MXScarna(seed):
0.385
Sensitivity
Contrafold:
0.481
MXScarna(seed):
0.196
Positive Predictive Value
Contrafold:
0.537
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
MXScarna(seed):
0.385
Sensitivity
CentroidFold:
0.369
MXScarna(seed):
0.196
Positive Predictive Value
CentroidFold:
0.600
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs MXScarna(seed)
Matthews Correlation Coefficient
Sfold:
0.469
MXScarna(seed):
0.385
Sensitivity
Sfold:
0.412
MXScarna(seed):
0.196
Positive Predictive Value
Sfold:
0.535
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs MXScarna(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
MXScarna(seed):
0.385
Sensitivity
MaxExpect:
0.443
MXScarna(seed):
0.196
Positive Predictive Value
MaxExpect:
0.488
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs MXScarna(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
MXScarna(seed):
0.385
Sensitivity
ProbKnot:
0.446
MXScarna(seed):
0.196
Positive Predictive Value
ProbKnot:
0.475
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs MXScarna(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
MXScarna(seed):
0.385
Sensitivity
UNAFold:
0.445
MXScarna(seed):
0.196
Positive Predictive Value
UNAFold:
0.446
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
MXScarna(seed):
0.385
Sensitivity
CentroidAlifold(seed):
0.213
MXScarna(seed):
0.196
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs MXScarna(seed)
Matthews Correlation Coefficient
Fold:
0.437
MXScarna(seed):
0.385
Sensitivity
Fold:
0.432
MXScarna(seed):
0.196
Positive Predictive Value
Fold:
0.444
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs MXScarna(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
MXScarna(seed):
0.385
Sensitivity
RNAfold:
0.422
MXScarna(seed):
0.196
Positive Predictive Value
RNAfold:
0.437
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
MXScarna(seed):
0.385
Sensitivity
CentroidHomfold‑LAST:
0.216
MXScarna(seed):
0.196
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
MXScarna(seed):
0.385
Sensitivity
PETfold_pre2.0(20):
0.211
MXScarna(seed):
0.196
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
MXScarna(seed):
0.385
Sensitivity
CentroidAlifold(20):
0.190
MXScarna(seed):
0.196
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs MXScarna(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
MXScarna(seed):
0.385
Sensitivity
PknotsRG:
0.320
MXScarna(seed):
0.196
Positive Predictive Value
PknotsRG:
0.465
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.978553375983
|
|
+
MXScarna(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAalifold(20):
0.381
Sensitivity
MXScarna(seed):
0.196
RNAalifold(20):
0.185
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAalifold(seed):
0.375
Sensitivity
MXScarna(seed):
0.196
RNAalifold(seed):
0.169
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Afold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Afold:
0.358
Sensitivity
MXScarna(seed):
0.196
Afold:
0.295
Positive Predictive Value
MXScarna(seed):
0.753
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs MXScarna(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
MXScarna(20):
0.359
Sensitivity
MXScarna(seed):
0.196
MXScarna(20):
0.181
Positive Predictive Value
MXScarna(seed):
0.753
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
CRWrnafold:
0.346
Sensitivity
MXScarna(seed):
0.196
CRWrnafold:
0.275
Positive Predictive Value
MXScarna(seed):
0.753
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAsubopt:
0.336
Sensitivity
MXScarna(seed):
0.196
RNAsubopt:
0.217
Positive Predictive Value
MXScarna(seed):
0.753
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs McQFold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
McQFold:
0.314
Sensitivity
MXScarna(seed):
0.196
McQFold:
0.205
Positive Predictive Value
MXScarna(seed):
0.753
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
TurboFold(20):
0.301
Sensitivity
MXScarna(seed):
0.196
TurboFold(20):
0.108
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
PPfold(20):
0.298
Sensitivity
MXScarna(seed):
0.196
PPfold(20):
0.101
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAshapes:
0.295
Sensitivity
MXScarna(seed):
0.196
RNAshapes:
0.180
Positive Predictive Value
MXScarna(seed):
0.753
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Carnac(20):
0.288
Sensitivity
MXScarna(seed):
0.196
Carnac(20):
0.093
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNASLOpt:
0.281
Sensitivity
MXScarna(seed):
0.196
RNASLOpt:
0.158
Positive Predictive Value
MXScarna(seed):
0.753
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RSpredict(20):
0.270
Sensitivity
MXScarna(seed):
0.196
RSpredict(20):
0.124
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Multilign(20):
0.251
Sensitivity
MXScarna(seed):
0.196
Multilign(20):
0.083
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAwolf:
0.251
Sensitivity
MXScarna(seed):
0.196
RNAwolf:
0.191
Positive Predictive Value
MXScarna(seed):
0.753
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Murlet(20):
0.230
Sensitivity
MXScarna(seed):
0.196
Murlet(20):
0.067
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Vsfold4:
0.217
Sensitivity
MXScarna(seed):
0.196
Vsfold4:
0.125
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Vsfold5:
0.211
Sensitivity
MXScarna(seed):
0.196
Vsfold5:
0.125
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RSpredict(seed):
0.212
Sensitivity
MXScarna(seed):
0.196
RSpredict(seed):
0.075
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
CMfinder(20):
0.210
Sensitivity
MXScarna(seed):
0.196
CMfinder(20):
0.055
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs HotKnots
Matthews Correlation Coefficient
MXScarna(seed):
0.385
HotKnots:
0.179
Sensitivity
MXScarna(seed):
0.196
HotKnots:
0.051
Positive Predictive Value
MXScarna(seed):
0.753
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNASampler(20):
0.176
Sensitivity
MXScarna(seed):
0.196
RNASampler(20):
0.035
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Cylofold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Cylofold:
0.155
Sensitivity
MXScarna(seed):
0.196
Cylofold:
0.040
Positive Predictive Value
MXScarna(seed):
0.753
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Pknots
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Pknots:
0.143
Sensitivity
MXScarna(seed):
0.196
Pknots:
0.033
Positive Predictive Value
MXScarna(seed):
0.753
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Mastr(20):
0.134
Sensitivity
MXScarna(seed):
0.196
Mastr(20):
0.023
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Alterna
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Alterna:
0.129
Sensitivity
MXScarna(seed):
0.196
Alterna:
0.030
Positive Predictive Value
MXScarna(seed):
0.753
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs MCFold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
MCFold:
0.125
Sensitivity
MXScarna(seed):
0.196
MCFold:
0.036
Positive Predictive Value
MXScarna(seed):
0.753
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RDfolder
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RDfolder:
0.122
Sensitivity
MXScarna(seed):
0.196
RDfolder:
0.025
Positive Predictive Value
MXScarna(seed):
0.753
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Murlet(seed):
0.079
Sensitivity
MXScarna(seed):
0.196
Murlet(seed):
0.008
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
CMfinder(seed):
0.080
Sensitivity
MXScarna(seed):
0.196
CMfinder(seed):
0.008
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
TurboFold(seed):
0.076
Sensitivity
MXScarna(seed):
0.196
TurboFold(seed):
0.007
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
PPfold(seed):
0.069
Sensitivity
MXScarna(seed):
0.196
PPfold(seed):
0.006
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Carnac(seed):
0.066
Sensitivity
MXScarna(seed):
0.196
Carnac(seed):
0.005
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNASampler(seed):
0.063
Sensitivity
MXScarna(seed):
0.196
RNASampler(seed):
0.005
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(seed):
0.385
NanoFolder:
0.052
Sensitivity
MXScarna(seed):
0.196
NanoFolder:
0.016
Positive Predictive Value
MXScarna(seed):
0.753
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Mastr(seed):
0.051
Sensitivity
MXScarna(seed):
0.196
Mastr(seed):
0.003
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Multilign(seed):
0.043
Sensitivity
MXScarna(seed):
0.196
Multilign(seed):
0.003
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAalifold(20) |
1987
ContextFold vs RNAalifold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
RNAalifold(20):
0.381
Sensitivity
ContextFold:
0.644
RNAalifold(20):
0.185
Positive Predictive Value
ContextFold:
0.718
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNAalifold(20)
Matthews Correlation Coefficient
IPknot:
0.548
RNAalifold(20):
0.381
Sensitivity
IPknot:
0.475
RNAalifold(20):
0.185
Positive Predictive Value
IPknot:
0.632
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAalifold(20):
0.381
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAalifold(20):
0.185
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNAalifold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
RNAalifold(20):
0.381
Sensitivity
Contrafold:
0.481
RNAalifold(20):
0.185
Positive Predictive Value
Contrafold:
0.537
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAalifold(20):
0.381
Sensitivity
CentroidFold:
0.369
RNAalifold(20):
0.185
Positive Predictive Value
CentroidFold:
0.600
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNAalifold(20)
Matthews Correlation Coefficient
Sfold:
0.469
RNAalifold(20):
0.381
Sensitivity
Sfold:
0.412
RNAalifold(20):
0.185
Positive Predictive Value
Sfold:
0.535
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNAalifold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAalifold(20):
0.381
Sensitivity
MaxExpect:
0.443
RNAalifold(20):
0.185
Positive Predictive Value
MaxExpect:
0.488
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNAalifold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAalifold(20):
0.381
Sensitivity
ProbKnot:
0.446
RNAalifold(20):
0.185
Positive Predictive Value
ProbKnot:
0.475
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNAalifold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
RNAalifold(20):
0.381
Sensitivity
UNAFold:
0.445
RNAalifold(20):
0.185
Positive Predictive Value
UNAFold:
0.446
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAalifold(20):
0.381
Sensitivity
CentroidAlifold(seed):
0.213
RNAalifold(20):
0.185
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNAalifold(20)
Matthews Correlation Coefficient
Fold:
0.437
RNAalifold(20):
0.381
Sensitivity
Fold:
0.432
RNAalifold(20):
0.185
Positive Predictive Value
Fold:
0.444
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNAalifold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
RNAalifold(20):
0.381
Sensitivity
RNAfold:
0.422
RNAalifold(20):
0.185
Positive Predictive Value
RNAfold:
0.437
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAalifold(20):
0.381
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAalifold(20):
0.185
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAalifold(20):
0.381
Sensitivity
PETfold_pre2.0(20):
0.211
RNAalifold(20):
0.185
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAalifold(20):
0.381
Sensitivity
CentroidAlifold(20):
0.190
RNAalifold(20):
0.185
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNAalifold(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAalifold(20):
0.381
Sensitivity
PknotsRG:
0.320
RNAalifold(20):
0.185
Positive Predictive Value
PknotsRG:
0.465
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 8.1815824276e-08
|
1987
MXScarna(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAalifold(20):
0.381
Sensitivity
MXScarna(seed):
0.196
RNAalifold(20):
0.185
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAalifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAalifold(seed):
0.375
Sensitivity
RNAalifold(20):
0.185
RNAalifold(seed):
0.169
Positive Predictive Value
RNAalifold(20):
0.787
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Afold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Afold:
0.358
Sensitivity
RNAalifold(20):
0.185
Afold:
0.295
Positive Predictive Value
RNAalifold(20):
0.787
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
MXScarna(20):
0.359
Sensitivity
RNAalifold(20):
0.185
MXScarna(20):
0.181
Positive Predictive Value
RNAalifold(20):
0.787
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
CRWrnafold:
0.346
Sensitivity
RNAalifold(20):
0.185
CRWrnafold:
0.275
Positive Predictive Value
RNAalifold(20):
0.787
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAsubopt:
0.336
Sensitivity
RNAalifold(20):
0.185
RNAsubopt:
0.217
Positive Predictive Value
RNAalifold(20):
0.787
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs McQFold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
McQFold:
0.314
Sensitivity
RNAalifold(20):
0.185
McQFold:
0.205
Positive Predictive Value
RNAalifold(20):
0.787
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
TurboFold(20):
0.301
Sensitivity
RNAalifold(20):
0.185
TurboFold(20):
0.108
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
PPfold(20):
0.298
Sensitivity
RNAalifold(20):
0.185
PPfold(20):
0.101
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAshapes:
0.295
Sensitivity
RNAalifold(20):
0.185
RNAshapes:
0.180
Positive Predictive Value
RNAalifold(20):
0.787
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Carnac(20):
0.288
Sensitivity
RNAalifold(20):
0.185
Carnac(20):
0.093
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNASLOpt:
0.281
Sensitivity
RNAalifold(20):
0.185
RNASLOpt:
0.158
Positive Predictive Value
RNAalifold(20):
0.787
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RSpredict(20):
0.270
Sensitivity
RNAalifold(20):
0.185
RSpredict(20):
0.124
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Multilign(20):
0.251
Sensitivity
RNAalifold(20):
0.185
Multilign(20):
0.083
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAwolf:
0.251
Sensitivity
RNAalifold(20):
0.185
RNAwolf:
0.191
Positive Predictive Value
RNAalifold(20):
0.787
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Murlet(20):
0.230
Sensitivity
RNAalifold(20):
0.185
Murlet(20):
0.067
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Vsfold4:
0.217
Sensitivity
RNAalifold(20):
0.185
Vsfold4:
0.125
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Vsfold5:
0.211
Sensitivity
RNAalifold(20):
0.185
Vsfold5:
0.125
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RSpredict(seed):
0.212
Sensitivity
RNAalifold(20):
0.185
RSpredict(seed):
0.075
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
CMfinder(20):
0.210
Sensitivity
RNAalifold(20):
0.185
CMfinder(20):
0.055
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(20):
0.381
HotKnots:
0.179
Sensitivity
RNAalifold(20):
0.185
HotKnots:
0.051
Positive Predictive Value
RNAalifold(20):
0.787
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNASampler(20):
0.176
Sensitivity
RNAalifold(20):
0.185
RNASampler(20):
0.035
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Cylofold:
0.155
Sensitivity
RNAalifold(20):
0.185
Cylofold:
0.040
Positive Predictive Value
RNAalifold(20):
0.787
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Pknots
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Pknots:
0.143
Sensitivity
RNAalifold(20):
0.185
Pknots:
0.033
Positive Predictive Value
RNAalifold(20):
0.787
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Mastr(20):
0.134
Sensitivity
RNAalifold(20):
0.185
Mastr(20):
0.023
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Alterna
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Alterna:
0.129
Sensitivity
RNAalifold(20):
0.185
Alterna:
0.030
Positive Predictive Value
RNAalifold(20):
0.787
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs MCFold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
MCFold:
0.125
Sensitivity
RNAalifold(20):
0.185
MCFold:
0.036
Positive Predictive Value
RNAalifold(20):
0.787
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RDfolder:
0.122
Sensitivity
RNAalifold(20):
0.185
RDfolder:
0.025
Positive Predictive Value
RNAalifold(20):
0.787
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Murlet(seed):
0.079
Sensitivity
RNAalifold(20):
0.185
Murlet(seed):
0.008
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
CMfinder(seed):
0.080
Sensitivity
RNAalifold(20):
0.185
CMfinder(seed):
0.008
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
TurboFold(seed):
0.076
Sensitivity
RNAalifold(20):
0.185
TurboFold(seed):
0.007
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
PPfold(seed):
0.069
Sensitivity
RNAalifold(20):
0.185
PPfold(seed):
0.006
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Carnac(seed):
0.066
Sensitivity
RNAalifold(20):
0.185
Carnac(seed):
0.005
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNASampler(seed):
0.063
Sensitivity
RNAalifold(20):
0.185
RNASampler(seed):
0.005
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(20):
0.381
NanoFolder:
0.052
Sensitivity
RNAalifold(20):
0.185
NanoFolder:
0.016
Positive Predictive Value
RNAalifold(20):
0.787
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Mastr(seed):
0.051
Sensitivity
RNAalifold(20):
0.185
Mastr(seed):
0.003
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Multilign(seed):
0.043
Sensitivity
RNAalifold(20):
0.185
Multilign(seed):
0.003
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAalifold(seed) |
1987
ContextFold vs RNAalifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
RNAalifold(seed):
0.375
Sensitivity
ContextFold:
0.644
RNAalifold(seed):
0.169
Positive Predictive Value
ContextFold:
0.718
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNAalifold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
RNAalifold(seed):
0.375
Sensitivity
IPknot:
0.475
RNAalifold(seed):
0.169
Positive Predictive Value
IPknot:
0.632
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAalifold(seed):
0.375
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAalifold(seed):
0.169
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNAalifold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
RNAalifold(seed):
0.375
Sensitivity
Contrafold:
0.481
RNAalifold(seed):
0.169
Positive Predictive Value
Contrafold:
0.537
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAalifold(seed):
0.375
Sensitivity
CentroidFold:
0.369
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidFold:
0.600
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNAalifold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
RNAalifold(seed):
0.375
Sensitivity
Sfold:
0.412
RNAalifold(seed):
0.169
Positive Predictive Value
Sfold:
0.535
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNAalifold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAalifold(seed):
0.375
Sensitivity
MaxExpect:
0.443
RNAalifold(seed):
0.169
Positive Predictive Value
MaxExpect:
0.488
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNAalifold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAalifold(seed):
0.375
Sensitivity
ProbKnot:
0.446
RNAalifold(seed):
0.169
Positive Predictive Value
ProbKnot:
0.475
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNAalifold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
RNAalifold(seed):
0.375
Sensitivity
UNAFold:
0.445
RNAalifold(seed):
0.169
Positive Predictive Value
UNAFold:
0.446
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAalifold(seed):
0.375
Sensitivity
CentroidAlifold(seed):
0.213
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNAalifold(seed)
Matthews Correlation Coefficient
Fold:
0.437
RNAalifold(seed):
0.375
Sensitivity
Fold:
0.432
RNAalifold(seed):
0.169
Positive Predictive Value
Fold:
0.444
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNAalifold(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
RNAalifold(seed):
0.375
Sensitivity
RNAfold:
0.422
RNAalifold(seed):
0.169
Positive Predictive Value
RNAfold:
0.437
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAalifold(seed):
0.375
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAalifold(seed):
0.375
Sensitivity
PETfold_pre2.0(20):
0.211
RNAalifold(seed):
0.169
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAalifold(seed):
0.375
Sensitivity
CentroidAlifold(20):
0.190
RNAalifold(seed):
0.169
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNAalifold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAalifold(seed):
0.375
Sensitivity
PknotsRG:
0.320
RNAalifold(seed):
0.169
Positive Predictive Value
PknotsRG:
0.465
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAalifold(seed):
0.375
Sensitivity
MXScarna(seed):
0.196
RNAalifold(seed):
0.169
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAalifold(seed):
0.375
Sensitivity
RNAalifold(20):
0.185
RNAalifold(seed):
0.169
Positive Predictive Value
RNAalifold(20):
0.787
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAalifold(seed) vs Afold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Afold:
0.358
Sensitivity
RNAalifold(seed):
0.169
Afold:
0.295
Positive Predictive Value
RNAalifold(seed):
0.835
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
MXScarna(20):
0.359
Sensitivity
RNAalifold(seed):
0.169
MXScarna(20):
0.181
Positive Predictive Value
RNAalifold(seed):
0.835
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
CRWrnafold:
0.346
Sensitivity
RNAalifold(seed):
0.169
CRWrnafold:
0.275
Positive Predictive Value
RNAalifold(seed):
0.835
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNAsubopt:
0.336
Sensitivity
RNAalifold(seed):
0.169
RNAsubopt:
0.217
Positive Predictive Value
RNAalifold(seed):
0.835
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs McQFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
McQFold:
0.314
Sensitivity
RNAalifold(seed):
0.169
McQFold:
0.205
Positive Predictive Value
RNAalifold(seed):
0.835
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
TurboFold(20):
0.301
Sensitivity
RNAalifold(seed):
0.169
TurboFold(20):
0.108
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
PPfold(20):
0.298
Sensitivity
RNAalifold(seed):
0.169
PPfold(20):
0.101
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNAshapes:
0.295
Sensitivity
RNAalifold(seed):
0.169
RNAshapes:
0.180
Positive Predictive Value
RNAalifold(seed):
0.835
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Carnac(20):
0.288
Sensitivity
RNAalifold(seed):
0.169
Carnac(20):
0.093
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNASLOpt:
0.281
Sensitivity
RNAalifold(seed):
0.169
RNASLOpt:
0.158
Positive Predictive Value
RNAalifold(seed):
0.835
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RSpredict(20):
0.270
Sensitivity
RNAalifold(seed):
0.169
RSpredict(20):
0.124
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Multilign(20):
0.251
Sensitivity
RNAalifold(seed):
0.169
Multilign(20):
0.083
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNAwolf:
0.251
Sensitivity
RNAalifold(seed):
0.169
RNAwolf:
0.191
Positive Predictive Value
RNAalifold(seed):
0.835
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Murlet(20):
0.230
Sensitivity
RNAalifold(seed):
0.169
Murlet(20):
0.067
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Vsfold4:
0.217
Sensitivity
RNAalifold(seed):
0.169
Vsfold4:
0.125
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Vsfold5:
0.211
Sensitivity
RNAalifold(seed):
0.169
Vsfold5:
0.125
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RSpredict(seed):
0.212
Sensitivity
RNAalifold(seed):
0.169
RSpredict(seed):
0.075
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
CMfinder(20):
0.210
Sensitivity
RNAalifold(seed):
0.169
CMfinder(20):
0.055
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
HotKnots:
0.179
Sensitivity
RNAalifold(seed):
0.169
HotKnots:
0.051
Positive Predictive Value
RNAalifold(seed):
0.835
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNASampler(20):
0.176
Sensitivity
RNAalifold(seed):
0.169
RNASampler(20):
0.035
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Cylofold:
0.155
Sensitivity
RNAalifold(seed):
0.169
Cylofold:
0.040
Positive Predictive Value
RNAalifold(seed):
0.835
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Pknots
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Pknots:
0.143
Sensitivity
RNAalifold(seed):
0.169
Pknots:
0.033
Positive Predictive Value
RNAalifold(seed):
0.835
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Mastr(20):
0.134
Sensitivity
RNAalifold(seed):
0.169
Mastr(20):
0.023
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Alterna
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Alterna:
0.129
Sensitivity
RNAalifold(seed):
0.169
Alterna:
0.030
Positive Predictive Value
RNAalifold(seed):
0.835
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs MCFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
MCFold:
0.125
Sensitivity
RNAalifold(seed):
0.169
MCFold:
0.036
Positive Predictive Value
RNAalifold(seed):
0.835
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RDfolder:
0.122
Sensitivity
RNAalifold(seed):
0.169
RDfolder:
0.025
Positive Predictive Value
RNAalifold(seed):
0.835
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Murlet(seed):
0.079
Sensitivity
RNAalifold(seed):
0.169
Murlet(seed):
0.008
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
CMfinder(seed):
0.080
Sensitivity
RNAalifold(seed):
0.169
CMfinder(seed):
0.008
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
TurboFold(seed):
0.076
Sensitivity
RNAalifold(seed):
0.169
TurboFold(seed):
0.007
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
PPfold(seed):
0.069
Sensitivity
RNAalifold(seed):
0.169
PPfold(seed):
0.006
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Carnac(seed):
0.066
Sensitivity
RNAalifold(seed):
0.169
Carnac(seed):
0.005
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNASampler(seed):
0.063
Sensitivity
RNAalifold(seed):
0.169
RNASampler(seed):
0.005
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
NanoFolder:
0.052
Sensitivity
RNAalifold(seed):
0.169
NanoFolder:
0.016
Positive Predictive Value
RNAalifold(seed):
0.835
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Mastr(seed):
0.051
Sensitivity
RNAalifold(seed):
0.169
Mastr(seed):
0.003
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Multilign(seed):
0.043
Sensitivity
RNAalifold(seed):
0.169
Multilign(seed):
0.003
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Afold |
1987
ContextFold vs Afold
Matthews Correlation Coefficient
ContextFold:
0.680
Afold:
0.358
Sensitivity
ContextFold:
0.644
Afold:
0.295
Positive Predictive Value
ContextFold:
0.718
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Afold
Matthews Correlation Coefficient
IPknot:
0.548
Afold:
0.358
Sensitivity
IPknot:
0.475
Afold:
0.295
Positive Predictive Value
IPknot:
0.632
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Afold:
0.358
Sensitivity
PETfold_pre2.0(seed):
0.343
Afold:
0.295
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Afold
Matthews Correlation Coefficient
Contrafold:
0.508
Afold:
0.358
Sensitivity
Contrafold:
0.481
Afold:
0.295
Positive Predictive Value
Contrafold:
0.537
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Afold
Matthews Correlation Coefficient
CentroidFold:
0.470
Afold:
0.358
Sensitivity
CentroidFold:
0.369
Afold:
0.295
Positive Predictive Value
CentroidFold:
0.600
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Afold
Matthews Correlation Coefficient
Sfold:
0.469
Afold:
0.358
Sensitivity
Sfold:
0.412
Afold:
0.295
Positive Predictive Value
Sfold:
0.535
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Afold
Matthews Correlation Coefficient
MaxExpect:
0.465
Afold:
0.358
Sensitivity
MaxExpect:
0.443
Afold:
0.295
Positive Predictive Value
MaxExpect:
0.488
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Afold
Matthews Correlation Coefficient
ProbKnot:
0.460
Afold:
0.358
Sensitivity
ProbKnot:
0.446
Afold:
0.295
Positive Predictive Value
ProbKnot:
0.475
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Afold
Matthews Correlation Coefficient
UNAFold:
0.445
Afold:
0.358
Sensitivity
UNAFold:
0.445
Afold:
0.295
Positive Predictive Value
UNAFold:
0.446
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Afold:
0.358
Sensitivity
CentroidAlifold(seed):
0.213
Afold:
0.295
Positive Predictive Value
CentroidAlifold(seed):
0.901
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Afold
Matthews Correlation Coefficient
Fold:
0.437
Afold:
0.358
Sensitivity
Fold:
0.432
Afold:
0.295
Positive Predictive Value
Fold:
0.444
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Afold
Matthews Correlation Coefficient
RNAfold:
0.429
Afold:
0.358
Sensitivity
RNAfold:
0.422
Afold:
0.295
Positive Predictive Value
RNAfold:
0.437
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Afold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Afold:
0.358
Sensitivity
CentroidHomfold‑LAST:
0.216
Afold:
0.295
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Afold:
0.358
Sensitivity
PETfold_pre2.0(20):
0.211
Afold:
0.295
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Afold:
0.358
Sensitivity
CentroidAlifold(20):
0.190
Afold:
0.295
Positive Predictive Value
CentroidAlifold(20):
0.891
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Afold
Matthews Correlation Coefficient
PknotsRG:
0.386
Afold:
0.358
Sensitivity
PknotsRG:
0.320
Afold:
0.295
Positive Predictive Value
PknotsRG:
0.465
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Afold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Afold:
0.358
Sensitivity
MXScarna(seed):
0.196
Afold:
0.295
Positive Predictive Value
MXScarna(seed):
0.753
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Afold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Afold:
0.358
Sensitivity
RNAalifold(20):
0.185
Afold:
0.295
Positive Predictive Value
RNAalifold(20):
0.787
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Afold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Afold:
0.358
Sensitivity
RNAalifold(seed):
0.169
Afold:
0.295
Positive Predictive Value
RNAalifold(seed):
0.835
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
MXScarna(20) vs Afold
Matthews Correlation Coefficient
MXScarna(20):
0.359
Afold:
0.358
Sensitivity
MXScarna(20):
0.181
Afold:
0.295
Positive Predictive Value
MXScarna(20):
0.710
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.00227948062498
|
+
Afold vs CRWrnafold
Matthews Correlation Coefficient
Afold:
0.358
CRWrnafold:
0.346
Sensitivity
Afold:
0.295
CRWrnafold:
0.275
Positive Predictive Value
Afold:
0.434
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNAsubopt
Matthews Correlation Coefficient
Afold:
0.358
RNAsubopt:
0.336
Sensitivity
Afold:
0.295
RNAsubopt:
0.217
Positive Predictive Value
Afold:
0.434
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs McQFold
Matthews Correlation Coefficient
Afold:
0.358
McQFold:
0.314
Sensitivity
Afold:
0.295
McQFold:
0.205
Positive Predictive Value
Afold:
0.434
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs TurboFold(20)
Matthews Correlation Coefficient
Afold:
0.358
TurboFold(20):
0.301
Sensitivity
Afold:
0.295
TurboFold(20):
0.108
Positive Predictive Value
Afold:
0.434
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs PPfold(20)
Matthews Correlation Coefficient
Afold:
0.358
PPfold(20):
0.298
Sensitivity
Afold:
0.295
PPfold(20):
0.101
Positive Predictive Value
Afold:
0.434
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNAshapes
Matthews Correlation Coefficient
Afold:
0.358
RNAshapes:
0.295
Sensitivity
Afold:
0.295
RNAshapes:
0.180
Positive Predictive Value
Afold:
0.434
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Carnac(20)
Matthews Correlation Coefficient
Afold:
0.358
Carnac(20):
0.288
Sensitivity
Afold:
0.295
Carnac(20):
0.093
Positive Predictive Value
Afold:
0.434
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNASLOpt
Matthews Correlation Coefficient
Afold:
0.358
RNASLOpt:
0.281
Sensitivity
Afold:
0.295
RNASLOpt:
0.158
Positive Predictive Value
Afold:
0.434
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RSpredict(20)
Matthews Correlation Coefficient
Afold:
0.358
RSpredict(20):
0.270
Sensitivity
Afold:
0.295
RSpredict(20):
0.124
Positive Predictive Value
Afold:
0.434
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Multilign(20)
Matthews Correlation Coefficient
Afold:
0.358
Multilign(20):
0.251
Sensitivity
Afold:
0.295
Multilign(20):
0.083
Positive Predictive Value
Afold:
0.434
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNAwolf
Matthews Correlation Coefficient
Afold:
0.358
RNAwolf:
0.251
Sensitivity
Afold:
0.295
RNAwolf:
0.191
Positive Predictive Value
Afold:
0.434
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Murlet(20)
Matthews Correlation Coefficient
Afold:
0.358
Murlet(20):
0.230
Sensitivity
Afold:
0.295
Murlet(20):
0.067
Positive Predictive Value
Afold:
0.434
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Vsfold4
Matthews Correlation Coefficient
Afold:
0.358
Vsfold4:
0.217
Sensitivity
Afold:
0.295
Vsfold4:
0.125
Positive Predictive Value
Afold:
0.434
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Vsfold5
Matthews Correlation Coefficient
Afold:
0.358
Vsfold5:
0.211
Sensitivity
Afold:
0.295
Vsfold5:
0.125
Positive Predictive Value
Afold:
0.434
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RSpredict(seed)
Matthews Correlation Coefficient
Afold:
0.358
RSpredict(seed):
0.212
Sensitivity
Afold:
0.295
RSpredict(seed):
0.075
Positive Predictive Value
Afold:
0.434
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs CMfinder(20)
Matthews Correlation Coefficient
Afold:
0.358
CMfinder(20):
0.210
Sensitivity
Afold:
0.295
CMfinder(20):
0.055
Positive Predictive Value
Afold:
0.434
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs HotKnots
Matthews Correlation Coefficient
Afold:
0.358
HotKnots:
0.179
Sensitivity
Afold:
0.295
HotKnots:
0.051
Positive Predictive Value
Afold:
0.434
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNASampler(20)
Matthews Correlation Coefficient
Afold:
0.358
RNASampler(20):
0.176
Sensitivity
Afold:
0.295
RNASampler(20):
0.035
Positive Predictive Value
Afold:
0.434
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Cylofold
Matthews Correlation Coefficient
Afold:
0.358
Cylofold:
0.155
Sensitivity
Afold:
0.295
Cylofold:
0.040
Positive Predictive Value
Afold:
0.434
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Pknots
Matthews Correlation Coefficient
Afold:
0.358
Pknots:
0.143
Sensitivity
Afold:
0.295
Pknots:
0.033
Positive Predictive Value
Afold:
0.434
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Mastr(20)
Matthews Correlation Coefficient
Afold:
0.358
Mastr(20):
0.134
Sensitivity
Afold:
0.295
Mastr(20):
0.023
Positive Predictive Value
Afold:
0.434
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Alterna
Matthews Correlation Coefficient
Afold:
0.358
Alterna:
0.129
Sensitivity
Afold:
0.295
Alterna:
0.030
Positive Predictive Value
Afold:
0.434
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs MCFold
Matthews Correlation Coefficient
Afold:
0.358
MCFold:
0.125
Sensitivity
Afold:
0.295
MCFold:
0.036
Positive Predictive Value
Afold:
0.434
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RDfolder
Matthews Correlation Coefficient
Afold:
0.358
RDfolder:
0.122
Sensitivity
Afold:
0.295
RDfolder:
0.025
Positive Predictive Value
Afold:
0.434
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Murlet(seed)
Matthews Correlation Coefficient
Afold:
0.358
Murlet(seed):
0.079
Sensitivity
Afold:
0.295
Murlet(seed):
0.008
Positive Predictive Value
Afold:
0.434
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs CMfinder(seed)
Matthews Correlation Coefficient
Afold:
0.358
CMfinder(seed):
0.080
Sensitivity
Afold:
0.295
CMfinder(seed):
0.008
Positive Predictive Value
Afold:
0.434
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs TurboFold(seed)
Matthews Correlation Coefficient
Afold:
0.358
TurboFold(seed):
0.076
Sensitivity
Afold:
0.295
TurboFold(seed):
0.007
Positive Predictive Value
Afold:
0.434
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs PPfold(seed)
Matthews Correlation Coefficient
Afold:
0.358
PPfold(seed):
0.069
Sensitivity
Afold:
0.295
PPfold(seed):
0.006
Positive Predictive Value
Afold:
0.434
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Carnac(seed)
Matthews Correlation Coefficient
Afold:
0.358
Carnac(seed):
0.066
Sensitivity
Afold:
0.295
Carnac(seed):
0.005
Positive Predictive Value
Afold:
0.434
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNASampler(seed)
Matthews Correlation Coefficient
Afold:
0.358
RNASampler(seed):
0.063
Sensitivity
Afold:
0.295
RNASampler(seed):
0.005
Positive Predictive Value
Afold:
0.434
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs NanoFolder
Matthews Correlation Coefficient
Afold:
0.358
NanoFolder:
0.052
Sensitivity
Afold:
0.295
NanoFolder:
0.016
Positive Predictive Value
Afold:
0.434
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Mastr(seed)
Matthews Correlation Coefficient
Afold:
0.358
Mastr(seed):
0.051
Sensitivity
Afold:
0.295
Mastr(seed):
0.003
Positive Predictive Value
Afold:
0.434
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Multilign(seed)
Matthews Correlation Coefficient
Afold:
0.358
Multilign(seed):
0.043
Sensitivity
Afold:
0.295
Multilign(seed):
0.003
Positive Predictive Value
Afold:
0.434
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MXScarna(20) |
1987
ContextFold vs MXScarna(20)
Matthews Correlation Coefficient
ContextFold:
0.680
MXScarna(20):
0.359
Sensitivity
ContextFold:
0.644
MXScarna(20):
0.181
Positive Predictive Value
ContextFold:
0.718
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs MXScarna(20)
Matthews Correlation Coefficient
IPknot:
0.548
MXScarna(20):
0.359
Sensitivity
IPknot:
0.475
MXScarna(20):
0.181
Positive Predictive Value
IPknot:
0.632
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MXScarna(20):
0.359
Sensitivity
PETfold_pre2.0(seed):
0.343
MXScarna(20):
0.181
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs MXScarna(20)
Matthews Correlation Coefficient
Contrafold:
0.508
MXScarna(20):
0.359
Sensitivity
Contrafold:
0.481
MXScarna(20):
0.181
Positive Predictive Value
Contrafold:
0.537
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs MXScarna(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
MXScarna(20):
0.359
Sensitivity
CentroidFold:
0.369
MXScarna(20):
0.181
Positive Predictive Value
CentroidFold:
0.600
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs MXScarna(20)
Matthews Correlation Coefficient
Sfold:
0.469
MXScarna(20):
0.359
Sensitivity
Sfold:
0.412
MXScarna(20):
0.181
Positive Predictive Value
Sfold:
0.535
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs MXScarna(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
MXScarna(20):
0.359
Sensitivity
MaxExpect:
0.443
MXScarna(20):
0.181
Positive Predictive Value
MaxExpect:
0.488
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs MXScarna(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
MXScarna(20):
0.359
Sensitivity
ProbKnot:
0.446
MXScarna(20):
0.181
Positive Predictive Value
ProbKnot:
0.475
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs MXScarna(20)
Matthews Correlation Coefficient
UNAFold:
0.445
MXScarna(20):
0.359
Sensitivity
UNAFold:
0.445
MXScarna(20):
0.181
Positive Predictive Value
UNAFold:
0.446
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
MXScarna(20):
0.359
Sensitivity
CentroidAlifold(seed):
0.213
MXScarna(20):
0.181
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs MXScarna(20)
Matthews Correlation Coefficient
Fold:
0.437
MXScarna(20):
0.359
Sensitivity
Fold:
0.432
MXScarna(20):
0.181
Positive Predictive Value
Fold:
0.444
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs MXScarna(20)
Matthews Correlation Coefficient
RNAfold:
0.429
MXScarna(20):
0.359
Sensitivity
RNAfold:
0.422
MXScarna(20):
0.181
Positive Predictive Value
RNAfold:
0.437
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs MXScarna(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
MXScarna(20):
0.359
Sensitivity
CentroidHomfold‑LAST:
0.216
MXScarna(20):
0.181
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
MXScarna(20):
0.359
Sensitivity
PETfold_pre2.0(20):
0.211
MXScarna(20):
0.181
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
MXScarna(20):
0.359
Sensitivity
CentroidAlifold(20):
0.190
MXScarna(20):
0.181
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs MXScarna(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
MXScarna(20):
0.359
Sensitivity
PknotsRG:
0.320
MXScarna(20):
0.181
Positive Predictive Value
PknotsRG:
0.465
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs MXScarna(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
MXScarna(20):
0.359
Sensitivity
MXScarna(seed):
0.196
MXScarna(20):
0.181
Positive Predictive Value
MXScarna(seed):
0.753
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
MXScarna(20):
0.359
Sensitivity
RNAalifold(20):
0.185
MXScarna(20):
0.181
Positive Predictive Value
RNAalifold(20):
0.787
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
MXScarna(20):
0.359
Sensitivity
RNAalifold(seed):
0.169
MXScarna(20):
0.181
Positive Predictive Value
RNAalifold(seed):
0.835
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Afold
Matthews Correlation Coefficient
MXScarna(20):
0.359
Afold:
0.358
Sensitivity
MXScarna(20):
0.181
Afold:
0.295
Positive Predictive Value
MXScarna(20):
0.710
Afold:
0.434
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.00227948062498
|
|
+
MXScarna(20) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(20):
0.359
CRWrnafold:
0.346
Sensitivity
MXScarna(20):
0.181
CRWrnafold:
0.275
Positive Predictive Value
MXScarna(20):
0.710
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNAsubopt:
0.336
Sensitivity
MXScarna(20):
0.181
RNAsubopt:
0.217
Positive Predictive Value
MXScarna(20):
0.710
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs McQFold
Matthews Correlation Coefficient
MXScarna(20):
0.359
McQFold:
0.314
Sensitivity
MXScarna(20):
0.181
McQFold:
0.205
Positive Predictive Value
MXScarna(20):
0.710
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
TurboFold(20):
0.301
Sensitivity
MXScarna(20):
0.181
TurboFold(20):
0.108
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
PPfold(20):
0.298
Sensitivity
MXScarna(20):
0.181
PPfold(20):
0.101
Positive Predictive Value
MXScarna(20):
0.710
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNAshapes:
0.295
Sensitivity
MXScarna(20):
0.181
RNAshapes:
0.180
Positive Predictive Value
MXScarna(20):
0.710
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Carnac(20):
0.288
Sensitivity
MXScarna(20):
0.181
Carnac(20):
0.093
Positive Predictive Value
MXScarna(20):
0.710
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNASLOpt:
0.281
Sensitivity
MXScarna(20):
0.181
RNASLOpt:
0.158
Positive Predictive Value
MXScarna(20):
0.710
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RSpredict(20):
0.270
Sensitivity
MXScarna(20):
0.181
RSpredict(20):
0.124
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Multilign(20):
0.251
Sensitivity
MXScarna(20):
0.181
Multilign(20):
0.083
Positive Predictive Value
MXScarna(20):
0.710
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNAwolf:
0.251
Sensitivity
MXScarna(20):
0.181
RNAwolf:
0.191
Positive Predictive Value
MXScarna(20):
0.710
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Murlet(20):
0.230
Sensitivity
MXScarna(20):
0.181
Murlet(20):
0.067
Positive Predictive Value
MXScarna(20):
0.710
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(20):
0.359
Vsfold4:
0.217
Sensitivity
MXScarna(20):
0.181
Vsfold4:
0.125
Positive Predictive Value
MXScarna(20):
0.710
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(20):
0.359
Vsfold5:
0.211
Sensitivity
MXScarna(20):
0.181
Vsfold5:
0.125
Positive Predictive Value
MXScarna(20):
0.710
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RSpredict(seed):
0.212
Sensitivity
MXScarna(20):
0.181
RSpredict(seed):
0.075
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
CMfinder(20):
0.210
Sensitivity
MXScarna(20):
0.181
CMfinder(20):
0.055
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs HotKnots
Matthews Correlation Coefficient
MXScarna(20):
0.359
HotKnots:
0.179
Sensitivity
MXScarna(20):
0.181
HotKnots:
0.051
Positive Predictive Value
MXScarna(20):
0.710
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNASampler(20):
0.176
Sensitivity
MXScarna(20):
0.181
RNASampler(20):
0.035
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Cylofold
Matthews Correlation Coefficient
MXScarna(20):
0.359
Cylofold:
0.155
Sensitivity
MXScarna(20):
0.181
Cylofold:
0.040
Positive Predictive Value
MXScarna(20):
0.710
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Pknots
Matthews Correlation Coefficient
MXScarna(20):
0.359
Pknots:
0.143
Sensitivity
MXScarna(20):
0.181
Pknots:
0.033
Positive Predictive Value
MXScarna(20):
0.710
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Mastr(20):
0.134
Sensitivity
MXScarna(20):
0.181
Mastr(20):
0.023
Positive Predictive Value
MXScarna(20):
0.710
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Alterna
Matthews Correlation Coefficient
MXScarna(20):
0.359
Alterna:
0.129
Sensitivity
MXScarna(20):
0.181
Alterna:
0.030
Positive Predictive Value
MXScarna(20):
0.710
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs MCFold
Matthews Correlation Coefficient
MXScarna(20):
0.359
MCFold:
0.125
Sensitivity
MXScarna(20):
0.181
MCFold:
0.036
Positive Predictive Value
MXScarna(20):
0.710
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RDfolder
Matthews Correlation Coefficient
MXScarna(20):
0.359
RDfolder:
0.122
Sensitivity
MXScarna(20):
0.181
RDfolder:
0.025
Positive Predictive Value
MXScarna(20):
0.710
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Murlet(seed):
0.079
Sensitivity
MXScarna(20):
0.181
Murlet(seed):
0.008
Positive Predictive Value
MXScarna(20):
0.710
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
CMfinder(seed):
0.080
Sensitivity
MXScarna(20):
0.181
CMfinder(seed):
0.008
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
TurboFold(seed):
0.076
Sensitivity
MXScarna(20):
0.181
TurboFold(seed):
0.007
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
PPfold(seed):
0.069
Sensitivity
MXScarna(20):
0.181
PPfold(seed):
0.006
Positive Predictive Value
MXScarna(20):
0.710
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Carnac(seed):
0.066
Sensitivity
MXScarna(20):
0.181
Carnac(seed):
0.005
Positive Predictive Value
MXScarna(20):
0.710
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNASampler(seed):
0.063
Sensitivity
MXScarna(20):
0.181
RNASampler(seed):
0.005
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(20):
0.359
NanoFolder:
0.052
Sensitivity
MXScarna(20):
0.181
NanoFolder:
0.016
Positive Predictive Value
MXScarna(20):
0.710
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Mastr(seed):
0.051
Sensitivity
MXScarna(20):
0.181
Mastr(seed):
0.003
Positive Predictive Value
MXScarna(20):
0.710
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Multilign(seed):
0.043
Sensitivity
MXScarna(20):
0.181
Multilign(seed):
0.003
Positive Predictive Value
MXScarna(20):
0.710
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CRWrnafold |
1987
ContextFold vs CRWrnafold
Matthews Correlation Coefficient
ContextFold:
0.680
CRWrnafold:
0.346
Sensitivity
ContextFold:
0.644
CRWrnafold:
0.275
Positive Predictive Value
ContextFold:
0.718
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CRWrnafold
Matthews Correlation Coefficient
IPknot:
0.548
CRWrnafold:
0.346
Sensitivity
IPknot:
0.475
CRWrnafold:
0.275
Positive Predictive Value
IPknot:
0.632
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CRWrnafold:
0.346
Sensitivity
PETfold_pre2.0(seed):
0.343
CRWrnafold:
0.275
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CRWrnafold
Matthews Correlation Coefficient
Contrafold:
0.508
CRWrnafold:
0.346
Sensitivity
Contrafold:
0.481
CRWrnafold:
0.275
Positive Predictive Value
Contrafold:
0.537
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs CRWrnafold
Matthews Correlation Coefficient
CentroidFold:
0.470
CRWrnafold:
0.346
Sensitivity
CentroidFold:
0.369
CRWrnafold:
0.275
Positive Predictive Value
CentroidFold:
0.600
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs CRWrnafold
Matthews Correlation Coefficient
Sfold:
0.469
CRWrnafold:
0.346
Sensitivity
Sfold:
0.412
CRWrnafold:
0.275
Positive Predictive Value
Sfold:
0.535
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs CRWrnafold
Matthews Correlation Coefficient
MaxExpect:
0.465
CRWrnafold:
0.346
Sensitivity
MaxExpect:
0.443
CRWrnafold:
0.275
Positive Predictive Value
MaxExpect:
0.488
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs CRWrnafold
Matthews Correlation Coefficient
ProbKnot:
0.460
CRWrnafold:
0.346
Sensitivity
ProbKnot:
0.446
CRWrnafold:
0.275
Positive Predictive Value
ProbKnot:
0.475
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs CRWrnafold
Matthews Correlation Coefficient
UNAFold:
0.445
CRWrnafold:
0.346
Sensitivity
UNAFold:
0.445
CRWrnafold:
0.275
Positive Predictive Value
UNAFold:
0.446
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CRWrnafold:
0.346
Sensitivity
CentroidAlifold(seed):
0.213
CRWrnafold:
0.275
Positive Predictive Value
CentroidAlifold(seed):
0.901
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs CRWrnafold
Matthews Correlation Coefficient
Fold:
0.437
CRWrnafold:
0.346
Sensitivity
Fold:
0.432
CRWrnafold:
0.275
Positive Predictive Value
Fold:
0.444
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs CRWrnafold
Matthews Correlation Coefficient
RNAfold:
0.429
CRWrnafold:
0.346
Sensitivity
RNAfold:
0.422
CRWrnafold:
0.275
Positive Predictive Value
RNAfold:
0.437
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs CRWrnafold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CRWrnafold:
0.346
Sensitivity
CentroidHomfold‑LAST:
0.216
CRWrnafold:
0.275
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CRWrnafold:
0.346
Sensitivity
PETfold_pre2.0(20):
0.211
CRWrnafold:
0.275
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
CRWrnafold:
0.346
Sensitivity
CentroidAlifold(20):
0.190
CRWrnafold:
0.275
Positive Predictive Value
CentroidAlifold(20):
0.891
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs CRWrnafold
Matthews Correlation Coefficient
PknotsRG:
0.386
CRWrnafold:
0.346
Sensitivity
PknotsRG:
0.320
CRWrnafold:
0.275
Positive Predictive Value
PknotsRG:
0.465
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
CRWrnafold:
0.346
Sensitivity
MXScarna(seed):
0.196
CRWrnafold:
0.275
Positive Predictive Value
MXScarna(seed):
0.753
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
CRWrnafold:
0.346
Sensitivity
RNAalifold(20):
0.185
CRWrnafold:
0.275
Positive Predictive Value
RNAalifold(20):
0.787
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
CRWrnafold:
0.346
Sensitivity
RNAalifold(seed):
0.169
CRWrnafold:
0.275
Positive Predictive Value
RNAalifold(seed):
0.835
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs CRWrnafold
Matthews Correlation Coefficient
Afold:
0.358
CRWrnafold:
0.346
Sensitivity
Afold:
0.295
CRWrnafold:
0.275
Positive Predictive Value
Afold:
0.434
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(20):
0.359
CRWrnafold:
0.346
Sensitivity
MXScarna(20):
0.181
CRWrnafold:
0.275
Positive Predictive Value
MXScarna(20):
0.710
CRWrnafold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
CRWrnafold vs RNAsubopt
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNAsubopt:
0.336
Sensitivity
CRWrnafold:
0.275
RNAsubopt:
0.217
Positive Predictive Value
CRWrnafold:
0.435
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs McQFold
Matthews Correlation Coefficient
CRWrnafold:
0.346
McQFold:
0.314
Sensitivity
CRWrnafold:
0.275
McQFold:
0.205
Positive Predictive Value
CRWrnafold:
0.435
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs TurboFold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
TurboFold(20):
0.301
Sensitivity
CRWrnafold:
0.275
TurboFold(20):
0.108
Positive Predictive Value
CRWrnafold:
0.435
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs PPfold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
PPfold(20):
0.298
Sensitivity
CRWrnafold:
0.275
PPfold(20):
0.101
Positive Predictive Value
CRWrnafold:
0.435
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNAshapes
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNAshapes:
0.295
Sensitivity
CRWrnafold:
0.275
RNAshapes:
0.180
Positive Predictive Value
CRWrnafold:
0.435
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Carnac(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Carnac(20):
0.288
Sensitivity
CRWrnafold:
0.275
Carnac(20):
0.093
Positive Predictive Value
CRWrnafold:
0.435
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNASLOpt
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNASLOpt:
0.281
Sensitivity
CRWrnafold:
0.275
RNASLOpt:
0.158
Positive Predictive Value
CRWrnafold:
0.435
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RSpredict(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RSpredict(20):
0.270
Sensitivity
CRWrnafold:
0.275
RSpredict(20):
0.124
Positive Predictive Value
CRWrnafold:
0.435
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Multilign(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Multilign(20):
0.251
Sensitivity
CRWrnafold:
0.275
Multilign(20):
0.083
Positive Predictive Value
CRWrnafold:
0.435
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNAwolf
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNAwolf:
0.251
Sensitivity
CRWrnafold:
0.275
RNAwolf:
0.191
Positive Predictive Value
CRWrnafold:
0.435
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Murlet(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Murlet(20):
0.230
Sensitivity
CRWrnafold:
0.275
Murlet(20):
0.067
Positive Predictive Value
CRWrnafold:
0.435
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Vsfold4
Matthews Correlation Coefficient
CRWrnafold:
0.346
Vsfold4:
0.217
Sensitivity
CRWrnafold:
0.275
Vsfold4:
0.125
Positive Predictive Value
CRWrnafold:
0.435
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Vsfold5
Matthews Correlation Coefficient
CRWrnafold:
0.346
Vsfold5:
0.211
Sensitivity
CRWrnafold:
0.275
Vsfold5:
0.125
Positive Predictive Value
CRWrnafold:
0.435
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RSpredict(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RSpredict(seed):
0.212
Sensitivity
CRWrnafold:
0.275
RSpredict(seed):
0.075
Positive Predictive Value
CRWrnafold:
0.435
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs CMfinder(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
CMfinder(20):
0.210
Sensitivity
CRWrnafold:
0.275
CMfinder(20):
0.055
Positive Predictive Value
CRWrnafold:
0.435
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs HotKnots
Matthews Correlation Coefficient
CRWrnafold:
0.346
HotKnots:
0.179
Sensitivity
CRWrnafold:
0.275
HotKnots:
0.051
Positive Predictive Value
CRWrnafold:
0.435
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNASampler(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNASampler(20):
0.176
Sensitivity
CRWrnafold:
0.275
RNASampler(20):
0.035
Positive Predictive Value
CRWrnafold:
0.435
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Cylofold
Matthews Correlation Coefficient
CRWrnafold:
0.346
Cylofold:
0.155
Sensitivity
CRWrnafold:
0.275
Cylofold:
0.040
Positive Predictive Value
CRWrnafold:
0.435
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Pknots
Matthews Correlation Coefficient
CRWrnafold:
0.346
Pknots:
0.143
Sensitivity
CRWrnafold:
0.275
Pknots:
0.033
Positive Predictive Value
CRWrnafold:
0.435
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Mastr(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Mastr(20):
0.134
Sensitivity
CRWrnafold:
0.275
Mastr(20):
0.023
Positive Predictive Value
CRWrnafold:
0.435
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Alterna
Matthews Correlation Coefficient
CRWrnafold:
0.346
Alterna:
0.129
Sensitivity
CRWrnafold:
0.275
Alterna:
0.030
Positive Predictive Value
CRWrnafold:
0.435
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs MCFold
Matthews Correlation Coefficient
CRWrnafold:
0.346
MCFold:
0.125
Sensitivity
CRWrnafold:
0.275
MCFold:
0.036
Positive Predictive Value
CRWrnafold:
0.435
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RDfolder
Matthews Correlation Coefficient
CRWrnafold:
0.346
RDfolder:
0.122
Sensitivity
CRWrnafold:
0.275
RDfolder:
0.025
Positive Predictive Value
CRWrnafold:
0.435
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Murlet(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Murlet(seed):
0.079
Sensitivity
CRWrnafold:
0.275
Murlet(seed):
0.008
Positive Predictive Value
CRWrnafold:
0.435
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs CMfinder(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
CMfinder(seed):
0.080
Sensitivity
CRWrnafold:
0.275
CMfinder(seed):
0.008
Positive Predictive Value
CRWrnafold:
0.435
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs TurboFold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
TurboFold(seed):
0.076
Sensitivity
CRWrnafold:
0.275
TurboFold(seed):
0.007
Positive Predictive Value
CRWrnafold:
0.435
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs PPfold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
PPfold(seed):
0.069
Sensitivity
CRWrnafold:
0.275
PPfold(seed):
0.006
Positive Predictive Value
CRWrnafold:
0.435
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Carnac(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Carnac(seed):
0.066
Sensitivity
CRWrnafold:
0.275
Carnac(seed):
0.005
Positive Predictive Value
CRWrnafold:
0.435
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNASampler(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNASampler(seed):
0.063
Sensitivity
CRWrnafold:
0.275
RNASampler(seed):
0.005
Positive Predictive Value
CRWrnafold:
0.435
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs NanoFolder
Matthews Correlation Coefficient
CRWrnafold:
0.346
NanoFolder:
0.052
Sensitivity
CRWrnafold:
0.275
NanoFolder:
0.016
Positive Predictive Value
CRWrnafold:
0.435
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Mastr(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Mastr(seed):
0.051
Sensitivity
CRWrnafold:
0.275
Mastr(seed):
0.003
Positive Predictive Value
CRWrnafold:
0.435
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Multilign(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Multilign(seed):
0.043
Sensitivity
CRWrnafold:
0.275
Multilign(seed):
0.003
Positive Predictive Value
CRWrnafold:
0.435
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAsubopt |
1987
ContextFold vs RNAsubopt
Matthews Correlation Coefficient
ContextFold:
0.680
RNAsubopt:
0.336
Sensitivity
ContextFold:
0.644
RNAsubopt:
0.217
Positive Predictive Value
ContextFold:
0.718
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNAsubopt
Matthews Correlation Coefficient
IPknot:
0.548
RNAsubopt:
0.336
Sensitivity
IPknot:
0.475
RNAsubopt:
0.217
Positive Predictive Value
IPknot:
0.632
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAsubopt:
0.336
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAsubopt:
0.217
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNAsubopt
Matthews Correlation Coefficient
Contrafold:
0.508
RNAsubopt:
0.336
Sensitivity
Contrafold:
0.481
RNAsubopt:
0.217
Positive Predictive Value
Contrafold:
0.537
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNAsubopt
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAsubopt:
0.336
Sensitivity
CentroidFold:
0.369
RNAsubopt:
0.217
Positive Predictive Value
CentroidFold:
0.600
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNAsubopt
Matthews Correlation Coefficient
Sfold:
0.469
RNAsubopt:
0.336
Sensitivity
Sfold:
0.412
RNAsubopt:
0.217
Positive Predictive Value
Sfold:
0.535
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNAsubopt
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAsubopt:
0.336
Sensitivity
MaxExpect:
0.443
RNAsubopt:
0.217
Positive Predictive Value
MaxExpect:
0.488
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNAsubopt
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAsubopt:
0.336
Sensitivity
ProbKnot:
0.446
RNAsubopt:
0.217
Positive Predictive Value
ProbKnot:
0.475
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNAsubopt
Matthews Correlation Coefficient
UNAFold:
0.445
RNAsubopt:
0.336
Sensitivity
UNAFold:
0.445
RNAsubopt:
0.217
Positive Predictive Value
UNAFold:
0.446
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAsubopt:
0.336
Sensitivity
CentroidAlifold(seed):
0.213
RNAsubopt:
0.217
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNAsubopt
Matthews Correlation Coefficient
Fold:
0.437
RNAsubopt:
0.336
Sensitivity
Fold:
0.432
RNAsubopt:
0.217
Positive Predictive Value
Fold:
0.444
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNAsubopt
Matthews Correlation Coefficient
RNAfold:
0.429
RNAsubopt:
0.336
Sensitivity
RNAfold:
0.422
RNAsubopt:
0.217
Positive Predictive Value
RNAfold:
0.437
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNAsubopt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAsubopt:
0.336
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAsubopt:
0.217
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAsubopt:
0.336
Sensitivity
PETfold_pre2.0(20):
0.211
RNAsubopt:
0.217
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAsubopt:
0.336
Sensitivity
CentroidAlifold(20):
0.190
RNAsubopt:
0.217
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNAsubopt
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAsubopt:
0.336
Sensitivity
PknotsRG:
0.320
RNAsubopt:
0.217
Positive Predictive Value
PknotsRG:
0.465
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAsubopt:
0.336
Sensitivity
MXScarna(seed):
0.196
RNAsubopt:
0.217
Positive Predictive Value
MXScarna(seed):
0.753
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAsubopt:
0.336
Sensitivity
RNAalifold(20):
0.185
RNAsubopt:
0.217
Positive Predictive Value
RNAalifold(20):
0.787
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNAsubopt:
0.336
Sensitivity
RNAalifold(seed):
0.169
RNAsubopt:
0.217
Positive Predictive Value
RNAalifold(seed):
0.835
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RNAsubopt
Matthews Correlation Coefficient
Afold:
0.358
RNAsubopt:
0.336
Sensitivity
Afold:
0.295
RNAsubopt:
0.217
Positive Predictive Value
Afold:
0.434
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNAsubopt:
0.336
Sensitivity
MXScarna(20):
0.181
RNAsubopt:
0.217
Positive Predictive Value
MXScarna(20):
0.710
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RNAsubopt
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNAsubopt:
0.336
Sensitivity
CRWrnafold:
0.275
RNAsubopt:
0.217
Positive Predictive Value
CRWrnafold:
0.435
RNAsubopt:
0.521
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAsubopt vs McQFold
Matthews Correlation Coefficient
RNAsubopt:
0.336
McQFold:
0.314
Sensitivity
RNAsubopt:
0.217
McQFold:
0.205
Positive Predictive Value
RNAsubopt:
0.521
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs TurboFold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
TurboFold(20):
0.301
Sensitivity
RNAsubopt:
0.217
TurboFold(20):
0.108
Positive Predictive Value
RNAsubopt:
0.521
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs PPfold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
PPfold(20):
0.298
Sensitivity
RNAsubopt:
0.217
PPfold(20):
0.101
Positive Predictive Value
RNAsubopt:
0.521
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNAshapes
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNAshapes:
0.295
Sensitivity
RNAsubopt:
0.217
RNAshapes:
0.180
Positive Predictive Value
RNAsubopt:
0.521
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Carnac(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Carnac(20):
0.288
Sensitivity
RNAsubopt:
0.217
Carnac(20):
0.093
Positive Predictive Value
RNAsubopt:
0.521
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNASLOpt
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNASLOpt:
0.281
Sensitivity
RNAsubopt:
0.217
RNASLOpt:
0.158
Positive Predictive Value
RNAsubopt:
0.521
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RSpredict(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RSpredict(20):
0.270
Sensitivity
RNAsubopt:
0.217
RSpredict(20):
0.124
Positive Predictive Value
RNAsubopt:
0.521
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Multilign(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Multilign(20):
0.251
Sensitivity
RNAsubopt:
0.217
Multilign(20):
0.083
Positive Predictive Value
RNAsubopt:
0.521
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNAwolf
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNAwolf:
0.251
Sensitivity
RNAsubopt:
0.217
RNAwolf:
0.191
Positive Predictive Value
RNAsubopt:
0.521
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Murlet(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Murlet(20):
0.230
Sensitivity
RNAsubopt:
0.217
Murlet(20):
0.067
Positive Predictive Value
RNAsubopt:
0.521
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Vsfold4
Matthews Correlation Coefficient
RNAsubopt:
0.336
Vsfold4:
0.217
Sensitivity
RNAsubopt:
0.217
Vsfold4:
0.125
Positive Predictive Value
RNAsubopt:
0.521
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Vsfold5
Matthews Correlation Coefficient
RNAsubopt:
0.336
Vsfold5:
0.211
Sensitivity
RNAsubopt:
0.217
Vsfold5:
0.125
Positive Predictive Value
RNAsubopt:
0.521
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RSpredict(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RSpredict(seed):
0.212
Sensitivity
RNAsubopt:
0.217
RSpredict(seed):
0.075
Positive Predictive Value
RNAsubopt:
0.521
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs CMfinder(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
CMfinder(20):
0.210
Sensitivity
RNAsubopt:
0.217
CMfinder(20):
0.055
Positive Predictive Value
RNAsubopt:
0.521
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs HotKnots
Matthews Correlation Coefficient
RNAsubopt:
0.336
HotKnots:
0.179
Sensitivity
RNAsubopt:
0.217
HotKnots:
0.051
Positive Predictive Value
RNAsubopt:
0.521
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNASampler(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNASampler(20):
0.176
Sensitivity
RNAsubopt:
0.217
RNASampler(20):
0.035
Positive Predictive Value
RNAsubopt:
0.521
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Cylofold
Matthews Correlation Coefficient
RNAsubopt:
0.336
Cylofold:
0.155
Sensitivity
RNAsubopt:
0.217
Cylofold:
0.040
Positive Predictive Value
RNAsubopt:
0.521
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Pknots
Matthews Correlation Coefficient
RNAsubopt:
0.336
Pknots:
0.143
Sensitivity
RNAsubopt:
0.217
Pknots:
0.033
Positive Predictive Value
RNAsubopt:
0.521
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Mastr(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Mastr(20):
0.134
Sensitivity
RNAsubopt:
0.217
Mastr(20):
0.023
Positive Predictive Value
RNAsubopt:
0.521
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Alterna
Matthews Correlation Coefficient
RNAsubopt:
0.336
Alterna:
0.129
Sensitivity
RNAsubopt:
0.217
Alterna:
0.030
Positive Predictive Value
RNAsubopt:
0.521
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs MCFold
Matthews Correlation Coefficient
RNAsubopt:
0.336
MCFold:
0.125
Sensitivity
RNAsubopt:
0.217
MCFold:
0.036
Positive Predictive Value
RNAsubopt:
0.521
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RDfolder
Matthews Correlation Coefficient
RNAsubopt:
0.336
RDfolder:
0.122
Sensitivity
RNAsubopt:
0.217
RDfolder:
0.025
Positive Predictive Value
RNAsubopt:
0.521
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Murlet(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Murlet(seed):
0.079
Sensitivity
RNAsubopt:
0.217
Murlet(seed):
0.008
Positive Predictive Value
RNAsubopt:
0.521
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs CMfinder(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
CMfinder(seed):
0.080
Sensitivity
RNAsubopt:
0.217
CMfinder(seed):
0.008
Positive Predictive Value
RNAsubopt:
0.521
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs TurboFold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
TurboFold(seed):
0.076
Sensitivity
RNAsubopt:
0.217
TurboFold(seed):
0.007
Positive Predictive Value
RNAsubopt:
0.521
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs PPfold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
PPfold(seed):
0.069
Sensitivity
RNAsubopt:
0.217
PPfold(seed):
0.006
Positive Predictive Value
RNAsubopt:
0.521
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Carnac(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Carnac(seed):
0.066
Sensitivity
RNAsubopt:
0.217
Carnac(seed):
0.005
Positive Predictive Value
RNAsubopt:
0.521
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNASampler(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNASampler(seed):
0.063
Sensitivity
RNAsubopt:
0.217
RNASampler(seed):
0.005
Positive Predictive Value
RNAsubopt:
0.521
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs NanoFolder
Matthews Correlation Coefficient
RNAsubopt:
0.336
NanoFolder:
0.052
Sensitivity
RNAsubopt:
0.217
NanoFolder:
0.016
Positive Predictive Value
RNAsubopt:
0.521
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Mastr(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Mastr(seed):
0.051
Sensitivity
RNAsubopt:
0.217
Mastr(seed):
0.003
Positive Predictive Value
RNAsubopt:
0.521
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Multilign(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Multilign(seed):
0.043
Sensitivity
RNAsubopt:
0.217
Multilign(seed):
0.003
Positive Predictive Value
RNAsubopt:
0.521
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| McQFold |
1987
ContextFold vs McQFold
Matthews Correlation Coefficient
ContextFold:
0.680
McQFold:
0.314
Sensitivity
ContextFold:
0.644
McQFold:
0.205
Positive Predictive Value
ContextFold:
0.718
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs McQFold
Matthews Correlation Coefficient
IPknot:
0.548
McQFold:
0.314
Sensitivity
IPknot:
0.475
McQFold:
0.205
Positive Predictive Value
IPknot:
0.632
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
McQFold:
0.314
Sensitivity
PETfold_pre2.0(seed):
0.343
McQFold:
0.205
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs McQFold
Matthews Correlation Coefficient
Contrafold:
0.508
McQFold:
0.314
Sensitivity
Contrafold:
0.481
McQFold:
0.205
Positive Predictive Value
Contrafold:
0.537
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs McQFold
Matthews Correlation Coefficient
CentroidFold:
0.470
McQFold:
0.314
Sensitivity
CentroidFold:
0.369
McQFold:
0.205
Positive Predictive Value
CentroidFold:
0.600
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs McQFold
Matthews Correlation Coefficient
Sfold:
0.469
McQFold:
0.314
Sensitivity
Sfold:
0.412
McQFold:
0.205
Positive Predictive Value
Sfold:
0.535
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs McQFold
Matthews Correlation Coefficient
MaxExpect:
0.465
McQFold:
0.314
Sensitivity
MaxExpect:
0.443
McQFold:
0.205
Positive Predictive Value
MaxExpect:
0.488
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs McQFold
Matthews Correlation Coefficient
ProbKnot:
0.460
McQFold:
0.314
Sensitivity
ProbKnot:
0.446
McQFold:
0.205
Positive Predictive Value
ProbKnot:
0.475
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs McQFold
Matthews Correlation Coefficient
UNAFold:
0.445
McQFold:
0.314
Sensitivity
UNAFold:
0.445
McQFold:
0.205
Positive Predictive Value
UNAFold:
0.446
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
McQFold:
0.314
Sensitivity
CentroidAlifold(seed):
0.213
McQFold:
0.205
Positive Predictive Value
CentroidAlifold(seed):
0.901
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs McQFold
Matthews Correlation Coefficient
Fold:
0.437
McQFold:
0.314
Sensitivity
Fold:
0.432
McQFold:
0.205
Positive Predictive Value
Fold:
0.444
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs McQFold
Matthews Correlation Coefficient
RNAfold:
0.429
McQFold:
0.314
Sensitivity
RNAfold:
0.422
McQFold:
0.205
Positive Predictive Value
RNAfold:
0.437
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs McQFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
McQFold:
0.314
Sensitivity
CentroidHomfold‑LAST:
0.216
McQFold:
0.205
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
McQFold:
0.314
Sensitivity
PETfold_pre2.0(20):
0.211
McQFold:
0.205
Positive Predictive Value
PETfold_pre2.0(20):
0.807
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
McQFold:
0.314
Sensitivity
CentroidAlifold(20):
0.190
McQFold:
0.205
Positive Predictive Value
CentroidAlifold(20):
0.891
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs McQFold
Matthews Correlation Coefficient
PknotsRG:
0.386
McQFold:
0.314
Sensitivity
PknotsRG:
0.320
McQFold:
0.205
Positive Predictive Value
PknotsRG:
0.465
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs McQFold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
McQFold:
0.314
Sensitivity
MXScarna(seed):
0.196
McQFold:
0.205
Positive Predictive Value
MXScarna(seed):
0.753
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs McQFold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
McQFold:
0.314
Sensitivity
RNAalifold(20):
0.185
McQFold:
0.205
Positive Predictive Value
RNAalifold(20):
0.787
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs McQFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
McQFold:
0.314
Sensitivity
RNAalifold(seed):
0.169
McQFold:
0.205
Positive Predictive Value
RNAalifold(seed):
0.835
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs McQFold
Matthews Correlation Coefficient
Afold:
0.358
McQFold:
0.314
Sensitivity
Afold:
0.295
McQFold:
0.205
Positive Predictive Value
Afold:
0.434
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs McQFold
Matthews Correlation Coefficient
MXScarna(20):
0.359
McQFold:
0.314
Sensitivity
MXScarna(20):
0.181
McQFold:
0.205
Positive Predictive Value
MXScarna(20):
0.710
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs McQFold
Matthews Correlation Coefficient
CRWrnafold:
0.346
McQFold:
0.314
Sensitivity
CRWrnafold:
0.275
McQFold:
0.205
Positive Predictive Value
CRWrnafold:
0.435
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs McQFold
Matthews Correlation Coefficient
RNAsubopt:
0.336
McQFold:
0.314
Sensitivity
RNAsubopt:
0.217
McQFold:
0.205
Positive Predictive Value
RNAsubopt:
0.521
McQFold:
0.480
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
McQFold vs TurboFold(20)
Matthews Correlation Coefficient
McQFold:
0.314
TurboFold(20):
0.301
Sensitivity
McQFold:
0.205
TurboFold(20):
0.108
Positive Predictive Value
McQFold:
0.480
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs PPfold(20)
Matthews Correlation Coefficient
McQFold:
0.314
PPfold(20):
0.298
Sensitivity
McQFold:
0.205
PPfold(20):
0.101
Positive Predictive Value
McQFold:
0.480
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNAshapes
Matthews Correlation Coefficient
McQFold:
0.314
RNAshapes:
0.295
Sensitivity
McQFold:
0.205
RNAshapes:
0.180
Positive Predictive Value
McQFold:
0.480
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Carnac(20)
Matthews Correlation Coefficient
McQFold:
0.314
Carnac(20):
0.288
Sensitivity
McQFold:
0.205
Carnac(20):
0.093
Positive Predictive Value
McQFold:
0.480
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNASLOpt
Matthews Correlation Coefficient
McQFold:
0.314
RNASLOpt:
0.281
Sensitivity
McQFold:
0.205
RNASLOpt:
0.158
Positive Predictive Value
McQFold:
0.480
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RSpredict(20)
Matthews Correlation Coefficient
McQFold:
0.314
RSpredict(20):
0.270
Sensitivity
McQFold:
0.205
RSpredict(20):
0.124
Positive Predictive Value
McQFold:
0.480
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Multilign(20)
Matthews Correlation Coefficient
McQFold:
0.314
Multilign(20):
0.251
Sensitivity
McQFold:
0.205
Multilign(20):
0.083
Positive Predictive Value
McQFold:
0.480
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNAwolf
Matthews Correlation Coefficient
McQFold:
0.314
RNAwolf:
0.251
Sensitivity
McQFold:
0.205
RNAwolf:
0.191
Positive Predictive Value
McQFold:
0.480
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Murlet(20)
Matthews Correlation Coefficient
McQFold:
0.314
Murlet(20):
0.230
Sensitivity
McQFold:
0.205
Murlet(20):
0.067
Positive Predictive Value
McQFold:
0.480
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Vsfold4
Matthews Correlation Coefficient
McQFold:
0.314
Vsfold4:
0.217
Sensitivity
McQFold:
0.205
Vsfold4:
0.125
Positive Predictive Value
McQFold:
0.480
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Vsfold5
Matthews Correlation Coefficient
McQFold:
0.314
Vsfold5:
0.211
Sensitivity
McQFold:
0.205
Vsfold5:
0.125
Positive Predictive Value
McQFold:
0.480
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RSpredict(seed)
Matthews Correlation Coefficient
McQFold:
0.314
RSpredict(seed):
0.212
Sensitivity
McQFold:
0.205
RSpredict(seed):
0.075
Positive Predictive Value
McQFold:
0.480
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs CMfinder(20)
Matthews Correlation Coefficient
McQFold:
0.314
CMfinder(20):
0.210
Sensitivity
McQFold:
0.205
CMfinder(20):
0.055
Positive Predictive Value
McQFold:
0.480
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs HotKnots
Matthews Correlation Coefficient
McQFold:
0.314
HotKnots:
0.179
Sensitivity
McQFold:
0.205
HotKnots:
0.051
Positive Predictive Value
McQFold:
0.480
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNASampler(20)
Matthews Correlation Coefficient
McQFold:
0.314
RNASampler(20):
0.176
Sensitivity
McQFold:
0.205
RNASampler(20):
0.035
Positive Predictive Value
McQFold:
0.480
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Cylofold
Matthews Correlation Coefficient
McQFold:
0.314
Cylofold:
0.155
Sensitivity
McQFold:
0.205
Cylofold:
0.040
Positive Predictive Value
McQFold:
0.480
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Pknots
Matthews Correlation Coefficient
McQFold:
0.314
Pknots:
0.143
Sensitivity
McQFold:
0.205
Pknots:
0.033
Positive Predictive Value
McQFold:
0.480
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Mastr(20)
Matthews Correlation Coefficient
McQFold:
0.314
Mastr(20):
0.134
Sensitivity
McQFold:
0.205
Mastr(20):
0.023
Positive Predictive Value
McQFold:
0.480
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Alterna
Matthews Correlation Coefficient
McQFold:
0.314
Alterna:
0.129
Sensitivity
McQFold:
0.205
Alterna:
0.030
Positive Predictive Value
McQFold:
0.480
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs MCFold
Matthews Correlation Coefficient
McQFold:
0.314
MCFold:
0.125
Sensitivity
McQFold:
0.205
MCFold:
0.036
Positive Predictive Value
McQFold:
0.480
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RDfolder
Matthews Correlation Coefficient
McQFold:
0.314
RDfolder:
0.122
Sensitivity
McQFold:
0.205
RDfolder:
0.025
Positive Predictive Value
McQFold:
0.480
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Murlet(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Murlet(seed):
0.079
Sensitivity
McQFold:
0.205
Murlet(seed):
0.008
Positive Predictive Value
McQFold:
0.480
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs CMfinder(seed)
Matthews Correlation Coefficient
McQFold:
0.314
CMfinder(seed):
0.080
Sensitivity
McQFold:
0.205
CMfinder(seed):
0.008
Positive Predictive Value
McQFold:
0.480
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs TurboFold(seed)
Matthews Correlation Coefficient
McQFold:
0.314
TurboFold(seed):
0.076
Sensitivity
McQFold:
0.205
TurboFold(seed):
0.007
Positive Predictive Value
McQFold:
0.480
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs PPfold(seed)
Matthews Correlation Coefficient
McQFold:
0.314
PPfold(seed):
0.069
Sensitivity
McQFold:
0.205
PPfold(seed):
0.006
Positive Predictive Value
McQFold:
0.480
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Carnac(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Carnac(seed):
0.066
Sensitivity
McQFold:
0.205
Carnac(seed):
0.005
Positive Predictive Value
McQFold:
0.480
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNASampler(seed)
Matthews Correlation Coefficient
McQFold:
0.314
RNASampler(seed):
0.063
Sensitivity
McQFold:
0.205
RNASampler(seed):
0.005
Positive Predictive Value
McQFold:
0.480
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs NanoFolder
Matthews Correlation Coefficient
McQFold:
0.314
NanoFolder:
0.052
Sensitivity
McQFold:
0.205
NanoFolder:
0.016
Positive Predictive Value
McQFold:
0.480
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Mastr(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Mastr(seed):
0.051
Sensitivity
McQFold:
0.205
Mastr(seed):
0.003
Positive Predictive Value
McQFold:
0.480
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Multilign(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Multilign(seed):
0.043
Sensitivity
McQFold:
0.205
Multilign(seed):
0.003
Positive Predictive Value
McQFold:
0.480
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| TurboFold(20) |
1987
ContextFold vs TurboFold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
TurboFold(20):
0.301
Sensitivity
ContextFold:
0.644
TurboFold(20):
0.108
Positive Predictive Value
ContextFold:
0.718
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs TurboFold(20)
Matthews Correlation Coefficient
IPknot:
0.548
TurboFold(20):
0.301
Sensitivity
IPknot:
0.475
TurboFold(20):
0.108
Positive Predictive Value
IPknot:
0.632
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
TurboFold(20):
0.301
Sensitivity
PETfold_pre2.0(seed):
0.343
TurboFold(20):
0.108
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs TurboFold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
TurboFold(20):
0.301
Sensitivity
Contrafold:
0.481
TurboFold(20):
0.108
Positive Predictive Value
Contrafold:
0.537
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs TurboFold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
TurboFold(20):
0.301
Sensitivity
CentroidFold:
0.369
TurboFold(20):
0.108
Positive Predictive Value
CentroidFold:
0.600
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs TurboFold(20)
Matthews Correlation Coefficient
Sfold:
0.469
TurboFold(20):
0.301
Sensitivity
Sfold:
0.412
TurboFold(20):
0.108
Positive Predictive Value
Sfold:
0.535
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs TurboFold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
TurboFold(20):
0.301
Sensitivity
MaxExpect:
0.443
TurboFold(20):
0.108
Positive Predictive Value
MaxExpect:
0.488
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs TurboFold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
TurboFold(20):
0.301
Sensitivity
ProbKnot:
0.446
TurboFold(20):
0.108
Positive Predictive Value
ProbKnot:
0.475
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs TurboFold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
TurboFold(20):
0.301
Sensitivity
UNAFold:
0.445
TurboFold(20):
0.108
Positive Predictive Value
UNAFold:
0.446
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
TurboFold(20):
0.301
Sensitivity
CentroidAlifold(seed):
0.213
TurboFold(20):
0.108
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs TurboFold(20)
Matthews Correlation Coefficient
Fold:
0.437
TurboFold(20):
0.301
Sensitivity
Fold:
0.432
TurboFold(20):
0.108
Positive Predictive Value
Fold:
0.444
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs TurboFold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
TurboFold(20):
0.301
Sensitivity
RNAfold:
0.422
TurboFold(20):
0.108
Positive Predictive Value
RNAfold:
0.437
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs TurboFold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
TurboFold(20):
0.301
Sensitivity
CentroidHomfold‑LAST:
0.216
TurboFold(20):
0.108
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
TurboFold(20):
0.301
Sensitivity
PETfold_pre2.0(20):
0.211
TurboFold(20):
0.108
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
TurboFold(20):
0.301
Sensitivity
CentroidAlifold(20):
0.190
TurboFold(20):
0.108
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs TurboFold(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
TurboFold(20):
0.301
Sensitivity
PknotsRG:
0.320
TurboFold(20):
0.108
Positive Predictive Value
PknotsRG:
0.465
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
TurboFold(20):
0.301
Sensitivity
MXScarna(seed):
0.196
TurboFold(20):
0.108
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
TurboFold(20):
0.301
Sensitivity
RNAalifold(20):
0.185
TurboFold(20):
0.108
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
TurboFold(20):
0.301
Sensitivity
RNAalifold(seed):
0.169
TurboFold(20):
0.108
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs TurboFold(20)
Matthews Correlation Coefficient
Afold:
0.358
TurboFold(20):
0.301
Sensitivity
Afold:
0.295
TurboFold(20):
0.108
Positive Predictive Value
Afold:
0.434
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
TurboFold(20):
0.301
Sensitivity
MXScarna(20):
0.181
TurboFold(20):
0.108
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs TurboFold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
TurboFold(20):
0.301
Sensitivity
CRWrnafold:
0.275
TurboFold(20):
0.108
Positive Predictive Value
CRWrnafold:
0.435
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs TurboFold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
TurboFold(20):
0.301
Sensitivity
RNAsubopt:
0.217
TurboFold(20):
0.108
Positive Predictive Value
RNAsubopt:
0.521
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs TurboFold(20)
Matthews Correlation Coefficient
McQFold:
0.314
TurboFold(20):
0.301
Sensitivity
McQFold:
0.205
TurboFold(20):
0.108
Positive Predictive Value
McQFold:
0.480
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
TurboFold(20) vs PPfold(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
PPfold(20):
0.298
Sensitivity
TurboFold(20):
0.108
PPfold(20):
0.101
Positive Predictive Value
TurboFold(20):
0.837
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
TurboFold(20) vs RNAshapes
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNAshapes:
0.295
Sensitivity
TurboFold(20):
0.108
RNAshapes:
0.180
Positive Predictive Value
TurboFold(20):
0.837
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
TurboFold(20) vs Carnac(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Carnac(20):
0.288
Sensitivity
TurboFold(20):
0.108
Carnac(20):
0.093
Positive Predictive Value
TurboFold(20):
0.837
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNASLOpt
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNASLOpt:
0.281
Sensitivity
TurboFold(20):
0.108
RNASLOpt:
0.158
Positive Predictive Value
TurboFold(20):
0.837
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RSpredict(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RSpredict(20):
0.270
Sensitivity
TurboFold(20):
0.108
RSpredict(20):
0.124
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Multilign(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Multilign(20):
0.251
Sensitivity
TurboFold(20):
0.108
Multilign(20):
0.083
Positive Predictive Value
TurboFold(20):
0.837
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNAwolf
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNAwolf:
0.251
Sensitivity
TurboFold(20):
0.108
RNAwolf:
0.191
Positive Predictive Value
TurboFold(20):
0.837
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Murlet(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Murlet(20):
0.230
Sensitivity
TurboFold(20):
0.108
Murlet(20):
0.067
Positive Predictive Value
TurboFold(20):
0.837
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Vsfold4
Matthews Correlation Coefficient
TurboFold(20):
0.301
Vsfold4:
0.217
Sensitivity
TurboFold(20):
0.108
Vsfold4:
0.125
Positive Predictive Value
TurboFold(20):
0.837
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Vsfold5
Matthews Correlation Coefficient
TurboFold(20):
0.301
Vsfold5:
0.211
Sensitivity
TurboFold(20):
0.108
Vsfold5:
0.125
Positive Predictive Value
TurboFold(20):
0.837
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RSpredict(seed):
0.212
Sensitivity
TurboFold(20):
0.108
RSpredict(seed):
0.075
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs CMfinder(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
CMfinder(20):
0.210
Sensitivity
TurboFold(20):
0.108
CMfinder(20):
0.055
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs HotKnots
Matthews Correlation Coefficient
TurboFold(20):
0.301
HotKnots:
0.179
Sensitivity
TurboFold(20):
0.108
HotKnots:
0.051
Positive Predictive Value
TurboFold(20):
0.837
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNASampler(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNASampler(20):
0.176
Sensitivity
TurboFold(20):
0.108
RNASampler(20):
0.035
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Cylofold
Matthews Correlation Coefficient
TurboFold(20):
0.301
Cylofold:
0.155
Sensitivity
TurboFold(20):
0.108
Cylofold:
0.040
Positive Predictive Value
TurboFold(20):
0.837
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Pknots
Matthews Correlation Coefficient
TurboFold(20):
0.301
Pknots:
0.143
Sensitivity
TurboFold(20):
0.108
Pknots:
0.033
Positive Predictive Value
TurboFold(20):
0.837
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Mastr(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Mastr(20):
0.134
Sensitivity
TurboFold(20):
0.108
Mastr(20):
0.023
Positive Predictive Value
TurboFold(20):
0.837
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Alterna
Matthews Correlation Coefficient
TurboFold(20):
0.301
Alterna:
0.129
Sensitivity
TurboFold(20):
0.108
Alterna:
0.030
Positive Predictive Value
TurboFold(20):
0.837
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs MCFold
Matthews Correlation Coefficient
TurboFold(20):
0.301
MCFold:
0.125
Sensitivity
TurboFold(20):
0.108
MCFold:
0.036
Positive Predictive Value
TurboFold(20):
0.837
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RDfolder
Matthews Correlation Coefficient
TurboFold(20):
0.301
RDfolder:
0.122
Sensitivity
TurboFold(20):
0.108
RDfolder:
0.025
Positive Predictive Value
TurboFold(20):
0.837
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Murlet(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Murlet(seed):
0.079
Sensitivity
TurboFold(20):
0.108
Murlet(seed):
0.008
Positive Predictive Value
TurboFold(20):
0.837
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
CMfinder(seed):
0.080
Sensitivity
TurboFold(20):
0.108
CMfinder(seed):
0.008
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
TurboFold(seed):
0.076
Sensitivity
TurboFold(20):
0.108
TurboFold(seed):
0.007
Positive Predictive Value
TurboFold(20):
0.837
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
PPfold(seed):
0.069
Sensitivity
TurboFold(20):
0.108
PPfold(seed):
0.006
Positive Predictive Value
TurboFold(20):
0.837
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Carnac(seed):
0.066
Sensitivity
TurboFold(20):
0.108
Carnac(seed):
0.005
Positive Predictive Value
TurboFold(20):
0.837
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNASampler(seed):
0.063
Sensitivity
TurboFold(20):
0.108
RNASampler(seed):
0.005
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(20):
0.301
NanoFolder:
0.052
Sensitivity
TurboFold(20):
0.108
NanoFolder:
0.016
Positive Predictive Value
TurboFold(20):
0.837
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Mastr(seed):
0.051
Sensitivity
TurboFold(20):
0.108
Mastr(seed):
0.003
Positive Predictive Value
TurboFold(20):
0.837
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Multilign(seed):
0.043
Sensitivity
TurboFold(20):
0.108
Multilign(seed):
0.003
Positive Predictive Value
TurboFold(20):
0.837
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PPfold(20) |
1987
ContextFold vs PPfold(20)
Matthews Correlation Coefficient
ContextFold:
0.680
PPfold(20):
0.298
Sensitivity
ContextFold:
0.644
PPfold(20):
0.101
Positive Predictive Value
ContextFold:
0.718
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs PPfold(20)
Matthews Correlation Coefficient
IPknot:
0.548
PPfold(20):
0.298
Sensitivity
IPknot:
0.475
PPfold(20):
0.101
Positive Predictive Value
IPknot:
0.632
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PPfold(20):
0.298
Sensitivity
PETfold_pre2.0(seed):
0.343
PPfold(20):
0.101
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs PPfold(20)
Matthews Correlation Coefficient
Contrafold:
0.508
PPfold(20):
0.298
Sensitivity
Contrafold:
0.481
PPfold(20):
0.101
Positive Predictive Value
Contrafold:
0.537
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs PPfold(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
PPfold(20):
0.298
Sensitivity
CentroidFold:
0.369
PPfold(20):
0.101
Positive Predictive Value
CentroidFold:
0.600
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs PPfold(20)
Matthews Correlation Coefficient
Sfold:
0.469
PPfold(20):
0.298
Sensitivity
Sfold:
0.412
PPfold(20):
0.101
Positive Predictive Value
Sfold:
0.535
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs PPfold(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
PPfold(20):
0.298
Sensitivity
MaxExpect:
0.443
PPfold(20):
0.101
Positive Predictive Value
MaxExpect:
0.488
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs PPfold(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
PPfold(20):
0.298
Sensitivity
ProbKnot:
0.446
PPfold(20):
0.101
Positive Predictive Value
ProbKnot:
0.475
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs PPfold(20)
Matthews Correlation Coefficient
UNAFold:
0.445
PPfold(20):
0.298
Sensitivity
UNAFold:
0.445
PPfold(20):
0.101
Positive Predictive Value
UNAFold:
0.446
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PPfold(20):
0.298
Sensitivity
CentroidAlifold(seed):
0.213
PPfold(20):
0.101
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs PPfold(20)
Matthews Correlation Coefficient
Fold:
0.437
PPfold(20):
0.298
Sensitivity
Fold:
0.432
PPfold(20):
0.101
Positive Predictive Value
Fold:
0.444
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs PPfold(20)
Matthews Correlation Coefficient
RNAfold:
0.429
PPfold(20):
0.298
Sensitivity
RNAfold:
0.422
PPfold(20):
0.101
Positive Predictive Value
RNAfold:
0.437
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs PPfold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PPfold(20):
0.298
Sensitivity
CentroidHomfold‑LAST:
0.216
PPfold(20):
0.101
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
PPfold(20):
0.298
Sensitivity
PETfold_pre2.0(20):
0.211
PPfold(20):
0.101
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
PPfold(20):
0.298
Sensitivity
CentroidAlifold(20):
0.190
PPfold(20):
0.101
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs PPfold(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
PPfold(20):
0.298
Sensitivity
PknotsRG:
0.320
PPfold(20):
0.101
Positive Predictive Value
PknotsRG:
0.465
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
PPfold(20):
0.298
Sensitivity
MXScarna(seed):
0.196
PPfold(20):
0.101
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
PPfold(20):
0.298
Sensitivity
RNAalifold(20):
0.185
PPfold(20):
0.101
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
PPfold(20):
0.298
Sensitivity
RNAalifold(seed):
0.169
PPfold(20):
0.101
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs PPfold(20)
Matthews Correlation Coefficient
Afold:
0.358
PPfold(20):
0.298
Sensitivity
Afold:
0.295
PPfold(20):
0.101
Positive Predictive Value
Afold:
0.434
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
PPfold(20):
0.298
Sensitivity
MXScarna(20):
0.181
PPfold(20):
0.101
Positive Predictive Value
MXScarna(20):
0.710
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs PPfold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
PPfold(20):
0.298
Sensitivity
CRWrnafold:
0.275
PPfold(20):
0.101
Positive Predictive Value
CRWrnafold:
0.435
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs PPfold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
PPfold(20):
0.298
Sensitivity
RNAsubopt:
0.217
PPfold(20):
0.101
Positive Predictive Value
RNAsubopt:
0.521
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs PPfold(20)
Matthews Correlation Coefficient
McQFold:
0.314
PPfold(20):
0.298
Sensitivity
McQFold:
0.205
PPfold(20):
0.101
Positive Predictive Value
McQFold:
0.480
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs PPfold(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
PPfold(20):
0.298
Sensitivity
TurboFold(20):
0.108
PPfold(20):
0.101
Positive Predictive Value
TurboFold(20):
0.837
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
|
+
PPfold(20) vs RNAshapes
Matthews Correlation Coefficient
PPfold(20):
0.298
RNAshapes:
0.295
Sensitivity
PPfold(20):
0.101
RNAshapes:
0.180
Positive Predictive Value
PPfold(20):
0.879
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 2.82781450476e-07
|
+
PPfold(20) vs Carnac(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Carnac(20):
0.288
Sensitivity
PPfold(20):
0.101
Carnac(20):
0.093
Positive Predictive Value
PPfold(20):
0.879
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNASLOpt
Matthews Correlation Coefficient
PPfold(20):
0.298
RNASLOpt:
0.281
Sensitivity
PPfold(20):
0.101
RNASLOpt:
0.158
Positive Predictive Value
PPfold(20):
0.879
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RSpredict(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
RSpredict(20):
0.270
Sensitivity
PPfold(20):
0.101
RSpredict(20):
0.124
Positive Predictive Value
PPfold(20):
0.879
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Multilign(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Multilign(20):
0.251
Sensitivity
PPfold(20):
0.101
Multilign(20):
0.083
Positive Predictive Value
PPfold(20):
0.879
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNAwolf
Matthews Correlation Coefficient
PPfold(20):
0.298
RNAwolf:
0.251
Sensitivity
PPfold(20):
0.101
RNAwolf:
0.191
Positive Predictive Value
PPfold(20):
0.879
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Murlet(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Murlet(20):
0.230
Sensitivity
PPfold(20):
0.101
Murlet(20):
0.067
Positive Predictive Value
PPfold(20):
0.879
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Vsfold4
Matthews Correlation Coefficient
PPfold(20):
0.298
Vsfold4:
0.217
Sensitivity
PPfold(20):
0.101
Vsfold4:
0.125
Positive Predictive Value
PPfold(20):
0.879
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Vsfold5
Matthews Correlation Coefficient
PPfold(20):
0.298
Vsfold5:
0.211
Sensitivity
PPfold(20):
0.101
Vsfold5:
0.125
Positive Predictive Value
PPfold(20):
0.879
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
RSpredict(seed):
0.212
Sensitivity
PPfold(20):
0.101
RSpredict(seed):
0.075
Positive Predictive Value
PPfold(20):
0.879
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs CMfinder(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
CMfinder(20):
0.210
Sensitivity
PPfold(20):
0.101
CMfinder(20):
0.055
Positive Predictive Value
PPfold(20):
0.879
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs HotKnots
Matthews Correlation Coefficient
PPfold(20):
0.298
HotKnots:
0.179
Sensitivity
PPfold(20):
0.101
HotKnots:
0.051
Positive Predictive Value
PPfold(20):
0.879
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNASampler(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
RNASampler(20):
0.176
Sensitivity
PPfold(20):
0.101
RNASampler(20):
0.035
Positive Predictive Value
PPfold(20):
0.879
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Cylofold
Matthews Correlation Coefficient
PPfold(20):
0.298
Cylofold:
0.155
Sensitivity
PPfold(20):
0.101
Cylofold:
0.040
Positive Predictive Value
PPfold(20):
0.879
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Pknots
Matthews Correlation Coefficient
PPfold(20):
0.298
Pknots:
0.143
Sensitivity
PPfold(20):
0.101
Pknots:
0.033
Positive Predictive Value
PPfold(20):
0.879
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Mastr(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Mastr(20):
0.134
Sensitivity
PPfold(20):
0.101
Mastr(20):
0.023
Positive Predictive Value
PPfold(20):
0.879
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Alterna
Matthews Correlation Coefficient
PPfold(20):
0.298
Alterna:
0.129
Sensitivity
PPfold(20):
0.101
Alterna:
0.030
Positive Predictive Value
PPfold(20):
0.879
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs MCFold
Matthews Correlation Coefficient
PPfold(20):
0.298
MCFold:
0.125
Sensitivity
PPfold(20):
0.101
MCFold:
0.036
Positive Predictive Value
PPfold(20):
0.879
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RDfolder
Matthews Correlation Coefficient
PPfold(20):
0.298
RDfolder:
0.122
Sensitivity
PPfold(20):
0.101
RDfolder:
0.025
Positive Predictive Value
PPfold(20):
0.879
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Murlet(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Murlet(seed):
0.079
Sensitivity
PPfold(20):
0.101
Murlet(seed):
0.008
Positive Predictive Value
PPfold(20):
0.879
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
CMfinder(seed):
0.080
Sensitivity
PPfold(20):
0.101
CMfinder(seed):
0.008
Positive Predictive Value
PPfold(20):
0.879
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
TurboFold(seed):
0.076
Sensitivity
PPfold(20):
0.101
TurboFold(seed):
0.007
Positive Predictive Value
PPfold(20):
0.879
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs PPfold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
PPfold(seed):
0.069
Sensitivity
PPfold(20):
0.101
PPfold(seed):
0.006
Positive Predictive Value
PPfold(20):
0.879
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Carnac(seed):
0.066
Sensitivity
PPfold(20):
0.101
Carnac(seed):
0.005
Positive Predictive Value
PPfold(20):
0.879
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
RNASampler(seed):
0.063
Sensitivity
PPfold(20):
0.101
RNASampler(seed):
0.005
Positive Predictive Value
PPfold(20):
0.879
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs NanoFolder
Matthews Correlation Coefficient
PPfold(20):
0.298
NanoFolder:
0.052
Sensitivity
PPfold(20):
0.101
NanoFolder:
0.016
Positive Predictive Value
PPfold(20):
0.879
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Mastr(seed):
0.051
Sensitivity
PPfold(20):
0.101
Mastr(seed):
0.003
Positive Predictive Value
PPfold(20):
0.879
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Multilign(seed):
0.043
Sensitivity
PPfold(20):
0.101
Multilign(seed):
0.003
Positive Predictive Value
PPfold(20):
0.879
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAshapes |
1987
ContextFold vs RNAshapes
Matthews Correlation Coefficient
ContextFold:
0.680
RNAshapes:
0.295
Sensitivity
ContextFold:
0.644
RNAshapes:
0.180
Positive Predictive Value
ContextFold:
0.718
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNAshapes
Matthews Correlation Coefficient
IPknot:
0.548
RNAshapes:
0.295
Sensitivity
IPknot:
0.475
RNAshapes:
0.180
Positive Predictive Value
IPknot:
0.632
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAshapes:
0.295
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAshapes:
0.180
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNAshapes
Matthews Correlation Coefficient
Contrafold:
0.508
RNAshapes:
0.295
Sensitivity
Contrafold:
0.481
RNAshapes:
0.180
Positive Predictive Value
Contrafold:
0.537
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNAshapes
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAshapes:
0.295
Sensitivity
CentroidFold:
0.369
RNAshapes:
0.180
Positive Predictive Value
CentroidFold:
0.600
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNAshapes
Matthews Correlation Coefficient
Sfold:
0.469
RNAshapes:
0.295
Sensitivity
Sfold:
0.412
RNAshapes:
0.180
Positive Predictive Value
Sfold:
0.535
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNAshapes
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAshapes:
0.295
Sensitivity
MaxExpect:
0.443
RNAshapes:
0.180
Positive Predictive Value
MaxExpect:
0.488
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNAshapes
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAshapes:
0.295
Sensitivity
ProbKnot:
0.446
RNAshapes:
0.180
Positive Predictive Value
ProbKnot:
0.475
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNAshapes
Matthews Correlation Coefficient
UNAFold:
0.445
RNAshapes:
0.295
Sensitivity
UNAFold:
0.445
RNAshapes:
0.180
Positive Predictive Value
UNAFold:
0.446
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAshapes:
0.295
Sensitivity
CentroidAlifold(seed):
0.213
RNAshapes:
0.180
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNAshapes
Matthews Correlation Coefficient
Fold:
0.437
RNAshapes:
0.295
Sensitivity
Fold:
0.432
RNAshapes:
0.180
Positive Predictive Value
Fold:
0.444
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNAshapes
Matthews Correlation Coefficient
RNAfold:
0.429
RNAshapes:
0.295
Sensitivity
RNAfold:
0.422
RNAshapes:
0.180
Positive Predictive Value
RNAfold:
0.437
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNAshapes
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAshapes:
0.295
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAshapes:
0.180
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAshapes:
0.295
Sensitivity
PETfold_pre2.0(20):
0.211
RNAshapes:
0.180
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAshapes:
0.295
Sensitivity
CentroidAlifold(20):
0.190
RNAshapes:
0.180
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNAshapes
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAshapes:
0.295
Sensitivity
PknotsRG:
0.320
RNAshapes:
0.180
Positive Predictive Value
PknotsRG:
0.465
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAshapes:
0.295
Sensitivity
MXScarna(seed):
0.196
RNAshapes:
0.180
Positive Predictive Value
MXScarna(seed):
0.753
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAshapes:
0.295
Sensitivity
RNAalifold(20):
0.185
RNAshapes:
0.180
Positive Predictive Value
RNAalifold(20):
0.787
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNAshapes:
0.295
Sensitivity
RNAalifold(seed):
0.169
RNAshapes:
0.180
Positive Predictive Value
RNAalifold(seed):
0.835
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RNAshapes
Matthews Correlation Coefficient
Afold:
0.358
RNAshapes:
0.295
Sensitivity
Afold:
0.295
RNAshapes:
0.180
Positive Predictive Value
Afold:
0.434
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNAshapes:
0.295
Sensitivity
MXScarna(20):
0.181
RNAshapes:
0.180
Positive Predictive Value
MXScarna(20):
0.710
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RNAshapes
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNAshapes:
0.295
Sensitivity
CRWrnafold:
0.275
RNAshapes:
0.180
Positive Predictive Value
CRWrnafold:
0.435
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RNAshapes
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNAshapes:
0.295
Sensitivity
RNAsubopt:
0.217
RNAshapes:
0.180
Positive Predictive Value
RNAsubopt:
0.521
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RNAshapes
Matthews Correlation Coefficient
McQFold:
0.314
RNAshapes:
0.295
Sensitivity
McQFold:
0.205
RNAshapes:
0.180
Positive Predictive Value
McQFold:
0.480
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RNAshapes
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNAshapes:
0.295
Sensitivity
TurboFold(20):
0.108
RNAshapes:
0.180
Positive Predictive Value
TurboFold(20):
0.837
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
1987
PPfold(20) vs RNAshapes
Matthews Correlation Coefficient
PPfold(20):
0.298
RNAshapes:
0.295
Sensitivity
PPfold(20):
0.101
RNAshapes:
0.180
Positive Predictive Value
PPfold(20):
0.879
RNAshapes:
0.484
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 2.82781450476e-07
|
|
+
RNAshapes vs Carnac(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Carnac(20):
0.288
Sensitivity
RNAshapes:
0.180
Carnac(20):
0.093
Positive Predictive Value
RNAshapes:
0.484
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 4.48494289621e-08
|
+
RNAshapes vs RNASLOpt
Matthews Correlation Coefficient
RNAshapes:
0.295
RNASLOpt:
0.281
Sensitivity
RNAshapes:
0.180
RNASLOpt:
0.158
Positive Predictive Value
RNAshapes:
0.484
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RSpredict(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
RSpredict(20):
0.270
Sensitivity
RNAshapes:
0.180
RSpredict(20):
0.124
Positive Predictive Value
RNAshapes:
0.484
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Multilign(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Multilign(20):
0.251
Sensitivity
RNAshapes:
0.180
Multilign(20):
0.083
Positive Predictive Value
RNAshapes:
0.484
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RNAwolf
Matthews Correlation Coefficient
RNAshapes:
0.295
RNAwolf:
0.251
Sensitivity
RNAshapes:
0.180
RNAwolf:
0.191
Positive Predictive Value
RNAshapes:
0.484
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Murlet(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Murlet(20):
0.230
Sensitivity
RNAshapes:
0.180
Murlet(20):
0.067
Positive Predictive Value
RNAshapes:
0.484
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Vsfold4
Matthews Correlation Coefficient
RNAshapes:
0.295
Vsfold4:
0.217
Sensitivity
RNAshapes:
0.180
Vsfold4:
0.125
Positive Predictive Value
RNAshapes:
0.484
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Vsfold5
Matthews Correlation Coefficient
RNAshapes:
0.295
Vsfold5:
0.211
Sensitivity
RNAshapes:
0.180
Vsfold5:
0.125
Positive Predictive Value
RNAshapes:
0.484
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RSpredict(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
RSpredict(seed):
0.212
Sensitivity
RNAshapes:
0.180
RSpredict(seed):
0.075
Positive Predictive Value
RNAshapes:
0.484
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs CMfinder(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
CMfinder(20):
0.210
Sensitivity
RNAshapes:
0.180
CMfinder(20):
0.055
Positive Predictive Value
RNAshapes:
0.484
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs HotKnots
Matthews Correlation Coefficient
RNAshapes:
0.295
HotKnots:
0.179
Sensitivity
RNAshapes:
0.180
HotKnots:
0.051
Positive Predictive Value
RNAshapes:
0.484
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RNASampler(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
RNASampler(20):
0.176
Sensitivity
RNAshapes:
0.180
RNASampler(20):
0.035
Positive Predictive Value
RNAshapes:
0.484
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Cylofold
Matthews Correlation Coefficient
RNAshapes:
0.295
Cylofold:
0.155
Sensitivity
RNAshapes:
0.180
Cylofold:
0.040
Positive Predictive Value
RNAshapes:
0.484
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Pknots
Matthews Correlation Coefficient
RNAshapes:
0.295
Pknots:
0.143
Sensitivity
RNAshapes:
0.180
Pknots:
0.033
Positive Predictive Value
RNAshapes:
0.484
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Mastr(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Mastr(20):
0.134
Sensitivity
RNAshapes:
0.180
Mastr(20):
0.023
Positive Predictive Value
RNAshapes:
0.484
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Alterna
Matthews Correlation Coefficient
RNAshapes:
0.295
Alterna:
0.129
Sensitivity
RNAshapes:
0.180
Alterna:
0.030
Positive Predictive Value
RNAshapes:
0.484
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs MCFold
Matthews Correlation Coefficient
RNAshapes:
0.295
MCFold:
0.125
Sensitivity
RNAshapes:
0.180
MCFold:
0.036
Positive Predictive Value
RNAshapes:
0.484
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RDfolder
Matthews Correlation Coefficient
RNAshapes:
0.295
RDfolder:
0.122
Sensitivity
RNAshapes:
0.180
RDfolder:
0.025
Positive Predictive Value
RNAshapes:
0.484
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Murlet(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Murlet(seed):
0.079
Sensitivity
RNAshapes:
0.180
Murlet(seed):
0.008
Positive Predictive Value
RNAshapes:
0.484
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs CMfinder(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
CMfinder(seed):
0.080
Sensitivity
RNAshapes:
0.180
CMfinder(seed):
0.008
Positive Predictive Value
RNAshapes:
0.484
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs TurboFold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
TurboFold(seed):
0.076
Sensitivity
RNAshapes:
0.180
TurboFold(seed):
0.007
Positive Predictive Value
RNAshapes:
0.484
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs PPfold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
PPfold(seed):
0.069
Sensitivity
RNAshapes:
0.180
PPfold(seed):
0.006
Positive Predictive Value
RNAshapes:
0.484
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Carnac(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Carnac(seed):
0.066
Sensitivity
RNAshapes:
0.180
Carnac(seed):
0.005
Positive Predictive Value
RNAshapes:
0.484
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RNASampler(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
RNASampler(seed):
0.063
Sensitivity
RNAshapes:
0.180
RNASampler(seed):
0.005
Positive Predictive Value
RNAshapes:
0.484
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs NanoFolder
Matthews Correlation Coefficient
RNAshapes:
0.295
NanoFolder:
0.052
Sensitivity
RNAshapes:
0.180
NanoFolder:
0.016
Positive Predictive Value
RNAshapes:
0.484
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Mastr(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Mastr(seed):
0.051
Sensitivity
RNAshapes:
0.180
Mastr(seed):
0.003
Positive Predictive Value
RNAshapes:
0.484
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Multilign(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Multilign(seed):
0.043
Sensitivity
RNAshapes:
0.180
Multilign(seed):
0.003
Positive Predictive Value
RNAshapes:
0.484
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Carnac(20) |
1987
ContextFold vs Carnac(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Carnac(20):
0.288
Sensitivity
ContextFold:
0.644
Carnac(20):
0.093
Positive Predictive Value
ContextFold:
0.718
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Carnac(20)
Matthews Correlation Coefficient
IPknot:
0.548
Carnac(20):
0.288
Sensitivity
IPknot:
0.475
Carnac(20):
0.093
Positive Predictive Value
IPknot:
0.632
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Carnac(20):
0.288
Sensitivity
PETfold_pre2.0(seed):
0.343
Carnac(20):
0.093
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Carnac(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Carnac(20):
0.288
Sensitivity
Contrafold:
0.481
Carnac(20):
0.093
Positive Predictive Value
Contrafold:
0.537
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Carnac(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Carnac(20):
0.288
Sensitivity
CentroidFold:
0.369
Carnac(20):
0.093
Positive Predictive Value
CentroidFold:
0.600
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Carnac(20)
Matthews Correlation Coefficient
Sfold:
0.469
Carnac(20):
0.288
Sensitivity
Sfold:
0.412
Carnac(20):
0.093
Positive Predictive Value
Sfold:
0.535
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Carnac(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Carnac(20):
0.288
Sensitivity
MaxExpect:
0.443
Carnac(20):
0.093
Positive Predictive Value
MaxExpect:
0.488
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Carnac(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Carnac(20):
0.288
Sensitivity
ProbKnot:
0.446
Carnac(20):
0.093
Positive Predictive Value
ProbKnot:
0.475
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Carnac(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Carnac(20):
0.288
Sensitivity
UNAFold:
0.445
Carnac(20):
0.093
Positive Predictive Value
UNAFold:
0.446
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Carnac(20):
0.288
Sensitivity
CentroidAlifold(seed):
0.213
Carnac(20):
0.093
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Carnac(20)
Matthews Correlation Coefficient
Fold:
0.437
Carnac(20):
0.288
Sensitivity
Fold:
0.432
Carnac(20):
0.093
Positive Predictive Value
Fold:
0.444
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Carnac(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Carnac(20):
0.288
Sensitivity
RNAfold:
0.422
Carnac(20):
0.093
Positive Predictive Value
RNAfold:
0.437
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Carnac(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Carnac(20):
0.288
Sensitivity
CentroidHomfold‑LAST:
0.216
Carnac(20):
0.093
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Carnac(20):
0.288
Sensitivity
PETfold_pre2.0(20):
0.211
Carnac(20):
0.093
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Carnac(20):
0.288
Sensitivity
CentroidAlifold(20):
0.190
Carnac(20):
0.093
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Carnac(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Carnac(20):
0.288
Sensitivity
PknotsRG:
0.320
Carnac(20):
0.093
Positive Predictive Value
PknotsRG:
0.465
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Carnac(20):
0.288
Sensitivity
MXScarna(seed):
0.196
Carnac(20):
0.093
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Carnac(20):
0.288
Sensitivity
RNAalifold(20):
0.185
Carnac(20):
0.093
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Carnac(20):
0.288
Sensitivity
RNAalifold(seed):
0.169
Carnac(20):
0.093
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Carnac(20)
Matthews Correlation Coefficient
Afold:
0.358
Carnac(20):
0.288
Sensitivity
Afold:
0.295
Carnac(20):
0.093
Positive Predictive Value
Afold:
0.434
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Carnac(20):
0.288
Sensitivity
MXScarna(20):
0.181
Carnac(20):
0.093
Positive Predictive Value
MXScarna(20):
0.710
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Carnac(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Carnac(20):
0.288
Sensitivity
CRWrnafold:
0.275
Carnac(20):
0.093
Positive Predictive Value
CRWrnafold:
0.435
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Carnac(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Carnac(20):
0.288
Sensitivity
RNAsubopt:
0.217
Carnac(20):
0.093
Positive Predictive Value
RNAsubopt:
0.521
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Carnac(20)
Matthews Correlation Coefficient
McQFold:
0.314
Carnac(20):
0.288
Sensitivity
McQFold:
0.205
Carnac(20):
0.093
Positive Predictive Value
McQFold:
0.480
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Carnac(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Carnac(20):
0.288
Sensitivity
TurboFold(20):
0.108
Carnac(20):
0.093
Positive Predictive Value
TurboFold(20):
0.837
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Carnac(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Carnac(20):
0.288
Sensitivity
PPfold(20):
0.101
Carnac(20):
0.093
Positive Predictive Value
PPfold(20):
0.879
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Carnac(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Carnac(20):
0.288
Sensitivity
RNAshapes:
0.180
Carnac(20):
0.093
Positive Predictive Value
RNAshapes:
0.484
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 4.48494289621e-08
|
|
+
Carnac(20) vs RNASLOpt
Matthews Correlation Coefficient
Carnac(20):
0.288
RNASLOpt:
0.281
Sensitivity
Carnac(20):
0.093
RNASLOpt:
0.158
Positive Predictive Value
Carnac(20):
0.892
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 6.06606238843e-08
|
+
Carnac(20) vs RSpredict(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
RSpredict(20):
0.270
Sensitivity
Carnac(20):
0.093
RSpredict(20):
0.124
Positive Predictive Value
Carnac(20):
0.892
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Multilign(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
Multilign(20):
0.251
Sensitivity
Carnac(20):
0.093
Multilign(20):
0.083
Positive Predictive Value
Carnac(20):
0.892
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNAwolf
Matthews Correlation Coefficient
Carnac(20):
0.288
RNAwolf:
0.251
Sensitivity
Carnac(20):
0.093
RNAwolf:
0.191
Positive Predictive Value
Carnac(20):
0.892
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Murlet(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
Murlet(20):
0.230
Sensitivity
Carnac(20):
0.093
Murlet(20):
0.067
Positive Predictive Value
Carnac(20):
0.892
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Vsfold4
Matthews Correlation Coefficient
Carnac(20):
0.288
Vsfold4:
0.217
Sensitivity
Carnac(20):
0.093
Vsfold4:
0.125
Positive Predictive Value
Carnac(20):
0.892
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Vsfold5
Matthews Correlation Coefficient
Carnac(20):
0.288
Vsfold5:
0.211
Sensitivity
Carnac(20):
0.093
Vsfold5:
0.125
Positive Predictive Value
Carnac(20):
0.892
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
RSpredict(seed):
0.212
Sensitivity
Carnac(20):
0.093
RSpredict(seed):
0.075
Positive Predictive Value
Carnac(20):
0.892
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs CMfinder(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
CMfinder(20):
0.210
Sensitivity
Carnac(20):
0.093
CMfinder(20):
0.055
Positive Predictive Value
Carnac(20):
0.892
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs HotKnots
Matthews Correlation Coefficient
Carnac(20):
0.288
HotKnots:
0.179
Sensitivity
Carnac(20):
0.093
HotKnots:
0.051
Positive Predictive Value
Carnac(20):
0.892
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNASampler(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
RNASampler(20):
0.176
Sensitivity
Carnac(20):
0.093
RNASampler(20):
0.035
Positive Predictive Value
Carnac(20):
0.892
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Cylofold
Matthews Correlation Coefficient
Carnac(20):
0.288
Cylofold:
0.155
Sensitivity
Carnac(20):
0.093
Cylofold:
0.040
Positive Predictive Value
Carnac(20):
0.892
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Pknots
Matthews Correlation Coefficient
Carnac(20):
0.288
Pknots:
0.143
Sensitivity
Carnac(20):
0.093
Pknots:
0.033
Positive Predictive Value
Carnac(20):
0.892
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Mastr(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
Mastr(20):
0.134
Sensitivity
Carnac(20):
0.093
Mastr(20):
0.023
Positive Predictive Value
Carnac(20):
0.892
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Alterna
Matthews Correlation Coefficient
Carnac(20):
0.288
Alterna:
0.129
Sensitivity
Carnac(20):
0.093
Alterna:
0.030
Positive Predictive Value
Carnac(20):
0.892
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs MCFold
Matthews Correlation Coefficient
Carnac(20):
0.288
MCFold:
0.125
Sensitivity
Carnac(20):
0.093
MCFold:
0.036
Positive Predictive Value
Carnac(20):
0.892
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RDfolder
Matthews Correlation Coefficient
Carnac(20):
0.288
RDfolder:
0.122
Sensitivity
Carnac(20):
0.093
RDfolder:
0.025
Positive Predictive Value
Carnac(20):
0.892
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Murlet(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Murlet(seed):
0.079
Sensitivity
Carnac(20):
0.093
Murlet(seed):
0.008
Positive Predictive Value
Carnac(20):
0.892
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
CMfinder(seed):
0.080
Sensitivity
Carnac(20):
0.093
CMfinder(seed):
0.008
Positive Predictive Value
Carnac(20):
0.892
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
TurboFold(seed):
0.076
Sensitivity
Carnac(20):
0.093
TurboFold(seed):
0.007
Positive Predictive Value
Carnac(20):
0.892
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs PPfold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
PPfold(seed):
0.069
Sensitivity
Carnac(20):
0.093
PPfold(seed):
0.006
Positive Predictive Value
Carnac(20):
0.892
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Carnac(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Carnac(seed):
0.066
Sensitivity
Carnac(20):
0.093
Carnac(seed):
0.005
Positive Predictive Value
Carnac(20):
0.892
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
RNASampler(seed):
0.063
Sensitivity
Carnac(20):
0.093
RNASampler(seed):
0.005
Positive Predictive Value
Carnac(20):
0.892
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs NanoFolder
Matthews Correlation Coefficient
Carnac(20):
0.288
NanoFolder:
0.052
Sensitivity
Carnac(20):
0.093
NanoFolder:
0.016
Positive Predictive Value
Carnac(20):
0.892
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Mastr(seed):
0.051
Sensitivity
Carnac(20):
0.093
Mastr(seed):
0.003
Positive Predictive Value
Carnac(20):
0.892
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Multilign(seed):
0.043
Sensitivity
Carnac(20):
0.093
Multilign(seed):
0.003
Positive Predictive Value
Carnac(20):
0.892
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNASLOpt |
1987
ContextFold vs RNASLOpt
Matthews Correlation Coefficient
ContextFold:
0.680
RNASLOpt:
0.281
Sensitivity
ContextFold:
0.644
RNASLOpt:
0.158
Positive Predictive Value
ContextFold:
0.718
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNASLOpt
Matthews Correlation Coefficient
IPknot:
0.548
RNASLOpt:
0.281
Sensitivity
IPknot:
0.475
RNASLOpt:
0.158
Positive Predictive Value
IPknot:
0.632
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNASLOpt:
0.281
Sensitivity
PETfold_pre2.0(seed):
0.343
RNASLOpt:
0.158
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNASLOpt
Matthews Correlation Coefficient
Contrafold:
0.508
RNASLOpt:
0.281
Sensitivity
Contrafold:
0.481
RNASLOpt:
0.158
Positive Predictive Value
Contrafold:
0.537
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNASLOpt
Matthews Correlation Coefficient
CentroidFold:
0.470
RNASLOpt:
0.281
Sensitivity
CentroidFold:
0.369
RNASLOpt:
0.158
Positive Predictive Value
CentroidFold:
0.600
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNASLOpt
Matthews Correlation Coefficient
Sfold:
0.469
RNASLOpt:
0.281
Sensitivity
Sfold:
0.412
RNASLOpt:
0.158
Positive Predictive Value
Sfold:
0.535
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNASLOpt
Matthews Correlation Coefficient
MaxExpect:
0.465
RNASLOpt:
0.281
Sensitivity
MaxExpect:
0.443
RNASLOpt:
0.158
Positive Predictive Value
MaxExpect:
0.488
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNASLOpt
Matthews Correlation Coefficient
ProbKnot:
0.460
RNASLOpt:
0.281
Sensitivity
ProbKnot:
0.446
RNASLOpt:
0.158
Positive Predictive Value
ProbKnot:
0.475
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNASLOpt
Matthews Correlation Coefficient
UNAFold:
0.445
RNASLOpt:
0.281
Sensitivity
UNAFold:
0.445
RNASLOpt:
0.158
Positive Predictive Value
UNAFold:
0.446
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNASLOpt:
0.281
Sensitivity
CentroidAlifold(seed):
0.213
RNASLOpt:
0.158
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNASLOpt
Matthews Correlation Coefficient
Fold:
0.437
RNASLOpt:
0.281
Sensitivity
Fold:
0.432
RNASLOpt:
0.158
Positive Predictive Value
Fold:
0.444
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNASLOpt
Matthews Correlation Coefficient
RNAfold:
0.429
RNASLOpt:
0.281
Sensitivity
RNAfold:
0.422
RNASLOpt:
0.158
Positive Predictive Value
RNAfold:
0.437
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNASLOpt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNASLOpt:
0.281
Sensitivity
CentroidHomfold‑LAST:
0.216
RNASLOpt:
0.158
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNASLOpt:
0.281
Sensitivity
PETfold_pre2.0(20):
0.211
RNASLOpt:
0.158
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNASLOpt:
0.281
Sensitivity
CentroidAlifold(20):
0.190
RNASLOpt:
0.158
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNASLOpt
Matthews Correlation Coefficient
PknotsRG:
0.386
RNASLOpt:
0.281
Sensitivity
PknotsRG:
0.320
RNASLOpt:
0.158
Positive Predictive Value
PknotsRG:
0.465
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNASLOpt:
0.281
Sensitivity
MXScarna(seed):
0.196
RNASLOpt:
0.158
Positive Predictive Value
MXScarna(seed):
0.753
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNASLOpt:
0.281
Sensitivity
RNAalifold(20):
0.185
RNASLOpt:
0.158
Positive Predictive Value
RNAalifold(20):
0.787
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNASLOpt:
0.281
Sensitivity
RNAalifold(seed):
0.169
RNASLOpt:
0.158
Positive Predictive Value
RNAalifold(seed):
0.835
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RNASLOpt
Matthews Correlation Coefficient
Afold:
0.358
RNASLOpt:
0.281
Sensitivity
Afold:
0.295
RNASLOpt:
0.158
Positive Predictive Value
Afold:
0.434
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNASLOpt:
0.281
Sensitivity
MXScarna(20):
0.181
RNASLOpt:
0.158
Positive Predictive Value
MXScarna(20):
0.710
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RNASLOpt
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNASLOpt:
0.281
Sensitivity
CRWrnafold:
0.275
RNASLOpt:
0.158
Positive Predictive Value
CRWrnafold:
0.435
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RNASLOpt
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNASLOpt:
0.281
Sensitivity
RNAsubopt:
0.217
RNASLOpt:
0.158
Positive Predictive Value
RNAsubopt:
0.521
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RNASLOpt
Matthews Correlation Coefficient
McQFold:
0.314
RNASLOpt:
0.281
Sensitivity
McQFold:
0.205
RNASLOpt:
0.158
Positive Predictive Value
McQFold:
0.480
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RNASLOpt
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNASLOpt:
0.281
Sensitivity
TurboFold(20):
0.108
RNASLOpt:
0.158
Positive Predictive Value
TurboFold(20):
0.837
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RNASLOpt
Matthews Correlation Coefficient
PPfold(20):
0.298
RNASLOpt:
0.281
Sensitivity
PPfold(20):
0.101
RNASLOpt:
0.158
Positive Predictive Value
PPfold(20):
0.879
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RNASLOpt
Matthews Correlation Coefficient
RNAshapes:
0.295
RNASLOpt:
0.281
Sensitivity
RNAshapes:
0.180
RNASLOpt:
0.158
Positive Predictive Value
RNAshapes:
0.484
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RNASLOpt
Matthews Correlation Coefficient
Carnac(20):
0.288
RNASLOpt:
0.281
Sensitivity
Carnac(20):
0.093
RNASLOpt:
0.158
Positive Predictive Value
Carnac(20):
0.892
RNASLOpt:
0.502
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 6.06606238843e-08
|
|
+
RNASLOpt vs RSpredict(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RSpredict(20):
0.270
Sensitivity
RNASLOpt:
0.158
RSpredict(20):
0.124
Positive Predictive Value
RNASLOpt:
0.502
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Multilign(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Multilign(20):
0.251
Sensitivity
RNASLOpt:
0.158
Multilign(20):
0.083
Positive Predictive Value
RNASLOpt:
0.502
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RNAwolf
Matthews Correlation Coefficient
RNASLOpt:
0.281
RNAwolf:
0.251
Sensitivity
RNASLOpt:
0.158
RNAwolf:
0.191
Positive Predictive Value
RNASLOpt:
0.502
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Murlet(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Murlet(20):
0.230
Sensitivity
RNASLOpt:
0.158
Murlet(20):
0.067
Positive Predictive Value
RNASLOpt:
0.502
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Vsfold4
Matthews Correlation Coefficient
RNASLOpt:
0.281
Vsfold4:
0.217
Sensitivity
RNASLOpt:
0.158
Vsfold4:
0.125
Positive Predictive Value
RNASLOpt:
0.502
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Vsfold5
Matthews Correlation Coefficient
RNASLOpt:
0.281
Vsfold5:
0.211
Sensitivity
RNASLOpt:
0.158
Vsfold5:
0.125
Positive Predictive Value
RNASLOpt:
0.502
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RSpredict(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RSpredict(seed):
0.212
Sensitivity
RNASLOpt:
0.158
RSpredict(seed):
0.075
Positive Predictive Value
RNASLOpt:
0.502
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs CMfinder(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
CMfinder(20):
0.210
Sensitivity
RNASLOpt:
0.158
CMfinder(20):
0.055
Positive Predictive Value
RNASLOpt:
0.502
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs HotKnots
Matthews Correlation Coefficient
RNASLOpt:
0.281
HotKnots:
0.179
Sensitivity
RNASLOpt:
0.158
HotKnots:
0.051
Positive Predictive Value
RNASLOpt:
0.502
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RNASampler(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RNASampler(20):
0.176
Sensitivity
RNASLOpt:
0.158
RNASampler(20):
0.035
Positive Predictive Value
RNASLOpt:
0.502
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Cylofold
Matthews Correlation Coefficient
RNASLOpt:
0.281
Cylofold:
0.155
Sensitivity
RNASLOpt:
0.158
Cylofold:
0.040
Positive Predictive Value
RNASLOpt:
0.502
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Pknots
Matthews Correlation Coefficient
RNASLOpt:
0.281
Pknots:
0.143
Sensitivity
RNASLOpt:
0.158
Pknots:
0.033
Positive Predictive Value
RNASLOpt:
0.502
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Mastr(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Mastr(20):
0.134
Sensitivity
RNASLOpt:
0.158
Mastr(20):
0.023
Positive Predictive Value
RNASLOpt:
0.502
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Alterna
Matthews Correlation Coefficient
RNASLOpt:
0.281
Alterna:
0.129
Sensitivity
RNASLOpt:
0.158
Alterna:
0.030
Positive Predictive Value
RNASLOpt:
0.502
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs MCFold
Matthews Correlation Coefficient
RNASLOpt:
0.281
MCFold:
0.125
Sensitivity
RNASLOpt:
0.158
MCFold:
0.036
Positive Predictive Value
RNASLOpt:
0.502
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RDfolder
Matthews Correlation Coefficient
RNASLOpt:
0.281
RDfolder:
0.122
Sensitivity
RNASLOpt:
0.158
RDfolder:
0.025
Positive Predictive Value
RNASLOpt:
0.502
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Murlet(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Murlet(seed):
0.079
Sensitivity
RNASLOpt:
0.158
Murlet(seed):
0.008
Positive Predictive Value
RNASLOpt:
0.502
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs CMfinder(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
CMfinder(seed):
0.080
Sensitivity
RNASLOpt:
0.158
CMfinder(seed):
0.008
Positive Predictive Value
RNASLOpt:
0.502
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs TurboFold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
TurboFold(seed):
0.076
Sensitivity
RNASLOpt:
0.158
TurboFold(seed):
0.007
Positive Predictive Value
RNASLOpt:
0.502
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs PPfold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
PPfold(seed):
0.069
Sensitivity
RNASLOpt:
0.158
PPfold(seed):
0.006
Positive Predictive Value
RNASLOpt:
0.502
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Carnac(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Carnac(seed):
0.066
Sensitivity
RNASLOpt:
0.158
Carnac(seed):
0.005
Positive Predictive Value
RNASLOpt:
0.502
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RNASampler(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RNASampler(seed):
0.063
Sensitivity
RNASLOpt:
0.158
RNASampler(seed):
0.005
Positive Predictive Value
RNASLOpt:
0.502
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs NanoFolder
Matthews Correlation Coefficient
RNASLOpt:
0.281
NanoFolder:
0.052
Sensitivity
RNASLOpt:
0.158
NanoFolder:
0.016
Positive Predictive Value
RNASLOpt:
0.502
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Mastr(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Mastr(seed):
0.051
Sensitivity
RNASLOpt:
0.158
Mastr(seed):
0.003
Positive Predictive Value
RNASLOpt:
0.502
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Multilign(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Multilign(seed):
0.043
Sensitivity
RNASLOpt:
0.158
Multilign(seed):
0.003
Positive Predictive Value
RNASLOpt:
0.502
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RSpredict(20) |
1987
ContextFold vs RSpredict(20)
Matthews Correlation Coefficient
ContextFold:
0.680
RSpredict(20):
0.270
Sensitivity
ContextFold:
0.644
RSpredict(20):
0.124
Positive Predictive Value
ContextFold:
0.718
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RSpredict(20)
Matthews Correlation Coefficient
IPknot:
0.548
RSpredict(20):
0.270
Sensitivity
IPknot:
0.475
RSpredict(20):
0.124
Positive Predictive Value
IPknot:
0.632
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RSpredict(20):
0.270
Sensitivity
PETfold_pre2.0(seed):
0.343
RSpredict(20):
0.124
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RSpredict(20)
Matthews Correlation Coefficient
Contrafold:
0.508
RSpredict(20):
0.270
Sensitivity
Contrafold:
0.481
RSpredict(20):
0.124
Positive Predictive Value
Contrafold:
0.537
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RSpredict(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
RSpredict(20):
0.270
Sensitivity
CentroidFold:
0.369
RSpredict(20):
0.124
Positive Predictive Value
CentroidFold:
0.600
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RSpredict(20)
Matthews Correlation Coefficient
Sfold:
0.469
RSpredict(20):
0.270
Sensitivity
Sfold:
0.412
RSpredict(20):
0.124
Positive Predictive Value
Sfold:
0.535
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RSpredict(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
RSpredict(20):
0.270
Sensitivity
MaxExpect:
0.443
RSpredict(20):
0.124
Positive Predictive Value
MaxExpect:
0.488
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RSpredict(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
RSpredict(20):
0.270
Sensitivity
ProbKnot:
0.446
RSpredict(20):
0.124
Positive Predictive Value
ProbKnot:
0.475
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RSpredict(20)
Matthews Correlation Coefficient
UNAFold:
0.445
RSpredict(20):
0.270
Sensitivity
UNAFold:
0.445
RSpredict(20):
0.124
Positive Predictive Value
UNAFold:
0.446
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RSpredict(20):
0.270
Sensitivity
CentroidAlifold(seed):
0.213
RSpredict(20):
0.124
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RSpredict(20)
Matthews Correlation Coefficient
Fold:
0.437
RSpredict(20):
0.270
Sensitivity
Fold:
0.432
RSpredict(20):
0.124
Positive Predictive Value
Fold:
0.444
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RSpredict(20)
Matthews Correlation Coefficient
RNAfold:
0.429
RSpredict(20):
0.270
Sensitivity
RNAfold:
0.422
RSpredict(20):
0.124
Positive Predictive Value
RNAfold:
0.437
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RSpredict(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RSpredict(20):
0.270
Sensitivity
CentroidHomfold‑LAST:
0.216
RSpredict(20):
0.124
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RSpredict(20):
0.270
Sensitivity
PETfold_pre2.0(20):
0.211
RSpredict(20):
0.124
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RSpredict(20):
0.270
Sensitivity
CentroidAlifold(20):
0.190
RSpredict(20):
0.124
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RSpredict(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
RSpredict(20):
0.270
Sensitivity
PknotsRG:
0.320
RSpredict(20):
0.124
Positive Predictive Value
PknotsRG:
0.465
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RSpredict(20):
0.270
Sensitivity
MXScarna(seed):
0.196
RSpredict(20):
0.124
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RSpredict(20):
0.270
Sensitivity
RNAalifold(20):
0.185
RSpredict(20):
0.124
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RSpredict(20):
0.270
Sensitivity
RNAalifold(seed):
0.169
RSpredict(20):
0.124
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RSpredict(20)
Matthews Correlation Coefficient
Afold:
0.358
RSpredict(20):
0.270
Sensitivity
Afold:
0.295
RSpredict(20):
0.124
Positive Predictive Value
Afold:
0.434
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RSpredict(20):
0.270
Sensitivity
MXScarna(20):
0.181
RSpredict(20):
0.124
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RSpredict(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RSpredict(20):
0.270
Sensitivity
CRWrnafold:
0.275
RSpredict(20):
0.124
Positive Predictive Value
CRWrnafold:
0.435
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RSpredict(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RSpredict(20):
0.270
Sensitivity
RNAsubopt:
0.217
RSpredict(20):
0.124
Positive Predictive Value
RNAsubopt:
0.521
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RSpredict(20)
Matthews Correlation Coefficient
McQFold:
0.314
RSpredict(20):
0.270
Sensitivity
McQFold:
0.205
RSpredict(20):
0.124
Positive Predictive Value
McQFold:
0.480
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RSpredict(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RSpredict(20):
0.270
Sensitivity
TurboFold(20):
0.108
RSpredict(20):
0.124
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RSpredict(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
RSpredict(20):
0.270
Sensitivity
PPfold(20):
0.101
RSpredict(20):
0.124
Positive Predictive Value
PPfold(20):
0.879
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RSpredict(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
RSpredict(20):
0.270
Sensitivity
RNAshapes:
0.180
RSpredict(20):
0.124
Positive Predictive Value
RNAshapes:
0.484
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RSpredict(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
RSpredict(20):
0.270
Sensitivity
Carnac(20):
0.093
RSpredict(20):
0.124
Positive Predictive Value
Carnac(20):
0.892
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs RSpredict(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RSpredict(20):
0.270
Sensitivity
RNASLOpt:
0.158
RSpredict(20):
0.124
Positive Predictive Value
RNASLOpt:
0.502
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RSpredict(20) vs Multilign(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Multilign(20):
0.251
Sensitivity
RSpredict(20):
0.124
Multilign(20):
0.083
Positive Predictive Value
RSpredict(20):
0.590
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RNAwolf
Matthews Correlation Coefficient
RSpredict(20):
0.270
RNAwolf:
0.251
Sensitivity
RSpredict(20):
0.124
RNAwolf:
0.191
Positive Predictive Value
RSpredict(20):
0.590
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Murlet(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Murlet(20):
0.230
Sensitivity
RSpredict(20):
0.124
Murlet(20):
0.067
Positive Predictive Value
RSpredict(20):
0.590
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Vsfold4
Matthews Correlation Coefficient
RSpredict(20):
0.270
Vsfold4:
0.217
Sensitivity
RSpredict(20):
0.124
Vsfold4:
0.125
Positive Predictive Value
RSpredict(20):
0.590
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(20):
0.270
Vsfold5:
0.211
Sensitivity
RSpredict(20):
0.124
Vsfold5:
0.125
Positive Predictive Value
RSpredict(20):
0.590
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
RSpredict(seed):
0.212
Sensitivity
RSpredict(20):
0.124
RSpredict(seed):
0.075
Positive Predictive Value
RSpredict(20):
0.590
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
CMfinder(20):
0.210
Sensitivity
RSpredict(20):
0.124
CMfinder(20):
0.055
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs HotKnots
Matthews Correlation Coefficient
RSpredict(20):
0.270
HotKnots:
0.179
Sensitivity
RSpredict(20):
0.124
HotKnots:
0.051
Positive Predictive Value
RSpredict(20):
0.590
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
RNASampler(20):
0.176
Sensitivity
RSpredict(20):
0.124
RNASampler(20):
0.035
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Cylofold
Matthews Correlation Coefficient
RSpredict(20):
0.270
Cylofold:
0.155
Sensitivity
RSpredict(20):
0.124
Cylofold:
0.040
Positive Predictive Value
RSpredict(20):
0.590
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Pknots
Matthews Correlation Coefficient
RSpredict(20):
0.270
Pknots:
0.143
Sensitivity
RSpredict(20):
0.124
Pknots:
0.033
Positive Predictive Value
RSpredict(20):
0.590
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Mastr(20):
0.134
Sensitivity
RSpredict(20):
0.124
Mastr(20):
0.023
Positive Predictive Value
RSpredict(20):
0.590
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Alterna
Matthews Correlation Coefficient
RSpredict(20):
0.270
Alterna:
0.129
Sensitivity
RSpredict(20):
0.124
Alterna:
0.030
Positive Predictive Value
RSpredict(20):
0.590
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs MCFold
Matthews Correlation Coefficient
RSpredict(20):
0.270
MCFold:
0.125
Sensitivity
RSpredict(20):
0.124
MCFold:
0.036
Positive Predictive Value
RSpredict(20):
0.590
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RDfolder
Matthews Correlation Coefficient
RSpredict(20):
0.270
RDfolder:
0.122
Sensitivity
RSpredict(20):
0.124
RDfolder:
0.025
Positive Predictive Value
RSpredict(20):
0.590
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Murlet(seed):
0.079
Sensitivity
RSpredict(20):
0.124
Murlet(seed):
0.008
Positive Predictive Value
RSpredict(20):
0.590
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
CMfinder(seed):
0.080
Sensitivity
RSpredict(20):
0.124
CMfinder(seed):
0.008
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
TurboFold(seed):
0.076
Sensitivity
RSpredict(20):
0.124
TurboFold(seed):
0.007
Positive Predictive Value
RSpredict(20):
0.590
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
PPfold(seed):
0.069
Sensitivity
RSpredict(20):
0.124
PPfold(seed):
0.006
Positive Predictive Value
RSpredict(20):
0.590
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Carnac(seed):
0.066
Sensitivity
RSpredict(20):
0.124
Carnac(seed):
0.005
Positive Predictive Value
RSpredict(20):
0.590
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
RNASampler(seed):
0.063
Sensitivity
RSpredict(20):
0.124
RNASampler(seed):
0.005
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(20):
0.270
NanoFolder:
0.052
Sensitivity
RSpredict(20):
0.124
NanoFolder:
0.016
Positive Predictive Value
RSpredict(20):
0.590
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Mastr(seed):
0.051
Sensitivity
RSpredict(20):
0.124
Mastr(seed):
0.003
Positive Predictive Value
RSpredict(20):
0.590
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Multilign(seed):
0.043
Sensitivity
RSpredict(20):
0.124
Multilign(seed):
0.003
Positive Predictive Value
RSpredict(20):
0.590
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Multilign(20) |
1987
ContextFold vs Multilign(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Multilign(20):
0.251
Sensitivity
ContextFold:
0.644
Multilign(20):
0.083
Positive Predictive Value
ContextFold:
0.718
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Multilign(20)
Matthews Correlation Coefficient
IPknot:
0.548
Multilign(20):
0.251
Sensitivity
IPknot:
0.475
Multilign(20):
0.083
Positive Predictive Value
IPknot:
0.632
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Multilign(20):
0.251
Sensitivity
PETfold_pre2.0(seed):
0.343
Multilign(20):
0.083
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Multilign(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Multilign(20):
0.251
Sensitivity
Contrafold:
0.481
Multilign(20):
0.083
Positive Predictive Value
Contrafold:
0.537
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Multilign(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Multilign(20):
0.251
Sensitivity
CentroidFold:
0.369
Multilign(20):
0.083
Positive Predictive Value
CentroidFold:
0.600
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Multilign(20)
Matthews Correlation Coefficient
Sfold:
0.469
Multilign(20):
0.251
Sensitivity
Sfold:
0.412
Multilign(20):
0.083
Positive Predictive Value
Sfold:
0.535
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Multilign(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Multilign(20):
0.251
Sensitivity
MaxExpect:
0.443
Multilign(20):
0.083
Positive Predictive Value
MaxExpect:
0.488
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Multilign(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Multilign(20):
0.251
Sensitivity
ProbKnot:
0.446
Multilign(20):
0.083
Positive Predictive Value
ProbKnot:
0.475
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Multilign(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Multilign(20):
0.251
Sensitivity
UNAFold:
0.445
Multilign(20):
0.083
Positive Predictive Value
UNAFold:
0.446
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Multilign(20):
0.251
Sensitivity
CentroidAlifold(seed):
0.213
Multilign(20):
0.083
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Multilign(20)
Matthews Correlation Coefficient
Fold:
0.437
Multilign(20):
0.251
Sensitivity
Fold:
0.432
Multilign(20):
0.083
Positive Predictive Value
Fold:
0.444
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Multilign(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Multilign(20):
0.251
Sensitivity
RNAfold:
0.422
Multilign(20):
0.083
Positive Predictive Value
RNAfold:
0.437
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Multilign(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Multilign(20):
0.251
Sensitivity
CentroidHomfold‑LAST:
0.216
Multilign(20):
0.083
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Multilign(20):
0.251
Sensitivity
PETfold_pre2.0(20):
0.211
Multilign(20):
0.083
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Multilign(20):
0.251
Sensitivity
CentroidAlifold(20):
0.190
Multilign(20):
0.083
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Multilign(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Multilign(20):
0.251
Sensitivity
PknotsRG:
0.320
Multilign(20):
0.083
Positive Predictive Value
PknotsRG:
0.465
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Multilign(20):
0.251
Sensitivity
MXScarna(seed):
0.196
Multilign(20):
0.083
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Multilign(20):
0.251
Sensitivity
RNAalifold(20):
0.185
Multilign(20):
0.083
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Multilign(20):
0.251
Sensitivity
RNAalifold(seed):
0.169
Multilign(20):
0.083
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Multilign(20)
Matthews Correlation Coefficient
Afold:
0.358
Multilign(20):
0.251
Sensitivity
Afold:
0.295
Multilign(20):
0.083
Positive Predictive Value
Afold:
0.434
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Multilign(20):
0.251
Sensitivity
MXScarna(20):
0.181
Multilign(20):
0.083
Positive Predictive Value
MXScarna(20):
0.710
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Multilign(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Multilign(20):
0.251
Sensitivity
CRWrnafold:
0.275
Multilign(20):
0.083
Positive Predictive Value
CRWrnafold:
0.435
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Multilign(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Multilign(20):
0.251
Sensitivity
RNAsubopt:
0.217
Multilign(20):
0.083
Positive Predictive Value
RNAsubopt:
0.521
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Multilign(20)
Matthews Correlation Coefficient
McQFold:
0.314
Multilign(20):
0.251
Sensitivity
McQFold:
0.205
Multilign(20):
0.083
Positive Predictive Value
McQFold:
0.480
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Multilign(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Multilign(20):
0.251
Sensitivity
TurboFold(20):
0.108
Multilign(20):
0.083
Positive Predictive Value
TurboFold(20):
0.837
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Multilign(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Multilign(20):
0.251
Sensitivity
PPfold(20):
0.101
Multilign(20):
0.083
Positive Predictive Value
PPfold(20):
0.879
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Multilign(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Multilign(20):
0.251
Sensitivity
RNAshapes:
0.180
Multilign(20):
0.083
Positive Predictive Value
RNAshapes:
0.484
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Multilign(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
Multilign(20):
0.251
Sensitivity
Carnac(20):
0.093
Multilign(20):
0.083
Positive Predictive Value
Carnac(20):
0.892
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Multilign(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Multilign(20):
0.251
Sensitivity
RNASLOpt:
0.158
Multilign(20):
0.083
Positive Predictive Value
RNASLOpt:
0.502
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Multilign(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Multilign(20):
0.251
Sensitivity
RSpredict(20):
0.124
Multilign(20):
0.083
Positive Predictive Value
RSpredict(20):
0.590
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
Multilign(20) vs RNAwolf
Matthews Correlation Coefficient
Multilign(20):
0.251
RNAwolf:
0.251
Sensitivity
Multilign(20):
0.083
RNAwolf:
0.191
Positive Predictive Value
Multilign(20):
0.762
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.412262207143
|
+
Multilign(20) vs Murlet(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
Murlet(20):
0.230
Sensitivity
Multilign(20):
0.083
Murlet(20):
0.067
Positive Predictive Value
Multilign(20):
0.762
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Vsfold4
Matthews Correlation Coefficient
Multilign(20):
0.251
Vsfold4:
0.217
Sensitivity
Multilign(20):
0.083
Vsfold4:
0.125
Positive Predictive Value
Multilign(20):
0.762
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Vsfold5
Matthews Correlation Coefficient
Multilign(20):
0.251
Vsfold5:
0.211
Sensitivity
Multilign(20):
0.083
Vsfold5:
0.125
Positive Predictive Value
Multilign(20):
0.762
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
RSpredict(seed):
0.212
Sensitivity
Multilign(20):
0.083
RSpredict(seed):
0.075
Positive Predictive Value
Multilign(20):
0.762
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs CMfinder(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
CMfinder(20):
0.210
Sensitivity
Multilign(20):
0.083
CMfinder(20):
0.055
Positive Predictive Value
Multilign(20):
0.762
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs HotKnots
Matthews Correlation Coefficient
Multilign(20):
0.251
HotKnots:
0.179
Sensitivity
Multilign(20):
0.083
HotKnots:
0.051
Positive Predictive Value
Multilign(20):
0.762
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RNASampler(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
RNASampler(20):
0.176
Sensitivity
Multilign(20):
0.083
RNASampler(20):
0.035
Positive Predictive Value
Multilign(20):
0.762
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Cylofold
Matthews Correlation Coefficient
Multilign(20):
0.251
Cylofold:
0.155
Sensitivity
Multilign(20):
0.083
Cylofold:
0.040
Positive Predictive Value
Multilign(20):
0.762
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Pknots
Matthews Correlation Coefficient
Multilign(20):
0.251
Pknots:
0.143
Sensitivity
Multilign(20):
0.083
Pknots:
0.033
Positive Predictive Value
Multilign(20):
0.762
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Mastr(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
Mastr(20):
0.134
Sensitivity
Multilign(20):
0.083
Mastr(20):
0.023
Positive Predictive Value
Multilign(20):
0.762
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Alterna
Matthews Correlation Coefficient
Multilign(20):
0.251
Alterna:
0.129
Sensitivity
Multilign(20):
0.083
Alterna:
0.030
Positive Predictive Value
Multilign(20):
0.762
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs MCFold
Matthews Correlation Coefficient
Multilign(20):
0.251
MCFold:
0.125
Sensitivity
Multilign(20):
0.083
MCFold:
0.036
Positive Predictive Value
Multilign(20):
0.762
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RDfolder
Matthews Correlation Coefficient
Multilign(20):
0.251
RDfolder:
0.122
Sensitivity
Multilign(20):
0.083
RDfolder:
0.025
Positive Predictive Value
Multilign(20):
0.762
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Murlet(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Murlet(seed):
0.079
Sensitivity
Multilign(20):
0.083
Murlet(seed):
0.008
Positive Predictive Value
Multilign(20):
0.762
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
CMfinder(seed):
0.080
Sensitivity
Multilign(20):
0.083
CMfinder(seed):
0.008
Positive Predictive Value
Multilign(20):
0.762
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
TurboFold(seed):
0.076
Sensitivity
Multilign(20):
0.083
TurboFold(seed):
0.007
Positive Predictive Value
Multilign(20):
0.762
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs PPfold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
PPfold(seed):
0.069
Sensitivity
Multilign(20):
0.083
PPfold(seed):
0.006
Positive Predictive Value
Multilign(20):
0.762
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Carnac(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Carnac(seed):
0.066
Sensitivity
Multilign(20):
0.083
Carnac(seed):
0.005
Positive Predictive Value
Multilign(20):
0.762
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
RNASampler(seed):
0.063
Sensitivity
Multilign(20):
0.083
RNASampler(seed):
0.005
Positive Predictive Value
Multilign(20):
0.762
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs NanoFolder
Matthews Correlation Coefficient
Multilign(20):
0.251
NanoFolder:
0.052
Sensitivity
Multilign(20):
0.083
NanoFolder:
0.016
Positive Predictive Value
Multilign(20):
0.762
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Mastr(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Mastr(seed):
0.051
Sensitivity
Multilign(20):
0.083
Mastr(seed):
0.003
Positive Predictive Value
Multilign(20):
0.762
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Multilign(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Multilign(seed):
0.043
Sensitivity
Multilign(20):
0.083
Multilign(seed):
0.003
Positive Predictive Value
Multilign(20):
0.762
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAwolf |
1987
ContextFold vs RNAwolf
Matthews Correlation Coefficient
ContextFold:
0.680
RNAwolf:
0.251
Sensitivity
ContextFold:
0.644
RNAwolf:
0.191
Positive Predictive Value
ContextFold:
0.718
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNAwolf
Matthews Correlation Coefficient
IPknot:
0.548
RNAwolf:
0.251
Sensitivity
IPknot:
0.475
RNAwolf:
0.191
Positive Predictive Value
IPknot:
0.632
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNAwolf:
0.251
Sensitivity
PETfold_pre2.0(seed):
0.343
RNAwolf:
0.191
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNAwolf
Matthews Correlation Coefficient
Contrafold:
0.508
RNAwolf:
0.251
Sensitivity
Contrafold:
0.481
RNAwolf:
0.191
Positive Predictive Value
Contrafold:
0.537
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNAwolf
Matthews Correlation Coefficient
CentroidFold:
0.470
RNAwolf:
0.251
Sensitivity
CentroidFold:
0.369
RNAwolf:
0.191
Positive Predictive Value
CentroidFold:
0.600
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNAwolf
Matthews Correlation Coefficient
Sfold:
0.469
RNAwolf:
0.251
Sensitivity
Sfold:
0.412
RNAwolf:
0.191
Positive Predictive Value
Sfold:
0.535
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNAwolf
Matthews Correlation Coefficient
MaxExpect:
0.465
RNAwolf:
0.251
Sensitivity
MaxExpect:
0.443
RNAwolf:
0.191
Positive Predictive Value
MaxExpect:
0.488
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNAwolf
Matthews Correlation Coefficient
ProbKnot:
0.460
RNAwolf:
0.251
Sensitivity
ProbKnot:
0.446
RNAwolf:
0.191
Positive Predictive Value
ProbKnot:
0.475
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNAwolf
Matthews Correlation Coefficient
UNAFold:
0.445
RNAwolf:
0.251
Sensitivity
UNAFold:
0.445
RNAwolf:
0.191
Positive Predictive Value
UNAFold:
0.446
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNAwolf:
0.251
Sensitivity
CentroidAlifold(seed):
0.213
RNAwolf:
0.191
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNAwolf
Matthews Correlation Coefficient
Fold:
0.437
RNAwolf:
0.251
Sensitivity
Fold:
0.432
RNAwolf:
0.191
Positive Predictive Value
Fold:
0.444
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNAwolf
Matthews Correlation Coefficient
RNAfold:
0.429
RNAwolf:
0.251
Sensitivity
RNAfold:
0.422
RNAwolf:
0.191
Positive Predictive Value
RNAfold:
0.437
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNAwolf
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNAwolf:
0.251
Sensitivity
CentroidHomfold‑LAST:
0.216
RNAwolf:
0.191
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNAwolf:
0.251
Sensitivity
PETfold_pre2.0(20):
0.211
RNAwolf:
0.191
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNAwolf:
0.251
Sensitivity
CentroidAlifold(20):
0.190
RNAwolf:
0.191
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNAwolf
Matthews Correlation Coefficient
PknotsRG:
0.386
RNAwolf:
0.251
Sensitivity
PknotsRG:
0.320
RNAwolf:
0.191
Positive Predictive Value
PknotsRG:
0.465
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNAwolf:
0.251
Sensitivity
MXScarna(seed):
0.196
RNAwolf:
0.191
Positive Predictive Value
MXScarna(seed):
0.753
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNAwolf:
0.251
Sensitivity
RNAalifold(20):
0.185
RNAwolf:
0.191
Positive Predictive Value
RNAalifold(20):
0.787
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNAwolf:
0.251
Sensitivity
RNAalifold(seed):
0.169
RNAwolf:
0.191
Positive Predictive Value
RNAalifold(seed):
0.835
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RNAwolf
Matthews Correlation Coefficient
Afold:
0.358
RNAwolf:
0.251
Sensitivity
Afold:
0.295
RNAwolf:
0.191
Positive Predictive Value
Afold:
0.434
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNAwolf:
0.251
Sensitivity
MXScarna(20):
0.181
RNAwolf:
0.191
Positive Predictive Value
MXScarna(20):
0.710
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RNAwolf
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNAwolf:
0.251
Sensitivity
CRWrnafold:
0.275
RNAwolf:
0.191
Positive Predictive Value
CRWrnafold:
0.435
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RNAwolf
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNAwolf:
0.251
Sensitivity
RNAsubopt:
0.217
RNAwolf:
0.191
Positive Predictive Value
RNAsubopt:
0.521
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RNAwolf
Matthews Correlation Coefficient
McQFold:
0.314
RNAwolf:
0.251
Sensitivity
McQFold:
0.205
RNAwolf:
0.191
Positive Predictive Value
McQFold:
0.480
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RNAwolf
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNAwolf:
0.251
Sensitivity
TurboFold(20):
0.108
RNAwolf:
0.191
Positive Predictive Value
TurboFold(20):
0.837
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RNAwolf
Matthews Correlation Coefficient
PPfold(20):
0.298
RNAwolf:
0.251
Sensitivity
PPfold(20):
0.101
RNAwolf:
0.191
Positive Predictive Value
PPfold(20):
0.879
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RNAwolf
Matthews Correlation Coefficient
RNAshapes:
0.295
RNAwolf:
0.251
Sensitivity
RNAshapes:
0.180
RNAwolf:
0.191
Positive Predictive Value
RNAshapes:
0.484
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RNAwolf
Matthews Correlation Coefficient
Carnac(20):
0.288
RNAwolf:
0.251
Sensitivity
Carnac(20):
0.093
RNAwolf:
0.191
Positive Predictive Value
Carnac(20):
0.892
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs RNAwolf
Matthews Correlation Coefficient
RNASLOpt:
0.281
RNAwolf:
0.251
Sensitivity
RNASLOpt:
0.158
RNAwolf:
0.191
Positive Predictive Value
RNASLOpt:
0.502
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs RNAwolf
Matthews Correlation Coefficient
RSpredict(20):
0.270
RNAwolf:
0.251
Sensitivity
RSpredict(20):
0.124
RNAwolf:
0.191
Positive Predictive Value
RSpredict(20):
0.590
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs RNAwolf
Matthews Correlation Coefficient
Multilign(20):
0.251
RNAwolf:
0.251
Sensitivity
Multilign(20):
0.083
RNAwolf:
0.191
Positive Predictive Value
Multilign(20):
0.762
RNAwolf:
0.330
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.412262207143
|
|
+
RNAwolf vs Murlet(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
Murlet(20):
0.230
Sensitivity
RNAwolf:
0.191
Murlet(20):
0.067
Positive Predictive Value
RNAwolf:
0.330
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Vsfold4
Matthews Correlation Coefficient
RNAwolf:
0.251
Vsfold4:
0.217
Sensitivity
RNAwolf:
0.191
Vsfold4:
0.125
Positive Predictive Value
RNAwolf:
0.330
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Vsfold5
Matthews Correlation Coefficient
RNAwolf:
0.251
Vsfold5:
0.211
Sensitivity
RNAwolf:
0.191
Vsfold5:
0.125
Positive Predictive Value
RNAwolf:
0.330
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RSpredict(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
RSpredict(seed):
0.212
Sensitivity
RNAwolf:
0.191
RSpredict(seed):
0.075
Positive Predictive Value
RNAwolf:
0.330
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs CMfinder(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
CMfinder(20):
0.210
Sensitivity
RNAwolf:
0.191
CMfinder(20):
0.055
Positive Predictive Value
RNAwolf:
0.330
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs HotKnots
Matthews Correlation Coefficient
RNAwolf:
0.251
HotKnots:
0.179
Sensitivity
RNAwolf:
0.191
HotKnots:
0.051
Positive Predictive Value
RNAwolf:
0.330
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RNASampler(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
RNASampler(20):
0.176
Sensitivity
RNAwolf:
0.191
RNASampler(20):
0.035
Positive Predictive Value
RNAwolf:
0.330
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Cylofold
Matthews Correlation Coefficient
RNAwolf:
0.251
Cylofold:
0.155
Sensitivity
RNAwolf:
0.191
Cylofold:
0.040
Positive Predictive Value
RNAwolf:
0.330
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Pknots
Matthews Correlation Coefficient
RNAwolf:
0.251
Pknots:
0.143
Sensitivity
RNAwolf:
0.191
Pknots:
0.033
Positive Predictive Value
RNAwolf:
0.330
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Mastr(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
Mastr(20):
0.134
Sensitivity
RNAwolf:
0.191
Mastr(20):
0.023
Positive Predictive Value
RNAwolf:
0.330
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Alterna
Matthews Correlation Coefficient
RNAwolf:
0.251
Alterna:
0.129
Sensitivity
RNAwolf:
0.191
Alterna:
0.030
Positive Predictive Value
RNAwolf:
0.330
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs MCFold
Matthews Correlation Coefficient
RNAwolf:
0.251
MCFold:
0.125
Sensitivity
RNAwolf:
0.191
MCFold:
0.036
Positive Predictive Value
RNAwolf:
0.330
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RDfolder
Matthews Correlation Coefficient
RNAwolf:
0.251
RDfolder:
0.122
Sensitivity
RNAwolf:
0.191
RDfolder:
0.025
Positive Predictive Value
RNAwolf:
0.330
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Murlet(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Murlet(seed):
0.079
Sensitivity
RNAwolf:
0.191
Murlet(seed):
0.008
Positive Predictive Value
RNAwolf:
0.330
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs CMfinder(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
CMfinder(seed):
0.080
Sensitivity
RNAwolf:
0.191
CMfinder(seed):
0.008
Positive Predictive Value
RNAwolf:
0.330
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs TurboFold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
TurboFold(seed):
0.076
Sensitivity
RNAwolf:
0.191
TurboFold(seed):
0.007
Positive Predictive Value
RNAwolf:
0.330
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs PPfold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
PPfold(seed):
0.069
Sensitivity
RNAwolf:
0.191
PPfold(seed):
0.006
Positive Predictive Value
RNAwolf:
0.330
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Carnac(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Carnac(seed):
0.066
Sensitivity
RNAwolf:
0.191
Carnac(seed):
0.005
Positive Predictive Value
RNAwolf:
0.330
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RNASampler(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
RNASampler(seed):
0.063
Sensitivity
RNAwolf:
0.191
RNASampler(seed):
0.005
Positive Predictive Value
RNAwolf:
0.330
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs NanoFolder
Matthews Correlation Coefficient
RNAwolf:
0.251
NanoFolder:
0.052
Sensitivity
RNAwolf:
0.191
NanoFolder:
0.016
Positive Predictive Value
RNAwolf:
0.330
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Mastr(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Mastr(seed):
0.051
Sensitivity
RNAwolf:
0.191
Mastr(seed):
0.003
Positive Predictive Value
RNAwolf:
0.330
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Multilign(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Multilign(seed):
0.043
Sensitivity
RNAwolf:
0.191
Multilign(seed):
0.003
Positive Predictive Value
RNAwolf:
0.330
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Murlet(20) |
1987
ContextFold vs Murlet(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Murlet(20):
0.230
Sensitivity
ContextFold:
0.644
Murlet(20):
0.067
Positive Predictive Value
ContextFold:
0.718
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Murlet(20)
Matthews Correlation Coefficient
IPknot:
0.548
Murlet(20):
0.230
Sensitivity
IPknot:
0.475
Murlet(20):
0.067
Positive Predictive Value
IPknot:
0.632
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Murlet(20):
0.230
Sensitivity
PETfold_pre2.0(seed):
0.343
Murlet(20):
0.067
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Murlet(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Murlet(20):
0.230
Sensitivity
Contrafold:
0.481
Murlet(20):
0.067
Positive Predictive Value
Contrafold:
0.537
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Murlet(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Murlet(20):
0.230
Sensitivity
CentroidFold:
0.369
Murlet(20):
0.067
Positive Predictive Value
CentroidFold:
0.600
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Murlet(20)
Matthews Correlation Coefficient
Sfold:
0.469
Murlet(20):
0.230
Sensitivity
Sfold:
0.412
Murlet(20):
0.067
Positive Predictive Value
Sfold:
0.535
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Murlet(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Murlet(20):
0.230
Sensitivity
MaxExpect:
0.443
Murlet(20):
0.067
Positive Predictive Value
MaxExpect:
0.488
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Murlet(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Murlet(20):
0.230
Sensitivity
ProbKnot:
0.446
Murlet(20):
0.067
Positive Predictive Value
ProbKnot:
0.475
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Murlet(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Murlet(20):
0.230
Sensitivity
UNAFold:
0.445
Murlet(20):
0.067
Positive Predictive Value
UNAFold:
0.446
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Murlet(20):
0.230
Sensitivity
CentroidAlifold(seed):
0.213
Murlet(20):
0.067
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Murlet(20)
Matthews Correlation Coefficient
Fold:
0.437
Murlet(20):
0.230
Sensitivity
Fold:
0.432
Murlet(20):
0.067
Positive Predictive Value
Fold:
0.444
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Murlet(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Murlet(20):
0.230
Sensitivity
RNAfold:
0.422
Murlet(20):
0.067
Positive Predictive Value
RNAfold:
0.437
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Murlet(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Murlet(20):
0.230
Sensitivity
CentroidHomfold‑LAST:
0.216
Murlet(20):
0.067
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Murlet(20):
0.230
Sensitivity
PETfold_pre2.0(20):
0.211
Murlet(20):
0.067
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Murlet(20):
0.230
Sensitivity
CentroidAlifold(20):
0.190
Murlet(20):
0.067
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Murlet(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Murlet(20):
0.230
Sensitivity
PknotsRG:
0.320
Murlet(20):
0.067
Positive Predictive Value
PknotsRG:
0.465
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Murlet(20):
0.230
Sensitivity
MXScarna(seed):
0.196
Murlet(20):
0.067
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Murlet(20):
0.230
Sensitivity
RNAalifold(20):
0.185
Murlet(20):
0.067
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Murlet(20):
0.230
Sensitivity
RNAalifold(seed):
0.169
Murlet(20):
0.067
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Murlet(20)
Matthews Correlation Coefficient
Afold:
0.358
Murlet(20):
0.230
Sensitivity
Afold:
0.295
Murlet(20):
0.067
Positive Predictive Value
Afold:
0.434
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Murlet(20):
0.230
Sensitivity
MXScarna(20):
0.181
Murlet(20):
0.067
Positive Predictive Value
MXScarna(20):
0.710
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Murlet(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Murlet(20):
0.230
Sensitivity
CRWrnafold:
0.275
Murlet(20):
0.067
Positive Predictive Value
CRWrnafold:
0.435
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Murlet(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Murlet(20):
0.230
Sensitivity
RNAsubopt:
0.217
Murlet(20):
0.067
Positive Predictive Value
RNAsubopt:
0.521
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Murlet(20)
Matthews Correlation Coefficient
McQFold:
0.314
Murlet(20):
0.230
Sensitivity
McQFold:
0.205
Murlet(20):
0.067
Positive Predictive Value
McQFold:
0.480
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Murlet(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Murlet(20):
0.230
Sensitivity
TurboFold(20):
0.108
Murlet(20):
0.067
Positive Predictive Value
TurboFold(20):
0.837
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Murlet(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Murlet(20):
0.230
Sensitivity
PPfold(20):
0.101
Murlet(20):
0.067
Positive Predictive Value
PPfold(20):
0.879
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Murlet(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Murlet(20):
0.230
Sensitivity
RNAshapes:
0.180
Murlet(20):
0.067
Positive Predictive Value
RNAshapes:
0.484
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Murlet(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
Murlet(20):
0.230
Sensitivity
Carnac(20):
0.093
Murlet(20):
0.067
Positive Predictive Value
Carnac(20):
0.892
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Murlet(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Murlet(20):
0.230
Sensitivity
RNASLOpt:
0.158
Murlet(20):
0.067
Positive Predictive Value
RNASLOpt:
0.502
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Murlet(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Murlet(20):
0.230
Sensitivity
RSpredict(20):
0.124
Murlet(20):
0.067
Positive Predictive Value
RSpredict(20):
0.590
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Murlet(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
Murlet(20):
0.230
Sensitivity
Multilign(20):
0.083
Murlet(20):
0.067
Positive Predictive Value
Multilign(20):
0.762
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Murlet(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
Murlet(20):
0.230
Sensitivity
RNAwolf:
0.191
Murlet(20):
0.067
Positive Predictive Value
RNAwolf:
0.330
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Murlet(20) vs Vsfold4
Matthews Correlation Coefficient
Murlet(20):
0.230
Vsfold4:
0.217
Sensitivity
Murlet(20):
0.067
Vsfold4:
0.125
Positive Predictive Value
Murlet(20):
0.787
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Vsfold5
Matthews Correlation Coefficient
Murlet(20):
0.230
Vsfold5:
0.211
Sensitivity
Murlet(20):
0.067
Vsfold5:
0.125
Positive Predictive Value
Murlet(20):
0.787
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
RSpredict(seed):
0.212
Sensitivity
Murlet(20):
0.067
RSpredict(seed):
0.075
Positive Predictive Value
Murlet(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs CMfinder(20)
Matthews Correlation Coefficient
Murlet(20):
0.230
CMfinder(20):
0.210
Sensitivity
Murlet(20):
0.067
CMfinder(20):
0.055
Positive Predictive Value
Murlet(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs HotKnots
Matthews Correlation Coefficient
Murlet(20):
0.230
HotKnots:
0.179
Sensitivity
Murlet(20):
0.067
HotKnots:
0.051
Positive Predictive Value
Murlet(20):
0.787
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RNASampler(20)
Matthews Correlation Coefficient
Murlet(20):
0.230
RNASampler(20):
0.176
Sensitivity
Murlet(20):
0.067
RNASampler(20):
0.035
Positive Predictive Value
Murlet(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Cylofold
Matthews Correlation Coefficient
Murlet(20):
0.230
Cylofold:
0.155
Sensitivity
Murlet(20):
0.067
Cylofold:
0.040
Positive Predictive Value
Murlet(20):
0.787
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Pknots
Matthews Correlation Coefficient
Murlet(20):
0.230
Pknots:
0.143
Sensitivity
Murlet(20):
0.067
Pknots:
0.033
Positive Predictive Value
Murlet(20):
0.787
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Mastr(20)
Matthews Correlation Coefficient
Murlet(20):
0.230
Mastr(20):
0.134
Sensitivity
Murlet(20):
0.067
Mastr(20):
0.023
Positive Predictive Value
Murlet(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Alterna
Matthews Correlation Coefficient
Murlet(20):
0.230
Alterna:
0.129
Sensitivity
Murlet(20):
0.067
Alterna:
0.030
Positive Predictive Value
Murlet(20):
0.787
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs MCFold
Matthews Correlation Coefficient
Murlet(20):
0.230
MCFold:
0.125
Sensitivity
Murlet(20):
0.067
MCFold:
0.036
Positive Predictive Value
Murlet(20):
0.787
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RDfolder
Matthews Correlation Coefficient
Murlet(20):
0.230
RDfolder:
0.122
Sensitivity
Murlet(20):
0.067
RDfolder:
0.025
Positive Predictive Value
Murlet(20):
0.787
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Murlet(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Murlet(seed):
0.079
Sensitivity
Murlet(20):
0.067
Murlet(seed):
0.008
Positive Predictive Value
Murlet(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
CMfinder(seed):
0.080
Sensitivity
Murlet(20):
0.067
CMfinder(seed):
0.008
Positive Predictive Value
Murlet(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
TurboFold(seed):
0.076
Sensitivity
Murlet(20):
0.067
TurboFold(seed):
0.007
Positive Predictive Value
Murlet(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
PPfold(seed):
0.069
Sensitivity
Murlet(20):
0.067
PPfold(seed):
0.006
Positive Predictive Value
Murlet(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Carnac(seed):
0.066
Sensitivity
Murlet(20):
0.067
Carnac(seed):
0.005
Positive Predictive Value
Murlet(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
RNASampler(seed):
0.063
Sensitivity
Murlet(20):
0.067
RNASampler(seed):
0.005
Positive Predictive Value
Murlet(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs NanoFolder
Matthews Correlation Coefficient
Murlet(20):
0.230
NanoFolder:
0.052
Sensitivity
Murlet(20):
0.067
NanoFolder:
0.016
Positive Predictive Value
Murlet(20):
0.787
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Mastr(seed):
0.051
Sensitivity
Murlet(20):
0.067
Mastr(seed):
0.003
Positive Predictive Value
Murlet(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Multilign(seed):
0.043
Sensitivity
Murlet(20):
0.067
Multilign(seed):
0.003
Positive Predictive Value
Murlet(20):
0.787
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Vsfold4 |
1987
ContextFold vs Vsfold4
Matthews Correlation Coefficient
ContextFold:
0.680
Vsfold4:
0.217
Sensitivity
ContextFold:
0.644
Vsfold4:
0.125
Positive Predictive Value
ContextFold:
0.718
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Vsfold4
Matthews Correlation Coefficient
IPknot:
0.548
Vsfold4:
0.217
Sensitivity
IPknot:
0.475
Vsfold4:
0.125
Positive Predictive Value
IPknot:
0.632
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Vsfold4:
0.217
Sensitivity
PETfold_pre2.0(seed):
0.343
Vsfold4:
0.125
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Vsfold4
Matthews Correlation Coefficient
Contrafold:
0.508
Vsfold4:
0.217
Sensitivity
Contrafold:
0.481
Vsfold4:
0.125
Positive Predictive Value
Contrafold:
0.537
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Vsfold4
Matthews Correlation Coefficient
CentroidFold:
0.470
Vsfold4:
0.217
Sensitivity
CentroidFold:
0.369
Vsfold4:
0.125
Positive Predictive Value
CentroidFold:
0.600
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Vsfold4
Matthews Correlation Coefficient
Sfold:
0.469
Vsfold4:
0.217
Sensitivity
Sfold:
0.412
Vsfold4:
0.125
Positive Predictive Value
Sfold:
0.535
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Vsfold4
Matthews Correlation Coefficient
MaxExpect:
0.465
Vsfold4:
0.217
Sensitivity
MaxExpect:
0.443
Vsfold4:
0.125
Positive Predictive Value
MaxExpect:
0.488
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Vsfold4
Matthews Correlation Coefficient
ProbKnot:
0.460
Vsfold4:
0.217
Sensitivity
ProbKnot:
0.446
Vsfold4:
0.125
Positive Predictive Value
ProbKnot:
0.475
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Vsfold4
Matthews Correlation Coefficient
UNAFold:
0.445
Vsfold4:
0.217
Sensitivity
UNAFold:
0.445
Vsfold4:
0.125
Positive Predictive Value
UNAFold:
0.446
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Vsfold4:
0.217
Sensitivity
CentroidAlifold(seed):
0.213
Vsfold4:
0.125
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Vsfold4
Matthews Correlation Coefficient
Fold:
0.437
Vsfold4:
0.217
Sensitivity
Fold:
0.432
Vsfold4:
0.125
Positive Predictive Value
Fold:
0.444
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Vsfold4
Matthews Correlation Coefficient
RNAfold:
0.429
Vsfold4:
0.217
Sensitivity
RNAfold:
0.422
Vsfold4:
0.125
Positive Predictive Value
RNAfold:
0.437
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Vsfold4
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Vsfold4:
0.217
Sensitivity
CentroidHomfold‑LAST:
0.216
Vsfold4:
0.125
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Vsfold4:
0.217
Sensitivity
PETfold_pre2.0(20):
0.211
Vsfold4:
0.125
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Vsfold4:
0.217
Sensitivity
CentroidAlifold(20):
0.190
Vsfold4:
0.125
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Vsfold4
Matthews Correlation Coefficient
PknotsRG:
0.386
Vsfold4:
0.217
Sensitivity
PknotsRG:
0.320
Vsfold4:
0.125
Positive Predictive Value
PknotsRG:
0.465
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Vsfold4:
0.217
Sensitivity
MXScarna(seed):
0.196
Vsfold4:
0.125
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Vsfold4:
0.217
Sensitivity
RNAalifold(20):
0.185
Vsfold4:
0.125
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Vsfold4:
0.217
Sensitivity
RNAalifold(seed):
0.169
Vsfold4:
0.125
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Vsfold4
Matthews Correlation Coefficient
Afold:
0.358
Vsfold4:
0.217
Sensitivity
Afold:
0.295
Vsfold4:
0.125
Positive Predictive Value
Afold:
0.434
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(20):
0.359
Vsfold4:
0.217
Sensitivity
MXScarna(20):
0.181
Vsfold4:
0.125
Positive Predictive Value
MXScarna(20):
0.710
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Vsfold4
Matthews Correlation Coefficient
CRWrnafold:
0.346
Vsfold4:
0.217
Sensitivity
CRWrnafold:
0.275
Vsfold4:
0.125
Positive Predictive Value
CRWrnafold:
0.435
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Vsfold4
Matthews Correlation Coefficient
RNAsubopt:
0.336
Vsfold4:
0.217
Sensitivity
RNAsubopt:
0.217
Vsfold4:
0.125
Positive Predictive Value
RNAsubopt:
0.521
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Vsfold4
Matthews Correlation Coefficient
McQFold:
0.314
Vsfold4:
0.217
Sensitivity
McQFold:
0.205
Vsfold4:
0.125
Positive Predictive Value
McQFold:
0.480
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Vsfold4
Matthews Correlation Coefficient
TurboFold(20):
0.301
Vsfold4:
0.217
Sensitivity
TurboFold(20):
0.108
Vsfold4:
0.125
Positive Predictive Value
TurboFold(20):
0.837
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Vsfold4
Matthews Correlation Coefficient
PPfold(20):
0.298
Vsfold4:
0.217
Sensitivity
PPfold(20):
0.101
Vsfold4:
0.125
Positive Predictive Value
PPfold(20):
0.879
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Vsfold4
Matthews Correlation Coefficient
RNAshapes:
0.295
Vsfold4:
0.217
Sensitivity
RNAshapes:
0.180
Vsfold4:
0.125
Positive Predictive Value
RNAshapes:
0.484
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Vsfold4
Matthews Correlation Coefficient
Carnac(20):
0.288
Vsfold4:
0.217
Sensitivity
Carnac(20):
0.093
Vsfold4:
0.125
Positive Predictive Value
Carnac(20):
0.892
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Vsfold4
Matthews Correlation Coefficient
RNASLOpt:
0.281
Vsfold4:
0.217
Sensitivity
RNASLOpt:
0.158
Vsfold4:
0.125
Positive Predictive Value
RNASLOpt:
0.502
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Vsfold4
Matthews Correlation Coefficient
RSpredict(20):
0.270
Vsfold4:
0.217
Sensitivity
RSpredict(20):
0.124
Vsfold4:
0.125
Positive Predictive Value
RSpredict(20):
0.590
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Vsfold4
Matthews Correlation Coefficient
Multilign(20):
0.251
Vsfold4:
0.217
Sensitivity
Multilign(20):
0.083
Vsfold4:
0.125
Positive Predictive Value
Multilign(20):
0.762
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Vsfold4
Matthews Correlation Coefficient
RNAwolf:
0.251
Vsfold4:
0.217
Sensitivity
RNAwolf:
0.191
Vsfold4:
0.125
Positive Predictive Value
RNAwolf:
0.330
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Vsfold4
Matthews Correlation Coefficient
Murlet(20):
0.230
Vsfold4:
0.217
Sensitivity
Murlet(20):
0.067
Vsfold4:
0.125
Positive Predictive Value
Murlet(20):
0.787
Vsfold4:
0.378
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Vsfold4 vs Vsfold5
Matthews Correlation Coefficient
Vsfold4:
0.217
Vsfold5:
0.211
Sensitivity
Vsfold4:
0.125
Vsfold5:
0.125
Positive Predictive Value
Vsfold4:
0.378
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RSpredict(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
RSpredict(seed):
0.212
Sensitivity
Vsfold4:
0.125
RSpredict(seed):
0.075
Positive Predictive Value
Vsfold4:
0.378
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
Vsfold4 vs CMfinder(20)
Matthews Correlation Coefficient
Vsfold4:
0.217
CMfinder(20):
0.210
Sensitivity
Vsfold4:
0.125
CMfinder(20):
0.055
Positive Predictive Value
Vsfold4:
0.378
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs HotKnots
Matthews Correlation Coefficient
Vsfold4:
0.217
HotKnots:
0.179
Sensitivity
Vsfold4:
0.125
HotKnots:
0.051
Positive Predictive Value
Vsfold4:
0.378
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold4:
0.217
RNASampler(20):
0.176
Sensitivity
Vsfold4:
0.125
RNASampler(20):
0.035
Positive Predictive Value
Vsfold4:
0.378
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Cylofold
Matthews Correlation Coefficient
Vsfold4:
0.217
Cylofold:
0.155
Sensitivity
Vsfold4:
0.125
Cylofold:
0.040
Positive Predictive Value
Vsfold4:
0.378
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Pknots
Matthews Correlation Coefficient
Vsfold4:
0.217
Pknots:
0.143
Sensitivity
Vsfold4:
0.125
Pknots:
0.033
Positive Predictive Value
Vsfold4:
0.378
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold4:
0.217
Mastr(20):
0.134
Sensitivity
Vsfold4:
0.125
Mastr(20):
0.023
Positive Predictive Value
Vsfold4:
0.378
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Alterna
Matthews Correlation Coefficient
Vsfold4:
0.217
Alterna:
0.129
Sensitivity
Vsfold4:
0.125
Alterna:
0.030
Positive Predictive Value
Vsfold4:
0.378
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs MCFold
Matthews Correlation Coefficient
Vsfold4:
0.217
MCFold:
0.125
Sensitivity
Vsfold4:
0.125
MCFold:
0.036
Positive Predictive Value
Vsfold4:
0.378
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RDfolder
Matthews Correlation Coefficient
Vsfold4:
0.217
RDfolder:
0.122
Sensitivity
Vsfold4:
0.125
RDfolder:
0.025
Positive Predictive Value
Vsfold4:
0.378
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Murlet(seed):
0.079
Sensitivity
Vsfold4:
0.125
Murlet(seed):
0.008
Positive Predictive Value
Vsfold4:
0.378
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
CMfinder(seed):
0.080
Sensitivity
Vsfold4:
0.125
CMfinder(seed):
0.008
Positive Predictive Value
Vsfold4:
0.378
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
TurboFold(seed):
0.076
Sensitivity
Vsfold4:
0.125
TurboFold(seed):
0.007
Positive Predictive Value
Vsfold4:
0.378
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
PPfold(seed):
0.069
Sensitivity
Vsfold4:
0.125
PPfold(seed):
0.006
Positive Predictive Value
Vsfold4:
0.378
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Carnac(seed):
0.066
Sensitivity
Vsfold4:
0.125
Carnac(seed):
0.005
Positive Predictive Value
Vsfold4:
0.378
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
RNASampler(seed):
0.063
Sensitivity
Vsfold4:
0.125
RNASampler(seed):
0.005
Positive Predictive Value
Vsfold4:
0.378
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs NanoFolder
Matthews Correlation Coefficient
Vsfold4:
0.217
NanoFolder:
0.052
Sensitivity
Vsfold4:
0.125
NanoFolder:
0.016
Positive Predictive Value
Vsfold4:
0.378
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Mastr(seed):
0.051
Sensitivity
Vsfold4:
0.125
Mastr(seed):
0.003
Positive Predictive Value
Vsfold4:
0.378
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Multilign(seed):
0.043
Sensitivity
Vsfold4:
0.125
Multilign(seed):
0.003
Positive Predictive Value
Vsfold4:
0.378
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Vsfold5 |
1987
ContextFold vs Vsfold5
Matthews Correlation Coefficient
ContextFold:
0.680
Vsfold5:
0.211
Sensitivity
ContextFold:
0.644
Vsfold5:
0.125
Positive Predictive Value
ContextFold:
0.718
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Vsfold5
Matthews Correlation Coefficient
IPknot:
0.548
Vsfold5:
0.211
Sensitivity
IPknot:
0.475
Vsfold5:
0.125
Positive Predictive Value
IPknot:
0.632
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Vsfold5:
0.211
Sensitivity
PETfold_pre2.0(seed):
0.343
Vsfold5:
0.125
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Vsfold5
Matthews Correlation Coefficient
Contrafold:
0.508
Vsfold5:
0.211
Sensitivity
Contrafold:
0.481
Vsfold5:
0.125
Positive Predictive Value
Contrafold:
0.537
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Vsfold5
Matthews Correlation Coefficient
CentroidFold:
0.470
Vsfold5:
0.211
Sensitivity
CentroidFold:
0.369
Vsfold5:
0.125
Positive Predictive Value
CentroidFold:
0.600
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Vsfold5
Matthews Correlation Coefficient
Sfold:
0.469
Vsfold5:
0.211
Sensitivity
Sfold:
0.412
Vsfold5:
0.125
Positive Predictive Value
Sfold:
0.535
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Vsfold5
Matthews Correlation Coefficient
MaxExpect:
0.465
Vsfold5:
0.211
Sensitivity
MaxExpect:
0.443
Vsfold5:
0.125
Positive Predictive Value
MaxExpect:
0.488
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Vsfold5
Matthews Correlation Coefficient
ProbKnot:
0.460
Vsfold5:
0.211
Sensitivity
ProbKnot:
0.446
Vsfold5:
0.125
Positive Predictive Value
ProbKnot:
0.475
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Vsfold5
Matthews Correlation Coefficient
UNAFold:
0.445
Vsfold5:
0.211
Sensitivity
UNAFold:
0.445
Vsfold5:
0.125
Positive Predictive Value
UNAFold:
0.446
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Vsfold5:
0.211
Sensitivity
CentroidAlifold(seed):
0.213
Vsfold5:
0.125
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Vsfold5
Matthews Correlation Coefficient
Fold:
0.437
Vsfold5:
0.211
Sensitivity
Fold:
0.432
Vsfold5:
0.125
Positive Predictive Value
Fold:
0.444
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Vsfold5
Matthews Correlation Coefficient
RNAfold:
0.429
Vsfold5:
0.211
Sensitivity
RNAfold:
0.422
Vsfold5:
0.125
Positive Predictive Value
RNAfold:
0.437
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Vsfold5
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Vsfold5:
0.211
Sensitivity
CentroidHomfold‑LAST:
0.216
Vsfold5:
0.125
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Vsfold5:
0.211
Sensitivity
PETfold_pre2.0(20):
0.211
Vsfold5:
0.125
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Vsfold5:
0.211
Sensitivity
CentroidAlifold(20):
0.190
Vsfold5:
0.125
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Vsfold5
Matthews Correlation Coefficient
PknotsRG:
0.386
Vsfold5:
0.211
Sensitivity
PknotsRG:
0.320
Vsfold5:
0.125
Positive Predictive Value
PknotsRG:
0.465
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Vsfold5:
0.211
Sensitivity
MXScarna(seed):
0.196
Vsfold5:
0.125
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Vsfold5:
0.211
Sensitivity
RNAalifold(20):
0.185
Vsfold5:
0.125
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Vsfold5:
0.211
Sensitivity
RNAalifold(seed):
0.169
Vsfold5:
0.125
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Vsfold5
Matthews Correlation Coefficient
Afold:
0.358
Vsfold5:
0.211
Sensitivity
Afold:
0.295
Vsfold5:
0.125
Positive Predictive Value
Afold:
0.434
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(20):
0.359
Vsfold5:
0.211
Sensitivity
MXScarna(20):
0.181
Vsfold5:
0.125
Positive Predictive Value
MXScarna(20):
0.710
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Vsfold5
Matthews Correlation Coefficient
CRWrnafold:
0.346
Vsfold5:
0.211
Sensitivity
CRWrnafold:
0.275
Vsfold5:
0.125
Positive Predictive Value
CRWrnafold:
0.435
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Vsfold5
Matthews Correlation Coefficient
RNAsubopt:
0.336
Vsfold5:
0.211
Sensitivity
RNAsubopt:
0.217
Vsfold5:
0.125
Positive Predictive Value
RNAsubopt:
0.521
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Vsfold5
Matthews Correlation Coefficient
McQFold:
0.314
Vsfold5:
0.211
Sensitivity
McQFold:
0.205
Vsfold5:
0.125
Positive Predictive Value
McQFold:
0.480
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Vsfold5
Matthews Correlation Coefficient
TurboFold(20):
0.301
Vsfold5:
0.211
Sensitivity
TurboFold(20):
0.108
Vsfold5:
0.125
Positive Predictive Value
TurboFold(20):
0.837
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Vsfold5
Matthews Correlation Coefficient
PPfold(20):
0.298
Vsfold5:
0.211
Sensitivity
PPfold(20):
0.101
Vsfold5:
0.125
Positive Predictive Value
PPfold(20):
0.879
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Vsfold5
Matthews Correlation Coefficient
RNAshapes:
0.295
Vsfold5:
0.211
Sensitivity
RNAshapes:
0.180
Vsfold5:
0.125
Positive Predictive Value
RNAshapes:
0.484
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Vsfold5
Matthews Correlation Coefficient
Carnac(20):
0.288
Vsfold5:
0.211
Sensitivity
Carnac(20):
0.093
Vsfold5:
0.125
Positive Predictive Value
Carnac(20):
0.892
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Vsfold5
Matthews Correlation Coefficient
RNASLOpt:
0.281
Vsfold5:
0.211
Sensitivity
RNASLOpt:
0.158
Vsfold5:
0.125
Positive Predictive Value
RNASLOpt:
0.502
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(20):
0.270
Vsfold5:
0.211
Sensitivity
RSpredict(20):
0.124
Vsfold5:
0.125
Positive Predictive Value
RSpredict(20):
0.590
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Vsfold5
Matthews Correlation Coefficient
Multilign(20):
0.251
Vsfold5:
0.211
Sensitivity
Multilign(20):
0.083
Vsfold5:
0.125
Positive Predictive Value
Multilign(20):
0.762
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Vsfold5
Matthews Correlation Coefficient
RNAwolf:
0.251
Vsfold5:
0.211
Sensitivity
RNAwolf:
0.191
Vsfold5:
0.125
Positive Predictive Value
RNAwolf:
0.330
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Vsfold5
Matthews Correlation Coefficient
Murlet(20):
0.230
Vsfold5:
0.211
Sensitivity
Murlet(20):
0.067
Vsfold5:
0.125
Positive Predictive Value
Murlet(20):
0.787
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Vsfold5
Matthews Correlation Coefficient
Vsfold4:
0.217
Vsfold5:
0.211
Sensitivity
Vsfold4:
0.125
Vsfold5:
0.125
Positive Predictive Value
Vsfold4:
0.378
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
RSpredict(seed) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Vsfold5:
0.211
Sensitivity
RSpredict(seed):
0.075
Vsfold5:
0.125
Positive Predictive Value
RSpredict(seed):
0.597
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.103865234064
|
+
Vsfold5 vs CMfinder(20)
Matthews Correlation Coefficient
Vsfold5:
0.211
CMfinder(20):
0.210
Sensitivity
Vsfold5:
0.125
CMfinder(20):
0.055
Positive Predictive Value
Vsfold5:
0.359
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 1.37177523642e-07
|
+
Vsfold5 vs HotKnots
Matthews Correlation Coefficient
Vsfold5:
0.211
HotKnots:
0.179
Sensitivity
Vsfold5:
0.125
HotKnots:
0.051
Positive Predictive Value
Vsfold5:
0.359
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold5:
0.211
RNASampler(20):
0.176
Sensitivity
Vsfold5:
0.125
RNASampler(20):
0.035
Positive Predictive Value
Vsfold5:
0.359
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Cylofold
Matthews Correlation Coefficient
Vsfold5:
0.211
Cylofold:
0.155
Sensitivity
Vsfold5:
0.125
Cylofold:
0.040
Positive Predictive Value
Vsfold5:
0.359
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Pknots
Matthews Correlation Coefficient
Vsfold5:
0.211
Pknots:
0.143
Sensitivity
Vsfold5:
0.125
Pknots:
0.033
Positive Predictive Value
Vsfold5:
0.359
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold5:
0.211
Mastr(20):
0.134
Sensitivity
Vsfold5:
0.125
Mastr(20):
0.023
Positive Predictive Value
Vsfold5:
0.359
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Alterna
Matthews Correlation Coefficient
Vsfold5:
0.211
Alterna:
0.129
Sensitivity
Vsfold5:
0.125
Alterna:
0.030
Positive Predictive Value
Vsfold5:
0.359
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs MCFold
Matthews Correlation Coefficient
Vsfold5:
0.211
MCFold:
0.125
Sensitivity
Vsfold5:
0.125
MCFold:
0.036
Positive Predictive Value
Vsfold5:
0.359
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs RDfolder
Matthews Correlation Coefficient
Vsfold5:
0.211
RDfolder:
0.122
Sensitivity
Vsfold5:
0.125
RDfolder:
0.025
Positive Predictive Value
Vsfold5:
0.359
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Murlet(seed):
0.079
Sensitivity
Vsfold5:
0.125
Murlet(seed):
0.008
Positive Predictive Value
Vsfold5:
0.359
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
CMfinder(seed):
0.080
Sensitivity
Vsfold5:
0.125
CMfinder(seed):
0.008
Positive Predictive Value
Vsfold5:
0.359
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
TurboFold(seed):
0.076
Sensitivity
Vsfold5:
0.125
TurboFold(seed):
0.007
Positive Predictive Value
Vsfold5:
0.359
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
PPfold(seed):
0.069
Sensitivity
Vsfold5:
0.125
PPfold(seed):
0.006
Positive Predictive Value
Vsfold5:
0.359
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Carnac(seed):
0.066
Sensitivity
Vsfold5:
0.125
Carnac(seed):
0.005
Positive Predictive Value
Vsfold5:
0.359
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
RNASampler(seed):
0.063
Sensitivity
Vsfold5:
0.125
RNASampler(seed):
0.005
Positive Predictive Value
Vsfold5:
0.359
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs NanoFolder
Matthews Correlation Coefficient
Vsfold5:
0.211
NanoFolder:
0.052
Sensitivity
Vsfold5:
0.125
NanoFolder:
0.016
Positive Predictive Value
Vsfold5:
0.359
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Mastr(seed):
0.051
Sensitivity
Vsfold5:
0.125
Mastr(seed):
0.003
Positive Predictive Value
Vsfold5:
0.359
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Multilign(seed):
0.043
Sensitivity
Vsfold5:
0.125
Multilign(seed):
0.003
Positive Predictive Value
Vsfold5:
0.359
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RSpredict(seed) |
1987
ContextFold vs RSpredict(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
RSpredict(seed):
0.212
Sensitivity
ContextFold:
0.644
RSpredict(seed):
0.075
Positive Predictive Value
ContextFold:
0.718
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RSpredict(seed)
Matthews Correlation Coefficient
IPknot:
0.548
RSpredict(seed):
0.212
Sensitivity
IPknot:
0.475
RSpredict(seed):
0.075
Positive Predictive Value
IPknot:
0.632
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RSpredict(seed):
0.212
Sensitivity
PETfold_pre2.0(seed):
0.343
RSpredict(seed):
0.075
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RSpredict(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
RSpredict(seed):
0.212
Sensitivity
Contrafold:
0.481
RSpredict(seed):
0.075
Positive Predictive Value
Contrafold:
0.537
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
RSpredict(seed):
0.212
Sensitivity
CentroidFold:
0.369
RSpredict(seed):
0.075
Positive Predictive Value
CentroidFold:
0.600
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RSpredict(seed)
Matthews Correlation Coefficient
Sfold:
0.469
RSpredict(seed):
0.212
Sensitivity
Sfold:
0.412
RSpredict(seed):
0.075
Positive Predictive Value
Sfold:
0.535
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RSpredict(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
RSpredict(seed):
0.212
Sensitivity
MaxExpect:
0.443
RSpredict(seed):
0.075
Positive Predictive Value
MaxExpect:
0.488
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RSpredict(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
RSpredict(seed):
0.212
Sensitivity
ProbKnot:
0.446
RSpredict(seed):
0.075
Positive Predictive Value
ProbKnot:
0.475
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RSpredict(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
RSpredict(seed):
0.212
Sensitivity
UNAFold:
0.445
RSpredict(seed):
0.075
Positive Predictive Value
UNAFold:
0.446
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RSpredict(seed):
0.212
Sensitivity
CentroidAlifold(seed):
0.213
RSpredict(seed):
0.075
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RSpredict(seed)
Matthews Correlation Coefficient
Fold:
0.437
RSpredict(seed):
0.212
Sensitivity
Fold:
0.432
RSpredict(seed):
0.075
Positive Predictive Value
Fold:
0.444
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RSpredict(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
RSpredict(seed):
0.212
Sensitivity
RNAfold:
0.422
RSpredict(seed):
0.075
Positive Predictive Value
RNAfold:
0.437
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RSpredict(seed):
0.212
Sensitivity
CentroidHomfold‑LAST:
0.216
RSpredict(seed):
0.075
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RSpredict(seed):
0.212
Sensitivity
PETfold_pre2.0(20):
0.211
RSpredict(seed):
0.075
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RSpredict(seed):
0.212
Sensitivity
CentroidAlifold(20):
0.190
RSpredict(seed):
0.075
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RSpredict(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
RSpredict(seed):
0.212
Sensitivity
PknotsRG:
0.320
RSpredict(seed):
0.075
Positive Predictive Value
PknotsRG:
0.465
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RSpredict(seed):
0.212
Sensitivity
MXScarna(seed):
0.196
RSpredict(seed):
0.075
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RSpredict(seed):
0.212
Sensitivity
RNAalifold(20):
0.185
RSpredict(seed):
0.075
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RSpredict(seed):
0.212
Sensitivity
RNAalifold(seed):
0.169
RSpredict(seed):
0.075
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RSpredict(seed)
Matthews Correlation Coefficient
Afold:
0.358
RSpredict(seed):
0.212
Sensitivity
Afold:
0.295
RSpredict(seed):
0.075
Positive Predictive Value
Afold:
0.434
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RSpredict(seed):
0.212
Sensitivity
MXScarna(20):
0.181
RSpredict(seed):
0.075
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RSpredict(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RSpredict(seed):
0.212
Sensitivity
CRWrnafold:
0.275
RSpredict(seed):
0.075
Positive Predictive Value
CRWrnafold:
0.435
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RSpredict(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RSpredict(seed):
0.212
Sensitivity
RNAsubopt:
0.217
RSpredict(seed):
0.075
Positive Predictive Value
RNAsubopt:
0.521
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RSpredict(seed)
Matthews Correlation Coefficient
McQFold:
0.314
RSpredict(seed):
0.212
Sensitivity
McQFold:
0.205
RSpredict(seed):
0.075
Positive Predictive Value
McQFold:
0.480
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RSpredict(seed):
0.212
Sensitivity
TurboFold(20):
0.108
RSpredict(seed):
0.075
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
RSpredict(seed):
0.212
Sensitivity
PPfold(20):
0.101
RSpredict(seed):
0.075
Positive Predictive Value
PPfold(20):
0.879
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RSpredict(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
RSpredict(seed):
0.212
Sensitivity
RNAshapes:
0.180
RSpredict(seed):
0.075
Positive Predictive Value
RNAshapes:
0.484
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
RSpredict(seed):
0.212
Sensitivity
Carnac(20):
0.093
RSpredict(seed):
0.075
Positive Predictive Value
Carnac(20):
0.892
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs RSpredict(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RSpredict(seed):
0.212
Sensitivity
RNASLOpt:
0.158
RSpredict(seed):
0.075
Positive Predictive Value
RNASLOpt:
0.502
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
RSpredict(seed):
0.212
Sensitivity
RSpredict(20):
0.124
RSpredict(seed):
0.075
Positive Predictive Value
RSpredict(20):
0.590
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
RSpredict(seed):
0.212
Sensitivity
Multilign(20):
0.083
RSpredict(seed):
0.075
Positive Predictive Value
Multilign(20):
0.762
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs RSpredict(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
RSpredict(seed):
0.212
Sensitivity
RNAwolf:
0.191
RSpredict(seed):
0.075
Positive Predictive Value
RNAwolf:
0.330
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
RSpredict(seed):
0.212
Sensitivity
Murlet(20):
0.067
RSpredict(seed):
0.075
Positive Predictive Value
Murlet(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs RSpredict(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
RSpredict(seed):
0.212
Sensitivity
Vsfold4:
0.125
RSpredict(seed):
0.075
Positive Predictive Value
Vsfold4:
0.378
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
1987
RSpredict(seed) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Vsfold5:
0.211
Sensitivity
RSpredict(seed):
0.075
Vsfold5:
0.125
Positive Predictive Value
RSpredict(seed):
0.597
Vsfold5:
0.359
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.103865234064
|
|
=
RSpredict(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
CMfinder(20):
0.210
Sensitivity
RSpredict(seed):
0.075
CMfinder(20):
0.055
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.00108987668019
|
+
RSpredict(seed) vs HotKnots
Matthews Correlation Coefficient
RSpredict(seed):
0.212
HotKnots:
0.179
Sensitivity
RSpredict(seed):
0.075
HotKnots:
0.051
Positive Predictive Value
RSpredict(seed):
0.597
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
RNASampler(20):
0.176
Sensitivity
RSpredict(seed):
0.075
RNASampler(20):
0.035
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Cylofold
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Cylofold:
0.155
Sensitivity
RSpredict(seed):
0.075
Cylofold:
0.040
Positive Predictive Value
RSpredict(seed):
0.597
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Pknots
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Pknots:
0.143
Sensitivity
RSpredict(seed):
0.075
Pknots:
0.033
Positive Predictive Value
RSpredict(seed):
0.597
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Mastr(20):
0.134
Sensitivity
RSpredict(seed):
0.075
Mastr(20):
0.023
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Alterna
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Alterna:
0.129
Sensitivity
RSpredict(seed):
0.075
Alterna:
0.030
Positive Predictive Value
RSpredict(seed):
0.597
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs MCFold
Matthews Correlation Coefficient
RSpredict(seed):
0.212
MCFold:
0.125
Sensitivity
RSpredict(seed):
0.075
MCFold:
0.036
Positive Predictive Value
RSpredict(seed):
0.597
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs RDfolder
Matthews Correlation Coefficient
RSpredict(seed):
0.212
RDfolder:
0.122
Sensitivity
RSpredict(seed):
0.075
RDfolder:
0.025
Positive Predictive Value
RSpredict(seed):
0.597
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Murlet(seed):
0.079
Sensitivity
RSpredict(seed):
0.075
Murlet(seed):
0.008
Positive Predictive Value
RSpredict(seed):
0.597
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
CMfinder(seed):
0.080
Sensitivity
RSpredict(seed):
0.075
CMfinder(seed):
0.008
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
TurboFold(seed):
0.076
Sensitivity
RSpredict(seed):
0.075
TurboFold(seed):
0.007
Positive Predictive Value
RSpredict(seed):
0.597
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
PPfold(seed):
0.069
Sensitivity
RSpredict(seed):
0.075
PPfold(seed):
0.006
Positive Predictive Value
RSpredict(seed):
0.597
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Carnac(seed):
0.066
Sensitivity
RSpredict(seed):
0.075
Carnac(seed):
0.005
Positive Predictive Value
RSpredict(seed):
0.597
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
RNASampler(seed):
0.063
Sensitivity
RSpredict(seed):
0.075
RNASampler(seed):
0.005
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(seed):
0.212
NanoFolder:
0.052
Sensitivity
RSpredict(seed):
0.075
NanoFolder:
0.016
Positive Predictive Value
RSpredict(seed):
0.597
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Mastr(seed):
0.051
Sensitivity
RSpredict(seed):
0.075
Mastr(seed):
0.003
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Multilign(seed):
0.043
Sensitivity
RSpredict(seed):
0.075
Multilign(seed):
0.003
Positive Predictive Value
RSpredict(seed):
0.597
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CMfinder(20) |
1987
ContextFold vs CMfinder(20)
Matthews Correlation Coefficient
ContextFold:
0.680
CMfinder(20):
0.210
Sensitivity
ContextFold:
0.644
CMfinder(20):
0.055
Positive Predictive Value
ContextFold:
0.718
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CMfinder(20)
Matthews Correlation Coefficient
IPknot:
0.548
CMfinder(20):
0.210
Sensitivity
IPknot:
0.475
CMfinder(20):
0.055
Positive Predictive Value
IPknot:
0.632
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CMfinder(20):
0.210
Sensitivity
PETfold_pre2.0(seed):
0.343
CMfinder(20):
0.055
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CMfinder(20)
Matthews Correlation Coefficient
Contrafold:
0.508
CMfinder(20):
0.210
Sensitivity
Contrafold:
0.481
CMfinder(20):
0.055
Positive Predictive Value
Contrafold:
0.537
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs CMfinder(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
CMfinder(20):
0.210
Sensitivity
CentroidFold:
0.369
CMfinder(20):
0.055
Positive Predictive Value
CentroidFold:
0.600
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs CMfinder(20)
Matthews Correlation Coefficient
Sfold:
0.469
CMfinder(20):
0.210
Sensitivity
Sfold:
0.412
CMfinder(20):
0.055
Positive Predictive Value
Sfold:
0.535
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs CMfinder(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
CMfinder(20):
0.210
Sensitivity
MaxExpect:
0.443
CMfinder(20):
0.055
Positive Predictive Value
MaxExpect:
0.488
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs CMfinder(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
CMfinder(20):
0.210
Sensitivity
ProbKnot:
0.446
CMfinder(20):
0.055
Positive Predictive Value
ProbKnot:
0.475
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs CMfinder(20)
Matthews Correlation Coefficient
UNAFold:
0.445
CMfinder(20):
0.210
Sensitivity
UNAFold:
0.445
CMfinder(20):
0.055
Positive Predictive Value
UNAFold:
0.446
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CMfinder(20):
0.210
Sensitivity
CentroidAlifold(seed):
0.213
CMfinder(20):
0.055
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs CMfinder(20)
Matthews Correlation Coefficient
Fold:
0.437
CMfinder(20):
0.210
Sensitivity
Fold:
0.432
CMfinder(20):
0.055
Positive Predictive Value
Fold:
0.444
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs CMfinder(20)
Matthews Correlation Coefficient
RNAfold:
0.429
CMfinder(20):
0.210
Sensitivity
RNAfold:
0.422
CMfinder(20):
0.055
Positive Predictive Value
RNAfold:
0.437
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs CMfinder(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CMfinder(20):
0.210
Sensitivity
CentroidHomfold‑LAST:
0.216
CMfinder(20):
0.055
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CMfinder(20):
0.210
Sensitivity
PETfold_pre2.0(20):
0.211
CMfinder(20):
0.055
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
CMfinder(20):
0.210
Sensitivity
CentroidAlifold(20):
0.190
CMfinder(20):
0.055
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs CMfinder(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
CMfinder(20):
0.210
Sensitivity
PknotsRG:
0.320
CMfinder(20):
0.055
Positive Predictive Value
PknotsRG:
0.465
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
CMfinder(20):
0.210
Sensitivity
MXScarna(seed):
0.196
CMfinder(20):
0.055
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
CMfinder(20):
0.210
Sensitivity
RNAalifold(20):
0.185
CMfinder(20):
0.055
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
CMfinder(20):
0.210
Sensitivity
RNAalifold(seed):
0.169
CMfinder(20):
0.055
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs CMfinder(20)
Matthews Correlation Coefficient
Afold:
0.358
CMfinder(20):
0.210
Sensitivity
Afold:
0.295
CMfinder(20):
0.055
Positive Predictive Value
Afold:
0.434
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
CMfinder(20):
0.210
Sensitivity
MXScarna(20):
0.181
CMfinder(20):
0.055
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs CMfinder(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
CMfinder(20):
0.210
Sensitivity
CRWrnafold:
0.275
CMfinder(20):
0.055
Positive Predictive Value
CRWrnafold:
0.435
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs CMfinder(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
CMfinder(20):
0.210
Sensitivity
RNAsubopt:
0.217
CMfinder(20):
0.055
Positive Predictive Value
RNAsubopt:
0.521
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs CMfinder(20)
Matthews Correlation Coefficient
McQFold:
0.314
CMfinder(20):
0.210
Sensitivity
McQFold:
0.205
CMfinder(20):
0.055
Positive Predictive Value
McQFold:
0.480
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs CMfinder(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
CMfinder(20):
0.210
Sensitivity
TurboFold(20):
0.108
CMfinder(20):
0.055
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs CMfinder(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
CMfinder(20):
0.210
Sensitivity
PPfold(20):
0.101
CMfinder(20):
0.055
Positive Predictive Value
PPfold(20):
0.879
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs CMfinder(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
CMfinder(20):
0.210
Sensitivity
RNAshapes:
0.180
CMfinder(20):
0.055
Positive Predictive Value
RNAshapes:
0.484
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs CMfinder(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
CMfinder(20):
0.210
Sensitivity
Carnac(20):
0.093
CMfinder(20):
0.055
Positive Predictive Value
Carnac(20):
0.892
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs CMfinder(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
CMfinder(20):
0.210
Sensitivity
RNASLOpt:
0.158
CMfinder(20):
0.055
Positive Predictive Value
RNASLOpt:
0.502
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
CMfinder(20):
0.210
Sensitivity
RSpredict(20):
0.124
CMfinder(20):
0.055
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs CMfinder(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
CMfinder(20):
0.210
Sensitivity
Multilign(20):
0.083
CMfinder(20):
0.055
Positive Predictive Value
Multilign(20):
0.762
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs CMfinder(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
CMfinder(20):
0.210
Sensitivity
RNAwolf:
0.191
CMfinder(20):
0.055
Positive Predictive Value
RNAwolf:
0.330
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs CMfinder(20)
Matthews Correlation Coefficient
Murlet(20):
0.230
CMfinder(20):
0.210
Sensitivity
Murlet(20):
0.067
CMfinder(20):
0.055
Positive Predictive Value
Murlet(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs CMfinder(20)
Matthews Correlation Coefficient
Vsfold4:
0.217
CMfinder(20):
0.210
Sensitivity
Vsfold4:
0.125
CMfinder(20):
0.055
Positive Predictive Value
Vsfold4:
0.378
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs CMfinder(20)
Matthews Correlation Coefficient
Vsfold5:
0.211
CMfinder(20):
0.210
Sensitivity
Vsfold5:
0.125
CMfinder(20):
0.055
Positive Predictive Value
Vsfold5:
0.359
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 1.37177523642e-07
|
1987
RSpredict(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
CMfinder(20):
0.210
Sensitivity
RSpredict(seed):
0.075
CMfinder(20):
0.055
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.00108987668019
|
|
+
CMfinder(20) vs HotKnots
Matthews Correlation Coefficient
CMfinder(20):
0.210
HotKnots:
0.179
Sensitivity
CMfinder(20):
0.055
HotKnots:
0.051
Positive Predictive Value
CMfinder(20):
0.794
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs RNASampler(20)
Matthews Correlation Coefficient
CMfinder(20):
0.210
RNASampler(20):
0.176
Sensitivity
CMfinder(20):
0.055
RNASampler(20):
0.035
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Cylofold
Matthews Correlation Coefficient
CMfinder(20):
0.210
Cylofold:
0.155
Sensitivity
CMfinder(20):
0.055
Cylofold:
0.040
Positive Predictive Value
CMfinder(20):
0.794
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Pknots
Matthews Correlation Coefficient
CMfinder(20):
0.210
Pknots:
0.143
Sensitivity
CMfinder(20):
0.055
Pknots:
0.033
Positive Predictive Value
CMfinder(20):
0.794
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Mastr(20)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Mastr(20):
0.134
Sensitivity
CMfinder(20):
0.055
Mastr(20):
0.023
Positive Predictive Value
CMfinder(20):
0.794
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Alterna
Matthews Correlation Coefficient
CMfinder(20):
0.210
Alterna:
0.129
Sensitivity
CMfinder(20):
0.055
Alterna:
0.030
Positive Predictive Value
CMfinder(20):
0.794
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs MCFold
Matthews Correlation Coefficient
CMfinder(20):
0.210
MCFold:
0.125
Sensitivity
CMfinder(20):
0.055
MCFold:
0.036
Positive Predictive Value
CMfinder(20):
0.794
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs RDfolder
Matthews Correlation Coefficient
CMfinder(20):
0.210
RDfolder:
0.122
Sensitivity
CMfinder(20):
0.055
RDfolder:
0.025
Positive Predictive Value
CMfinder(20):
0.794
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Murlet(seed):
0.079
Sensitivity
CMfinder(20):
0.055
Murlet(seed):
0.008
Positive Predictive Value
CMfinder(20):
0.794
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
CMfinder(seed):
0.080
Sensitivity
CMfinder(20):
0.055
CMfinder(seed):
0.008
Positive Predictive Value
CMfinder(20):
0.794
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
TurboFold(seed):
0.076
Sensitivity
CMfinder(20):
0.055
TurboFold(seed):
0.007
Positive Predictive Value
CMfinder(20):
0.794
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
PPfold(seed):
0.069
Sensitivity
CMfinder(20):
0.055
PPfold(seed):
0.006
Positive Predictive Value
CMfinder(20):
0.794
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Carnac(seed):
0.066
Sensitivity
CMfinder(20):
0.055
Carnac(seed):
0.005
Positive Predictive Value
CMfinder(20):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
RNASampler(seed):
0.063
Sensitivity
CMfinder(20):
0.055
RNASampler(seed):
0.005
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(20):
0.210
NanoFolder:
0.052
Sensitivity
CMfinder(20):
0.055
NanoFolder:
0.016
Positive Predictive Value
CMfinder(20):
0.794
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Mastr(seed):
0.051
Sensitivity
CMfinder(20):
0.055
Mastr(seed):
0.003
Positive Predictive Value
CMfinder(20):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Multilign(seed):
0.043
Sensitivity
CMfinder(20):
0.055
Multilign(seed):
0.003
Positive Predictive Value
CMfinder(20):
0.794
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| HotKnots |
1987
ContextFold vs HotKnots
Matthews Correlation Coefficient
ContextFold:
0.680
HotKnots:
0.179
Sensitivity
ContextFold:
0.644
HotKnots:
0.051
Positive Predictive Value
ContextFold:
0.718
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs HotKnots
Matthews Correlation Coefficient
IPknot:
0.548
HotKnots:
0.179
Sensitivity
IPknot:
0.475
HotKnots:
0.051
Positive Predictive Value
IPknot:
0.632
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
HotKnots:
0.179
Sensitivity
PETfold_pre2.0(seed):
0.343
HotKnots:
0.051
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs HotKnots
Matthews Correlation Coefficient
Contrafold:
0.508
HotKnots:
0.179
Sensitivity
Contrafold:
0.481
HotKnots:
0.051
Positive Predictive Value
Contrafold:
0.537
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs HotKnots
Matthews Correlation Coefficient
CentroidFold:
0.470
HotKnots:
0.179
Sensitivity
CentroidFold:
0.369
HotKnots:
0.051
Positive Predictive Value
CentroidFold:
0.600
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs HotKnots
Matthews Correlation Coefficient
Sfold:
0.469
HotKnots:
0.179
Sensitivity
Sfold:
0.412
HotKnots:
0.051
Positive Predictive Value
Sfold:
0.535
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs HotKnots
Matthews Correlation Coefficient
MaxExpect:
0.465
HotKnots:
0.179
Sensitivity
MaxExpect:
0.443
HotKnots:
0.051
Positive Predictive Value
MaxExpect:
0.488
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs HotKnots
Matthews Correlation Coefficient
ProbKnot:
0.460
HotKnots:
0.179
Sensitivity
ProbKnot:
0.446
HotKnots:
0.051
Positive Predictive Value
ProbKnot:
0.475
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs HotKnots
Matthews Correlation Coefficient
UNAFold:
0.445
HotKnots:
0.179
Sensitivity
UNAFold:
0.445
HotKnots:
0.051
Positive Predictive Value
UNAFold:
0.446
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
HotKnots:
0.179
Sensitivity
CentroidAlifold(seed):
0.213
HotKnots:
0.051
Positive Predictive Value
CentroidAlifold(seed):
0.901
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs HotKnots
Matthews Correlation Coefficient
Fold:
0.437
HotKnots:
0.179
Sensitivity
Fold:
0.432
HotKnots:
0.051
Positive Predictive Value
Fold:
0.444
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs HotKnots
Matthews Correlation Coefficient
RNAfold:
0.429
HotKnots:
0.179
Sensitivity
RNAfold:
0.422
HotKnots:
0.051
Positive Predictive Value
RNAfold:
0.437
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs HotKnots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
HotKnots:
0.179
Sensitivity
CentroidHomfold‑LAST:
0.216
HotKnots:
0.051
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
HotKnots:
0.179
Sensitivity
PETfold_pre2.0(20):
0.211
HotKnots:
0.051
Positive Predictive Value
PETfold_pre2.0(20):
0.807
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
HotKnots:
0.179
Sensitivity
CentroidAlifold(20):
0.190
HotKnots:
0.051
Positive Predictive Value
CentroidAlifold(20):
0.891
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs HotKnots
Matthews Correlation Coefficient
PknotsRG:
0.386
HotKnots:
0.179
Sensitivity
PknotsRG:
0.320
HotKnots:
0.051
Positive Predictive Value
PknotsRG:
0.465
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs HotKnots
Matthews Correlation Coefficient
MXScarna(seed):
0.385
HotKnots:
0.179
Sensitivity
MXScarna(seed):
0.196
HotKnots:
0.051
Positive Predictive Value
MXScarna(seed):
0.753
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(20):
0.381
HotKnots:
0.179
Sensitivity
RNAalifold(20):
0.185
HotKnots:
0.051
Positive Predictive Value
RNAalifold(20):
0.787
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
HotKnots:
0.179
Sensitivity
RNAalifold(seed):
0.169
HotKnots:
0.051
Positive Predictive Value
RNAalifold(seed):
0.835
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs HotKnots
Matthews Correlation Coefficient
Afold:
0.358
HotKnots:
0.179
Sensitivity
Afold:
0.295
HotKnots:
0.051
Positive Predictive Value
Afold:
0.434
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs HotKnots
Matthews Correlation Coefficient
MXScarna(20):
0.359
HotKnots:
0.179
Sensitivity
MXScarna(20):
0.181
HotKnots:
0.051
Positive Predictive Value
MXScarna(20):
0.710
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs HotKnots
Matthews Correlation Coefficient
CRWrnafold:
0.346
HotKnots:
0.179
Sensitivity
CRWrnafold:
0.275
HotKnots:
0.051
Positive Predictive Value
CRWrnafold:
0.435
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs HotKnots
Matthews Correlation Coefficient
RNAsubopt:
0.336
HotKnots:
0.179
Sensitivity
RNAsubopt:
0.217
HotKnots:
0.051
Positive Predictive Value
RNAsubopt:
0.521
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs HotKnots
Matthews Correlation Coefficient
McQFold:
0.314
HotKnots:
0.179
Sensitivity
McQFold:
0.205
HotKnots:
0.051
Positive Predictive Value
McQFold:
0.480
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs HotKnots
Matthews Correlation Coefficient
TurboFold(20):
0.301
HotKnots:
0.179
Sensitivity
TurboFold(20):
0.108
HotKnots:
0.051
Positive Predictive Value
TurboFold(20):
0.837
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs HotKnots
Matthews Correlation Coefficient
PPfold(20):
0.298
HotKnots:
0.179
Sensitivity
PPfold(20):
0.101
HotKnots:
0.051
Positive Predictive Value
PPfold(20):
0.879
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs HotKnots
Matthews Correlation Coefficient
RNAshapes:
0.295
HotKnots:
0.179
Sensitivity
RNAshapes:
0.180
HotKnots:
0.051
Positive Predictive Value
RNAshapes:
0.484
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs HotKnots
Matthews Correlation Coefficient
Carnac(20):
0.288
HotKnots:
0.179
Sensitivity
Carnac(20):
0.093
HotKnots:
0.051
Positive Predictive Value
Carnac(20):
0.892
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs HotKnots
Matthews Correlation Coefficient
RNASLOpt:
0.281
HotKnots:
0.179
Sensitivity
RNASLOpt:
0.158
HotKnots:
0.051
Positive Predictive Value
RNASLOpt:
0.502
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs HotKnots
Matthews Correlation Coefficient
RSpredict(20):
0.270
HotKnots:
0.179
Sensitivity
RSpredict(20):
0.124
HotKnots:
0.051
Positive Predictive Value
RSpredict(20):
0.590
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs HotKnots
Matthews Correlation Coefficient
Multilign(20):
0.251
HotKnots:
0.179
Sensitivity
Multilign(20):
0.083
HotKnots:
0.051
Positive Predictive Value
Multilign(20):
0.762
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs HotKnots
Matthews Correlation Coefficient
RNAwolf:
0.251
HotKnots:
0.179
Sensitivity
RNAwolf:
0.191
HotKnots:
0.051
Positive Predictive Value
RNAwolf:
0.330
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs HotKnots
Matthews Correlation Coefficient
Murlet(20):
0.230
HotKnots:
0.179
Sensitivity
Murlet(20):
0.067
HotKnots:
0.051
Positive Predictive Value
Murlet(20):
0.787
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs HotKnots
Matthews Correlation Coefficient
Vsfold4:
0.217
HotKnots:
0.179
Sensitivity
Vsfold4:
0.125
HotKnots:
0.051
Positive Predictive Value
Vsfold4:
0.378
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs HotKnots
Matthews Correlation Coefficient
Vsfold5:
0.211
HotKnots:
0.179
Sensitivity
Vsfold5:
0.125
HotKnots:
0.051
Positive Predictive Value
Vsfold5:
0.359
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs HotKnots
Matthews Correlation Coefficient
RSpredict(seed):
0.212
HotKnots:
0.179
Sensitivity
RSpredict(seed):
0.075
HotKnots:
0.051
Positive Predictive Value
RSpredict(seed):
0.597
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs HotKnots
Matthews Correlation Coefficient
CMfinder(20):
0.210
HotKnots:
0.179
Sensitivity
CMfinder(20):
0.055
HotKnots:
0.051
Positive Predictive Value
CMfinder(20):
0.794
HotKnots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
HotKnots vs RNASampler(20)
Matthews Correlation Coefficient
HotKnots:
0.179
RNASampler(20):
0.176
Sensitivity
HotKnots:
0.051
RNASampler(20):
0.035
Positive Predictive Value
HotKnots:
0.624
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 5.7281986208e-07
|
+
HotKnots vs Cylofold
Matthews Correlation Coefficient
HotKnots:
0.179
Cylofold:
0.155
Sensitivity
HotKnots:
0.051
Cylofold:
0.040
Positive Predictive Value
HotKnots:
0.624
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Pknots
Matthews Correlation Coefficient
HotKnots:
0.179
Pknots:
0.143
Sensitivity
HotKnots:
0.051
Pknots:
0.033
Positive Predictive Value
HotKnots:
0.624
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Mastr(20)
Matthews Correlation Coefficient
HotKnots:
0.179
Mastr(20):
0.134
Sensitivity
HotKnots:
0.051
Mastr(20):
0.023
Positive Predictive Value
HotKnots:
0.624
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Alterna
Matthews Correlation Coefficient
HotKnots:
0.179
Alterna:
0.129
Sensitivity
HotKnots:
0.051
Alterna:
0.030
Positive Predictive Value
HotKnots:
0.624
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs MCFold
Matthews Correlation Coefficient
HotKnots:
0.179
MCFold:
0.125
Sensitivity
HotKnots:
0.051
MCFold:
0.036
Positive Predictive Value
HotKnots:
0.624
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs RDfolder
Matthews Correlation Coefficient
HotKnots:
0.179
RDfolder:
0.122
Sensitivity
HotKnots:
0.051
RDfolder:
0.025
Positive Predictive Value
HotKnots:
0.624
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Murlet(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Murlet(seed):
0.079
Sensitivity
HotKnots:
0.051
Murlet(seed):
0.008
Positive Predictive Value
HotKnots:
0.624
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs CMfinder(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
CMfinder(seed):
0.080
Sensitivity
HotKnots:
0.051
CMfinder(seed):
0.008
Positive Predictive Value
HotKnots:
0.624
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs TurboFold(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
TurboFold(seed):
0.076
Sensitivity
HotKnots:
0.051
TurboFold(seed):
0.007
Positive Predictive Value
HotKnots:
0.624
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs PPfold(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
PPfold(seed):
0.069
Sensitivity
HotKnots:
0.051
PPfold(seed):
0.006
Positive Predictive Value
HotKnots:
0.624
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Carnac(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Carnac(seed):
0.066
Sensitivity
HotKnots:
0.051
Carnac(seed):
0.005
Positive Predictive Value
HotKnots:
0.624
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs RNASampler(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
RNASampler(seed):
0.063
Sensitivity
HotKnots:
0.051
RNASampler(seed):
0.005
Positive Predictive Value
HotKnots:
0.624
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs NanoFolder
Matthews Correlation Coefficient
HotKnots:
0.179
NanoFolder:
0.052
Sensitivity
HotKnots:
0.051
NanoFolder:
0.016
Positive Predictive Value
HotKnots:
0.624
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Mastr(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Mastr(seed):
0.051
Sensitivity
HotKnots:
0.051
Mastr(seed):
0.003
Positive Predictive Value
HotKnots:
0.624
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Multilign(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Multilign(seed):
0.043
Sensitivity
HotKnots:
0.051
Multilign(seed):
0.003
Positive Predictive Value
HotKnots:
0.624
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNASampler(20) |
1987
ContextFold vs RNASampler(20)
Matthews Correlation Coefficient
ContextFold:
0.680
RNASampler(20):
0.176
Sensitivity
ContextFold:
0.644
RNASampler(20):
0.035
Positive Predictive Value
ContextFold:
0.718
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNASampler(20)
Matthews Correlation Coefficient
IPknot:
0.548
RNASampler(20):
0.176
Sensitivity
IPknot:
0.475
RNASampler(20):
0.035
Positive Predictive Value
IPknot:
0.632
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNASampler(20):
0.176
Sensitivity
PETfold_pre2.0(seed):
0.343
RNASampler(20):
0.035
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNASampler(20)
Matthews Correlation Coefficient
Contrafold:
0.508
RNASampler(20):
0.176
Sensitivity
Contrafold:
0.481
RNASampler(20):
0.035
Positive Predictive Value
Contrafold:
0.537
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNASampler(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNASampler(20):
0.176
Sensitivity
CentroidFold:
0.369
RNASampler(20):
0.035
Positive Predictive Value
CentroidFold:
0.600
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNASampler(20)
Matthews Correlation Coefficient
Sfold:
0.469
RNASampler(20):
0.176
Sensitivity
Sfold:
0.412
RNASampler(20):
0.035
Positive Predictive Value
Sfold:
0.535
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNASampler(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNASampler(20):
0.176
Sensitivity
MaxExpect:
0.443
RNASampler(20):
0.035
Positive Predictive Value
MaxExpect:
0.488
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNASampler(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNASampler(20):
0.176
Sensitivity
ProbKnot:
0.446
RNASampler(20):
0.035
Positive Predictive Value
ProbKnot:
0.475
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNASampler(20)
Matthews Correlation Coefficient
UNAFold:
0.445
RNASampler(20):
0.176
Sensitivity
UNAFold:
0.445
RNASampler(20):
0.035
Positive Predictive Value
UNAFold:
0.446
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNASampler(20):
0.176
Sensitivity
CentroidAlifold(seed):
0.213
RNASampler(20):
0.035
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNASampler(20)
Matthews Correlation Coefficient
Fold:
0.437
RNASampler(20):
0.176
Sensitivity
Fold:
0.432
RNASampler(20):
0.035
Positive Predictive Value
Fold:
0.444
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNASampler(20)
Matthews Correlation Coefficient
RNAfold:
0.429
RNASampler(20):
0.176
Sensitivity
RNAfold:
0.422
RNASampler(20):
0.035
Positive Predictive Value
RNAfold:
0.437
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNASampler(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNASampler(20):
0.176
Sensitivity
CentroidHomfold‑LAST:
0.216
RNASampler(20):
0.035
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNASampler(20):
0.176
Sensitivity
PETfold_pre2.0(20):
0.211
RNASampler(20):
0.035
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNASampler(20):
0.176
Sensitivity
CentroidAlifold(20):
0.190
RNASampler(20):
0.035
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNASampler(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNASampler(20):
0.176
Sensitivity
PknotsRG:
0.320
RNASampler(20):
0.035
Positive Predictive Value
PknotsRG:
0.465
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNASampler(20):
0.176
Sensitivity
MXScarna(seed):
0.196
RNASampler(20):
0.035
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNASampler(20):
0.176
Sensitivity
RNAalifold(20):
0.185
RNASampler(20):
0.035
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNASampler(20):
0.176
Sensitivity
RNAalifold(seed):
0.169
RNASampler(20):
0.035
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RNASampler(20)
Matthews Correlation Coefficient
Afold:
0.358
RNASampler(20):
0.176
Sensitivity
Afold:
0.295
RNASampler(20):
0.035
Positive Predictive Value
Afold:
0.434
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNASampler(20):
0.176
Sensitivity
MXScarna(20):
0.181
RNASampler(20):
0.035
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RNASampler(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNASampler(20):
0.176
Sensitivity
CRWrnafold:
0.275
RNASampler(20):
0.035
Positive Predictive Value
CRWrnafold:
0.435
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RNASampler(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNASampler(20):
0.176
Sensitivity
RNAsubopt:
0.217
RNASampler(20):
0.035
Positive Predictive Value
RNAsubopt:
0.521
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RNASampler(20)
Matthews Correlation Coefficient
McQFold:
0.314
RNASampler(20):
0.176
Sensitivity
McQFold:
0.205
RNASampler(20):
0.035
Positive Predictive Value
McQFold:
0.480
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RNASampler(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNASampler(20):
0.176
Sensitivity
TurboFold(20):
0.108
RNASampler(20):
0.035
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RNASampler(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
RNASampler(20):
0.176
Sensitivity
PPfold(20):
0.101
RNASampler(20):
0.035
Positive Predictive Value
PPfold(20):
0.879
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RNASampler(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
RNASampler(20):
0.176
Sensitivity
RNAshapes:
0.180
RNASampler(20):
0.035
Positive Predictive Value
RNAshapes:
0.484
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RNASampler(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
RNASampler(20):
0.176
Sensitivity
Carnac(20):
0.093
RNASampler(20):
0.035
Positive Predictive Value
Carnac(20):
0.892
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs RNASampler(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RNASampler(20):
0.176
Sensitivity
RNASLOpt:
0.158
RNASampler(20):
0.035
Positive Predictive Value
RNASLOpt:
0.502
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
RNASampler(20):
0.176
Sensitivity
RSpredict(20):
0.124
RNASampler(20):
0.035
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs RNASampler(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
RNASampler(20):
0.176
Sensitivity
Multilign(20):
0.083
RNASampler(20):
0.035
Positive Predictive Value
Multilign(20):
0.762
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs RNASampler(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
RNASampler(20):
0.176
Sensitivity
RNAwolf:
0.191
RNASampler(20):
0.035
Positive Predictive Value
RNAwolf:
0.330
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs RNASampler(20)
Matthews Correlation Coefficient
Murlet(20):
0.230
RNASampler(20):
0.176
Sensitivity
Murlet(20):
0.067
RNASampler(20):
0.035
Positive Predictive Value
Murlet(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold4:
0.217
RNASampler(20):
0.176
Sensitivity
Vsfold4:
0.125
RNASampler(20):
0.035
Positive Predictive Value
Vsfold4:
0.378
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold5:
0.211
RNASampler(20):
0.176
Sensitivity
Vsfold5:
0.125
RNASampler(20):
0.035
Positive Predictive Value
Vsfold5:
0.359
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
RNASampler(20):
0.176
Sensitivity
RSpredict(seed):
0.075
RNASampler(20):
0.035
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs RNASampler(20)
Matthews Correlation Coefficient
CMfinder(20):
0.210
RNASampler(20):
0.176
Sensitivity
CMfinder(20):
0.055
RNASampler(20):
0.035
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs RNASampler(20)
Matthews Correlation Coefficient
HotKnots:
0.179
RNASampler(20):
0.176
Sensitivity
HotKnots:
0.051
RNASampler(20):
0.035
Positive Predictive Value
HotKnots:
0.624
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 5.7281986208e-07
|
|
+
RNASampler(20) vs Cylofold
Matthews Correlation Coefficient
RNASampler(20):
0.176
Cylofold:
0.155
Sensitivity
RNASampler(20):
0.035
Cylofold:
0.040
Positive Predictive Value
RNASampler(20):
0.877
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Pknots
Matthews Correlation Coefficient
RNASampler(20):
0.176
Pknots:
0.143
Sensitivity
RNASampler(20):
0.035
Pknots:
0.033
Positive Predictive Value
RNASampler(20):
0.877
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Mastr(20)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Mastr(20):
0.134
Sensitivity
RNASampler(20):
0.035
Mastr(20):
0.023
Positive Predictive Value
RNASampler(20):
0.877
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Alterna
Matthews Correlation Coefficient
RNASampler(20):
0.176
Alterna:
0.129
Sensitivity
RNASampler(20):
0.035
Alterna:
0.030
Positive Predictive Value
RNASampler(20):
0.877
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs MCFold
Matthews Correlation Coefficient
RNASampler(20):
0.176
MCFold:
0.125
Sensitivity
RNASampler(20):
0.035
MCFold:
0.036
Positive Predictive Value
RNASampler(20):
0.877
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs RDfolder
Matthews Correlation Coefficient
RNASampler(20):
0.176
RDfolder:
0.122
Sensitivity
RNASampler(20):
0.035
RDfolder:
0.025
Positive Predictive Value
RNASampler(20):
0.877
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Murlet(seed):
0.079
Sensitivity
RNASampler(20):
0.035
Murlet(seed):
0.008
Positive Predictive Value
RNASampler(20):
0.877
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
CMfinder(seed):
0.080
Sensitivity
RNASampler(20):
0.035
CMfinder(seed):
0.008
Positive Predictive Value
RNASampler(20):
0.877
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
TurboFold(seed):
0.076
Sensitivity
RNASampler(20):
0.035
TurboFold(seed):
0.007
Positive Predictive Value
RNASampler(20):
0.877
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
PPfold(seed):
0.069
Sensitivity
RNASampler(20):
0.035
PPfold(seed):
0.006
Positive Predictive Value
RNASampler(20):
0.877
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Carnac(seed):
0.066
Sensitivity
RNASampler(20):
0.035
Carnac(seed):
0.005
Positive Predictive Value
RNASampler(20):
0.877
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
RNASampler(seed):
0.063
Sensitivity
RNASampler(20):
0.035
RNASampler(seed):
0.005
Positive Predictive Value
RNASampler(20):
0.877
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(20):
0.176
NanoFolder:
0.052
Sensitivity
RNASampler(20):
0.035
NanoFolder:
0.016
Positive Predictive Value
RNASampler(20):
0.877
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Mastr(seed):
0.051
Sensitivity
RNASampler(20):
0.035
Mastr(seed):
0.003
Positive Predictive Value
RNASampler(20):
0.877
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Multilign(seed):
0.043
Sensitivity
RNASampler(20):
0.035
Multilign(seed):
0.003
Positive Predictive Value
RNASampler(20):
0.877
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Cylofold |
1987
ContextFold vs Cylofold
Matthews Correlation Coefficient
ContextFold:
0.680
Cylofold:
0.155
Sensitivity
ContextFold:
0.644
Cylofold:
0.040
Positive Predictive Value
ContextFold:
0.718
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Cylofold
Matthews Correlation Coefficient
IPknot:
0.548
Cylofold:
0.155
Sensitivity
IPknot:
0.475
Cylofold:
0.040
Positive Predictive Value
IPknot:
0.632
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Cylofold:
0.155
Sensitivity
PETfold_pre2.0(seed):
0.343
Cylofold:
0.040
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Cylofold
Matthews Correlation Coefficient
Contrafold:
0.508
Cylofold:
0.155
Sensitivity
Contrafold:
0.481
Cylofold:
0.040
Positive Predictive Value
Contrafold:
0.537
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Cylofold
Matthews Correlation Coefficient
CentroidFold:
0.470
Cylofold:
0.155
Sensitivity
CentroidFold:
0.369
Cylofold:
0.040
Positive Predictive Value
CentroidFold:
0.600
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Cylofold
Matthews Correlation Coefficient
Sfold:
0.469
Cylofold:
0.155
Sensitivity
Sfold:
0.412
Cylofold:
0.040
Positive Predictive Value
Sfold:
0.535
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Cylofold
Matthews Correlation Coefficient
MaxExpect:
0.465
Cylofold:
0.155
Sensitivity
MaxExpect:
0.443
Cylofold:
0.040
Positive Predictive Value
MaxExpect:
0.488
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Cylofold
Matthews Correlation Coefficient
ProbKnot:
0.460
Cylofold:
0.155
Sensitivity
ProbKnot:
0.446
Cylofold:
0.040
Positive Predictive Value
ProbKnot:
0.475
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Cylofold
Matthews Correlation Coefficient
UNAFold:
0.445
Cylofold:
0.155
Sensitivity
UNAFold:
0.445
Cylofold:
0.040
Positive Predictive Value
UNAFold:
0.446
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Cylofold:
0.155
Sensitivity
CentroidAlifold(seed):
0.213
Cylofold:
0.040
Positive Predictive Value
CentroidAlifold(seed):
0.901
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Cylofold
Matthews Correlation Coefficient
Fold:
0.437
Cylofold:
0.155
Sensitivity
Fold:
0.432
Cylofold:
0.040
Positive Predictive Value
Fold:
0.444
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Cylofold
Matthews Correlation Coefficient
RNAfold:
0.429
Cylofold:
0.155
Sensitivity
RNAfold:
0.422
Cylofold:
0.040
Positive Predictive Value
RNAfold:
0.437
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Cylofold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Cylofold:
0.155
Sensitivity
CentroidHomfold‑LAST:
0.216
Cylofold:
0.040
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Cylofold:
0.155
Sensitivity
PETfold_pre2.0(20):
0.211
Cylofold:
0.040
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Cylofold:
0.155
Sensitivity
CentroidAlifold(20):
0.190
Cylofold:
0.040
Positive Predictive Value
CentroidAlifold(20):
0.891
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Cylofold
Matthews Correlation Coefficient
PknotsRG:
0.386
Cylofold:
0.155
Sensitivity
PknotsRG:
0.320
Cylofold:
0.040
Positive Predictive Value
PknotsRG:
0.465
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Cylofold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Cylofold:
0.155
Sensitivity
MXScarna(seed):
0.196
Cylofold:
0.040
Positive Predictive Value
MXScarna(seed):
0.753
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Cylofold:
0.155
Sensitivity
RNAalifold(20):
0.185
Cylofold:
0.040
Positive Predictive Value
RNAalifold(20):
0.787
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Cylofold:
0.155
Sensitivity
RNAalifold(seed):
0.169
Cylofold:
0.040
Positive Predictive Value
RNAalifold(seed):
0.835
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Cylofold
Matthews Correlation Coefficient
Afold:
0.358
Cylofold:
0.155
Sensitivity
Afold:
0.295
Cylofold:
0.040
Positive Predictive Value
Afold:
0.434
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Cylofold
Matthews Correlation Coefficient
MXScarna(20):
0.359
Cylofold:
0.155
Sensitivity
MXScarna(20):
0.181
Cylofold:
0.040
Positive Predictive Value
MXScarna(20):
0.710
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Cylofold
Matthews Correlation Coefficient
CRWrnafold:
0.346
Cylofold:
0.155
Sensitivity
CRWrnafold:
0.275
Cylofold:
0.040
Positive Predictive Value
CRWrnafold:
0.435
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Cylofold
Matthews Correlation Coefficient
RNAsubopt:
0.336
Cylofold:
0.155
Sensitivity
RNAsubopt:
0.217
Cylofold:
0.040
Positive Predictive Value
RNAsubopt:
0.521
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Cylofold
Matthews Correlation Coefficient
McQFold:
0.314
Cylofold:
0.155
Sensitivity
McQFold:
0.205
Cylofold:
0.040
Positive Predictive Value
McQFold:
0.480
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Cylofold
Matthews Correlation Coefficient
TurboFold(20):
0.301
Cylofold:
0.155
Sensitivity
TurboFold(20):
0.108
Cylofold:
0.040
Positive Predictive Value
TurboFold(20):
0.837
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Cylofold
Matthews Correlation Coefficient
PPfold(20):
0.298
Cylofold:
0.155
Sensitivity
PPfold(20):
0.101
Cylofold:
0.040
Positive Predictive Value
PPfold(20):
0.879
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Cylofold
Matthews Correlation Coefficient
RNAshapes:
0.295
Cylofold:
0.155
Sensitivity
RNAshapes:
0.180
Cylofold:
0.040
Positive Predictive Value
RNAshapes:
0.484
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Cylofold
Matthews Correlation Coefficient
Carnac(20):
0.288
Cylofold:
0.155
Sensitivity
Carnac(20):
0.093
Cylofold:
0.040
Positive Predictive Value
Carnac(20):
0.892
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Cylofold
Matthews Correlation Coefficient
RNASLOpt:
0.281
Cylofold:
0.155
Sensitivity
RNASLOpt:
0.158
Cylofold:
0.040
Positive Predictive Value
RNASLOpt:
0.502
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Cylofold
Matthews Correlation Coefficient
RSpredict(20):
0.270
Cylofold:
0.155
Sensitivity
RSpredict(20):
0.124
Cylofold:
0.040
Positive Predictive Value
RSpredict(20):
0.590
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Cylofold
Matthews Correlation Coefficient
Multilign(20):
0.251
Cylofold:
0.155
Sensitivity
Multilign(20):
0.083
Cylofold:
0.040
Positive Predictive Value
Multilign(20):
0.762
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Cylofold
Matthews Correlation Coefficient
RNAwolf:
0.251
Cylofold:
0.155
Sensitivity
RNAwolf:
0.191
Cylofold:
0.040
Positive Predictive Value
RNAwolf:
0.330
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Cylofold
Matthews Correlation Coefficient
Murlet(20):
0.230
Cylofold:
0.155
Sensitivity
Murlet(20):
0.067
Cylofold:
0.040
Positive Predictive Value
Murlet(20):
0.787
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Cylofold
Matthews Correlation Coefficient
Vsfold4:
0.217
Cylofold:
0.155
Sensitivity
Vsfold4:
0.125
Cylofold:
0.040
Positive Predictive Value
Vsfold4:
0.378
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Cylofold
Matthews Correlation Coefficient
Vsfold5:
0.211
Cylofold:
0.155
Sensitivity
Vsfold5:
0.125
Cylofold:
0.040
Positive Predictive Value
Vsfold5:
0.359
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Cylofold
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Cylofold:
0.155
Sensitivity
RSpredict(seed):
0.075
Cylofold:
0.040
Positive Predictive Value
RSpredict(seed):
0.597
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Cylofold
Matthews Correlation Coefficient
CMfinder(20):
0.210
Cylofold:
0.155
Sensitivity
CMfinder(20):
0.055
Cylofold:
0.040
Positive Predictive Value
CMfinder(20):
0.794
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Cylofold
Matthews Correlation Coefficient
HotKnots:
0.179
Cylofold:
0.155
Sensitivity
HotKnots:
0.051
Cylofold:
0.040
Positive Predictive Value
HotKnots:
0.624
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Cylofold
Matthews Correlation Coefficient
RNASampler(20):
0.176
Cylofold:
0.155
Sensitivity
RNASampler(20):
0.035
Cylofold:
0.040
Positive Predictive Value
RNASampler(20):
0.877
Cylofold:
0.606
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Cylofold vs Pknots
Matthews Correlation Coefficient
Cylofold:
0.155
Pknots:
0.143
Sensitivity
Cylofold:
0.040
Pknots:
0.033
Positive Predictive Value
Cylofold:
0.606
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Mastr(20)
Matthews Correlation Coefficient
Cylofold:
0.155
Mastr(20):
0.134
Sensitivity
Cylofold:
0.040
Mastr(20):
0.023
Positive Predictive Value
Cylofold:
0.606
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Alterna
Matthews Correlation Coefficient
Cylofold:
0.155
Alterna:
0.129
Sensitivity
Cylofold:
0.040
Alterna:
0.030
Positive Predictive Value
Cylofold:
0.606
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs MCFold
Matthews Correlation Coefficient
Cylofold:
0.155
MCFold:
0.125
Sensitivity
Cylofold:
0.040
MCFold:
0.036
Positive Predictive Value
Cylofold:
0.606
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs RDfolder
Matthews Correlation Coefficient
Cylofold:
0.155
RDfolder:
0.122
Sensitivity
Cylofold:
0.040
RDfolder:
0.025
Positive Predictive Value
Cylofold:
0.606
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Murlet(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Murlet(seed):
0.079
Sensitivity
Cylofold:
0.040
Murlet(seed):
0.008
Positive Predictive Value
Cylofold:
0.606
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs CMfinder(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
CMfinder(seed):
0.080
Sensitivity
Cylofold:
0.040
CMfinder(seed):
0.008
Positive Predictive Value
Cylofold:
0.606
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs TurboFold(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
TurboFold(seed):
0.076
Sensitivity
Cylofold:
0.040
TurboFold(seed):
0.007
Positive Predictive Value
Cylofold:
0.606
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs PPfold(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
PPfold(seed):
0.069
Sensitivity
Cylofold:
0.040
PPfold(seed):
0.006
Positive Predictive Value
Cylofold:
0.606
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Carnac(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Carnac(seed):
0.066
Sensitivity
Cylofold:
0.040
Carnac(seed):
0.005
Positive Predictive Value
Cylofold:
0.606
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs RNASampler(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
RNASampler(seed):
0.063
Sensitivity
Cylofold:
0.040
RNASampler(seed):
0.005
Positive Predictive Value
Cylofold:
0.606
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs NanoFolder
Matthews Correlation Coefficient
Cylofold:
0.155
NanoFolder:
0.052
Sensitivity
Cylofold:
0.040
NanoFolder:
0.016
Positive Predictive Value
Cylofold:
0.606
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Mastr(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Mastr(seed):
0.051
Sensitivity
Cylofold:
0.040
Mastr(seed):
0.003
Positive Predictive Value
Cylofold:
0.606
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Multilign(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Multilign(seed):
0.043
Sensitivity
Cylofold:
0.040
Multilign(seed):
0.003
Positive Predictive Value
Cylofold:
0.606
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Pknots |
1987
ContextFold vs Pknots
Matthews Correlation Coefficient
ContextFold:
0.680
Pknots:
0.143
Sensitivity
ContextFold:
0.644
Pknots:
0.033
Positive Predictive Value
ContextFold:
0.718
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Pknots
Matthews Correlation Coefficient
IPknot:
0.548
Pknots:
0.143
Sensitivity
IPknot:
0.475
Pknots:
0.033
Positive Predictive Value
IPknot:
0.632
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Pknots:
0.143
Sensitivity
PETfold_pre2.0(seed):
0.343
Pknots:
0.033
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Pknots
Matthews Correlation Coefficient
Contrafold:
0.508
Pknots:
0.143
Sensitivity
Contrafold:
0.481
Pknots:
0.033
Positive Predictive Value
Contrafold:
0.537
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Pknots
Matthews Correlation Coefficient
CentroidFold:
0.470
Pknots:
0.143
Sensitivity
CentroidFold:
0.369
Pknots:
0.033
Positive Predictive Value
CentroidFold:
0.600
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Pknots
Matthews Correlation Coefficient
Sfold:
0.469
Pknots:
0.143
Sensitivity
Sfold:
0.412
Pknots:
0.033
Positive Predictive Value
Sfold:
0.535
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Pknots
Matthews Correlation Coefficient
MaxExpect:
0.465
Pknots:
0.143
Sensitivity
MaxExpect:
0.443
Pknots:
0.033
Positive Predictive Value
MaxExpect:
0.488
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Pknots
Matthews Correlation Coefficient
ProbKnot:
0.460
Pknots:
0.143
Sensitivity
ProbKnot:
0.446
Pknots:
0.033
Positive Predictive Value
ProbKnot:
0.475
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Pknots
Matthews Correlation Coefficient
UNAFold:
0.445
Pknots:
0.143
Sensitivity
UNAFold:
0.445
Pknots:
0.033
Positive Predictive Value
UNAFold:
0.446
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Pknots:
0.143
Sensitivity
CentroidAlifold(seed):
0.213
Pknots:
0.033
Positive Predictive Value
CentroidAlifold(seed):
0.901
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Pknots
Matthews Correlation Coefficient
Fold:
0.437
Pknots:
0.143
Sensitivity
Fold:
0.432
Pknots:
0.033
Positive Predictive Value
Fold:
0.444
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Pknots
Matthews Correlation Coefficient
RNAfold:
0.429
Pknots:
0.143
Sensitivity
RNAfold:
0.422
Pknots:
0.033
Positive Predictive Value
RNAfold:
0.437
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Pknots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Pknots:
0.143
Sensitivity
CentroidHomfold‑LAST:
0.216
Pknots:
0.033
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Pknots:
0.143
Sensitivity
PETfold_pre2.0(20):
0.211
Pknots:
0.033
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Pknots:
0.143
Sensitivity
CentroidAlifold(20):
0.190
Pknots:
0.033
Positive Predictive Value
CentroidAlifold(20):
0.891
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Pknots
Matthews Correlation Coefficient
PknotsRG:
0.386
Pknots:
0.143
Sensitivity
PknotsRG:
0.320
Pknots:
0.033
Positive Predictive Value
PknotsRG:
0.465
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Pknots
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Pknots:
0.143
Sensitivity
MXScarna(seed):
0.196
Pknots:
0.033
Positive Predictive Value
MXScarna(seed):
0.753
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Pknots
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Pknots:
0.143
Sensitivity
RNAalifold(20):
0.185
Pknots:
0.033
Positive Predictive Value
RNAalifold(20):
0.787
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Pknots
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Pknots:
0.143
Sensitivity
RNAalifold(seed):
0.169
Pknots:
0.033
Positive Predictive Value
RNAalifold(seed):
0.835
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Pknots
Matthews Correlation Coefficient
Afold:
0.358
Pknots:
0.143
Sensitivity
Afold:
0.295
Pknots:
0.033
Positive Predictive Value
Afold:
0.434
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Pknots
Matthews Correlation Coefficient
MXScarna(20):
0.359
Pknots:
0.143
Sensitivity
MXScarna(20):
0.181
Pknots:
0.033
Positive Predictive Value
MXScarna(20):
0.710
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Pknots
Matthews Correlation Coefficient
CRWrnafold:
0.346
Pknots:
0.143
Sensitivity
CRWrnafold:
0.275
Pknots:
0.033
Positive Predictive Value
CRWrnafold:
0.435
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Pknots
Matthews Correlation Coefficient
RNAsubopt:
0.336
Pknots:
0.143
Sensitivity
RNAsubopt:
0.217
Pknots:
0.033
Positive Predictive Value
RNAsubopt:
0.521
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Pknots
Matthews Correlation Coefficient
McQFold:
0.314
Pknots:
0.143
Sensitivity
McQFold:
0.205
Pknots:
0.033
Positive Predictive Value
McQFold:
0.480
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Pknots
Matthews Correlation Coefficient
TurboFold(20):
0.301
Pknots:
0.143
Sensitivity
TurboFold(20):
0.108
Pknots:
0.033
Positive Predictive Value
TurboFold(20):
0.837
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Pknots
Matthews Correlation Coefficient
PPfold(20):
0.298
Pknots:
0.143
Sensitivity
PPfold(20):
0.101
Pknots:
0.033
Positive Predictive Value
PPfold(20):
0.879
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Pknots
Matthews Correlation Coefficient
RNAshapes:
0.295
Pknots:
0.143
Sensitivity
RNAshapes:
0.180
Pknots:
0.033
Positive Predictive Value
RNAshapes:
0.484
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Pknots
Matthews Correlation Coefficient
Carnac(20):
0.288
Pknots:
0.143
Sensitivity
Carnac(20):
0.093
Pknots:
0.033
Positive Predictive Value
Carnac(20):
0.892
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Pknots
Matthews Correlation Coefficient
RNASLOpt:
0.281
Pknots:
0.143
Sensitivity
RNASLOpt:
0.158
Pknots:
0.033
Positive Predictive Value
RNASLOpt:
0.502
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Pknots
Matthews Correlation Coefficient
RSpredict(20):
0.270
Pknots:
0.143
Sensitivity
RSpredict(20):
0.124
Pknots:
0.033
Positive Predictive Value
RSpredict(20):
0.590
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Pknots
Matthews Correlation Coefficient
Multilign(20):
0.251
Pknots:
0.143
Sensitivity
Multilign(20):
0.083
Pknots:
0.033
Positive Predictive Value
Multilign(20):
0.762
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Pknots
Matthews Correlation Coefficient
RNAwolf:
0.251
Pknots:
0.143
Sensitivity
RNAwolf:
0.191
Pknots:
0.033
Positive Predictive Value
RNAwolf:
0.330
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Pknots
Matthews Correlation Coefficient
Murlet(20):
0.230
Pknots:
0.143
Sensitivity
Murlet(20):
0.067
Pknots:
0.033
Positive Predictive Value
Murlet(20):
0.787
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Pknots
Matthews Correlation Coefficient
Vsfold4:
0.217
Pknots:
0.143
Sensitivity
Vsfold4:
0.125
Pknots:
0.033
Positive Predictive Value
Vsfold4:
0.378
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Pknots
Matthews Correlation Coefficient
Vsfold5:
0.211
Pknots:
0.143
Sensitivity
Vsfold5:
0.125
Pknots:
0.033
Positive Predictive Value
Vsfold5:
0.359
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Pknots
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Pknots:
0.143
Sensitivity
RSpredict(seed):
0.075
Pknots:
0.033
Positive Predictive Value
RSpredict(seed):
0.597
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Pknots
Matthews Correlation Coefficient
CMfinder(20):
0.210
Pknots:
0.143
Sensitivity
CMfinder(20):
0.055
Pknots:
0.033
Positive Predictive Value
CMfinder(20):
0.794
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Pknots
Matthews Correlation Coefficient
HotKnots:
0.179
Pknots:
0.143
Sensitivity
HotKnots:
0.051
Pknots:
0.033
Positive Predictive Value
HotKnots:
0.624
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Pknots
Matthews Correlation Coefficient
RNASampler(20):
0.176
Pknots:
0.143
Sensitivity
RNASampler(20):
0.035
Pknots:
0.033
Positive Predictive Value
RNASampler(20):
0.877
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Pknots
Matthews Correlation Coefficient
Cylofold:
0.155
Pknots:
0.143
Sensitivity
Cylofold:
0.040
Pknots:
0.033
Positive Predictive Value
Cylofold:
0.606
Pknots:
0.624
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Pknots vs Mastr(20)
Matthews Correlation Coefficient
Pknots:
0.143
Mastr(20):
0.134
Sensitivity
Pknots:
0.033
Mastr(20):
0.023
Positive Predictive Value
Pknots:
0.624
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Alterna
Matthews Correlation Coefficient
Pknots:
0.143
Alterna:
0.129
Sensitivity
Pknots:
0.033
Alterna:
0.030
Positive Predictive Value
Pknots:
0.624
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs MCFold
Matthews Correlation Coefficient
Pknots:
0.143
MCFold:
0.125
Sensitivity
Pknots:
0.033
MCFold:
0.036
Positive Predictive Value
Pknots:
0.624
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs RDfolder
Matthews Correlation Coefficient
Pknots:
0.143
RDfolder:
0.122
Sensitivity
Pknots:
0.033
RDfolder:
0.025
Positive Predictive Value
Pknots:
0.624
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Murlet(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Murlet(seed):
0.079
Sensitivity
Pknots:
0.033
Murlet(seed):
0.008
Positive Predictive Value
Pknots:
0.624
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs CMfinder(seed)
Matthews Correlation Coefficient
Pknots:
0.143
CMfinder(seed):
0.080
Sensitivity
Pknots:
0.033
CMfinder(seed):
0.008
Positive Predictive Value
Pknots:
0.624
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs TurboFold(seed)
Matthews Correlation Coefficient
Pknots:
0.143
TurboFold(seed):
0.076
Sensitivity
Pknots:
0.033
TurboFold(seed):
0.007
Positive Predictive Value
Pknots:
0.624
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs PPfold(seed)
Matthews Correlation Coefficient
Pknots:
0.143
PPfold(seed):
0.069
Sensitivity
Pknots:
0.033
PPfold(seed):
0.006
Positive Predictive Value
Pknots:
0.624
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Carnac(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Carnac(seed):
0.066
Sensitivity
Pknots:
0.033
Carnac(seed):
0.005
Positive Predictive Value
Pknots:
0.624
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs RNASampler(seed)
Matthews Correlation Coefficient
Pknots:
0.143
RNASampler(seed):
0.063
Sensitivity
Pknots:
0.033
RNASampler(seed):
0.005
Positive Predictive Value
Pknots:
0.624
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs NanoFolder
Matthews Correlation Coefficient
Pknots:
0.143
NanoFolder:
0.052
Sensitivity
Pknots:
0.033
NanoFolder:
0.016
Positive Predictive Value
Pknots:
0.624
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Mastr(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Mastr(seed):
0.051
Sensitivity
Pknots:
0.033
Mastr(seed):
0.003
Positive Predictive Value
Pknots:
0.624
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Multilign(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Multilign(seed):
0.043
Sensitivity
Pknots:
0.033
Multilign(seed):
0.003
Positive Predictive Value
Pknots:
0.624
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Mastr(20) |
1987
ContextFold vs Mastr(20)
Matthews Correlation Coefficient
ContextFold:
0.680
Mastr(20):
0.134
Sensitivity
ContextFold:
0.644
Mastr(20):
0.023
Positive Predictive Value
ContextFold:
0.718
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Mastr(20)
Matthews Correlation Coefficient
IPknot:
0.548
Mastr(20):
0.134
Sensitivity
IPknot:
0.475
Mastr(20):
0.023
Positive Predictive Value
IPknot:
0.632
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Mastr(20):
0.134
Sensitivity
PETfold_pre2.0(seed):
0.343
Mastr(20):
0.023
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Mastr(20)
Matthews Correlation Coefficient
Contrafold:
0.508
Mastr(20):
0.134
Sensitivity
Contrafold:
0.481
Mastr(20):
0.023
Positive Predictive Value
Contrafold:
0.537
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Mastr(20)
Matthews Correlation Coefficient
CentroidFold:
0.470
Mastr(20):
0.134
Sensitivity
CentroidFold:
0.369
Mastr(20):
0.023
Positive Predictive Value
CentroidFold:
0.600
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Mastr(20)
Matthews Correlation Coefficient
Sfold:
0.469
Mastr(20):
0.134
Sensitivity
Sfold:
0.412
Mastr(20):
0.023
Positive Predictive Value
Sfold:
0.535
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Mastr(20)
Matthews Correlation Coefficient
MaxExpect:
0.465
Mastr(20):
0.134
Sensitivity
MaxExpect:
0.443
Mastr(20):
0.023
Positive Predictive Value
MaxExpect:
0.488
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Mastr(20)
Matthews Correlation Coefficient
ProbKnot:
0.460
Mastr(20):
0.134
Sensitivity
ProbKnot:
0.446
Mastr(20):
0.023
Positive Predictive Value
ProbKnot:
0.475
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Mastr(20)
Matthews Correlation Coefficient
UNAFold:
0.445
Mastr(20):
0.134
Sensitivity
UNAFold:
0.445
Mastr(20):
0.023
Positive Predictive Value
UNAFold:
0.446
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Mastr(20):
0.134
Sensitivity
CentroidAlifold(seed):
0.213
Mastr(20):
0.023
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Mastr(20)
Matthews Correlation Coefficient
Fold:
0.437
Mastr(20):
0.134
Sensitivity
Fold:
0.432
Mastr(20):
0.023
Positive Predictive Value
Fold:
0.444
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Mastr(20)
Matthews Correlation Coefficient
RNAfold:
0.429
Mastr(20):
0.134
Sensitivity
RNAfold:
0.422
Mastr(20):
0.023
Positive Predictive Value
RNAfold:
0.437
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Mastr(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Mastr(20):
0.134
Sensitivity
CentroidHomfold‑LAST:
0.216
Mastr(20):
0.023
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Mastr(20):
0.134
Sensitivity
PETfold_pre2.0(20):
0.211
Mastr(20):
0.023
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Mastr(20):
0.134
Sensitivity
CentroidAlifold(20):
0.190
Mastr(20):
0.023
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Mastr(20)
Matthews Correlation Coefficient
PknotsRG:
0.386
Mastr(20):
0.134
Sensitivity
PknotsRG:
0.320
Mastr(20):
0.023
Positive Predictive Value
PknotsRG:
0.465
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Mastr(20):
0.134
Sensitivity
MXScarna(seed):
0.196
Mastr(20):
0.023
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Mastr(20):
0.134
Sensitivity
RNAalifold(20):
0.185
Mastr(20):
0.023
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Mastr(20):
0.134
Sensitivity
RNAalifold(seed):
0.169
Mastr(20):
0.023
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Mastr(20)
Matthews Correlation Coefficient
Afold:
0.358
Mastr(20):
0.134
Sensitivity
Afold:
0.295
Mastr(20):
0.023
Positive Predictive Value
Afold:
0.434
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Mastr(20):
0.134
Sensitivity
MXScarna(20):
0.181
Mastr(20):
0.023
Positive Predictive Value
MXScarna(20):
0.710
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Mastr(20)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Mastr(20):
0.134
Sensitivity
CRWrnafold:
0.275
Mastr(20):
0.023
Positive Predictive Value
CRWrnafold:
0.435
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Mastr(20)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Mastr(20):
0.134
Sensitivity
RNAsubopt:
0.217
Mastr(20):
0.023
Positive Predictive Value
RNAsubopt:
0.521
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Mastr(20)
Matthews Correlation Coefficient
McQFold:
0.314
Mastr(20):
0.134
Sensitivity
McQFold:
0.205
Mastr(20):
0.023
Positive Predictive Value
McQFold:
0.480
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Mastr(20)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Mastr(20):
0.134
Sensitivity
TurboFold(20):
0.108
Mastr(20):
0.023
Positive Predictive Value
TurboFold(20):
0.837
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Mastr(20)
Matthews Correlation Coefficient
PPfold(20):
0.298
Mastr(20):
0.134
Sensitivity
PPfold(20):
0.101
Mastr(20):
0.023
Positive Predictive Value
PPfold(20):
0.879
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Mastr(20)
Matthews Correlation Coefficient
RNAshapes:
0.295
Mastr(20):
0.134
Sensitivity
RNAshapes:
0.180
Mastr(20):
0.023
Positive Predictive Value
RNAshapes:
0.484
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Mastr(20)
Matthews Correlation Coefficient
Carnac(20):
0.288
Mastr(20):
0.134
Sensitivity
Carnac(20):
0.093
Mastr(20):
0.023
Positive Predictive Value
Carnac(20):
0.892
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Mastr(20)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Mastr(20):
0.134
Sensitivity
RNASLOpt:
0.158
Mastr(20):
0.023
Positive Predictive Value
RNASLOpt:
0.502
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Mastr(20):
0.134
Sensitivity
RSpredict(20):
0.124
Mastr(20):
0.023
Positive Predictive Value
RSpredict(20):
0.590
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Mastr(20)
Matthews Correlation Coefficient
Multilign(20):
0.251
Mastr(20):
0.134
Sensitivity
Multilign(20):
0.083
Mastr(20):
0.023
Positive Predictive Value
Multilign(20):
0.762
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Mastr(20)
Matthews Correlation Coefficient
RNAwolf:
0.251
Mastr(20):
0.134
Sensitivity
RNAwolf:
0.191
Mastr(20):
0.023
Positive Predictive Value
RNAwolf:
0.330
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Mastr(20)
Matthews Correlation Coefficient
Murlet(20):
0.230
Mastr(20):
0.134
Sensitivity
Murlet(20):
0.067
Mastr(20):
0.023
Positive Predictive Value
Murlet(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold4:
0.217
Mastr(20):
0.134
Sensitivity
Vsfold4:
0.125
Mastr(20):
0.023
Positive Predictive Value
Vsfold4:
0.378
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold5:
0.211
Mastr(20):
0.134
Sensitivity
Vsfold5:
0.125
Mastr(20):
0.023
Positive Predictive Value
Vsfold5:
0.359
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Mastr(20):
0.134
Sensitivity
RSpredict(seed):
0.075
Mastr(20):
0.023
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Mastr(20)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Mastr(20):
0.134
Sensitivity
CMfinder(20):
0.055
Mastr(20):
0.023
Positive Predictive Value
CMfinder(20):
0.794
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Mastr(20)
Matthews Correlation Coefficient
HotKnots:
0.179
Mastr(20):
0.134
Sensitivity
HotKnots:
0.051
Mastr(20):
0.023
Positive Predictive Value
HotKnots:
0.624
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Mastr(20)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Mastr(20):
0.134
Sensitivity
RNASampler(20):
0.035
Mastr(20):
0.023
Positive Predictive Value
RNASampler(20):
0.877
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Mastr(20)
Matthews Correlation Coefficient
Cylofold:
0.155
Mastr(20):
0.134
Sensitivity
Cylofold:
0.040
Mastr(20):
0.023
Positive Predictive Value
Cylofold:
0.606
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs Mastr(20)
Matthews Correlation Coefficient
Pknots:
0.143
Mastr(20):
0.134
Sensitivity
Pknots:
0.033
Mastr(20):
0.023
Positive Predictive Value
Pknots:
0.624
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Mastr(20) vs Alterna
Matthews Correlation Coefficient
Mastr(20):
0.134
Alterna:
0.129
Sensitivity
Mastr(20):
0.023
Alterna:
0.030
Positive Predictive Value
Mastr(20):
0.790
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs MCFold
Matthews Correlation Coefficient
Mastr(20):
0.134
MCFold:
0.125
Sensitivity
Mastr(20):
0.023
MCFold:
0.036
Positive Predictive Value
Mastr(20):
0.790
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs RDfolder
Matthews Correlation Coefficient
Mastr(20):
0.134
RDfolder:
0.122
Sensitivity
Mastr(20):
0.023
RDfolder:
0.025
Positive Predictive Value
Mastr(20):
0.790
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Murlet(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Murlet(seed):
0.079
Sensitivity
Mastr(20):
0.023
Murlet(seed):
0.008
Positive Predictive Value
Mastr(20):
0.790
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
CMfinder(seed):
0.080
Sensitivity
Mastr(20):
0.023
CMfinder(seed):
0.008
Positive Predictive Value
Mastr(20):
0.790
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
TurboFold(seed):
0.076
Sensitivity
Mastr(20):
0.023
TurboFold(seed):
0.007
Positive Predictive Value
Mastr(20):
0.790
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs PPfold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
PPfold(seed):
0.069
Sensitivity
Mastr(20):
0.023
PPfold(seed):
0.006
Positive Predictive Value
Mastr(20):
0.790
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Carnac(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Carnac(seed):
0.066
Sensitivity
Mastr(20):
0.023
Carnac(seed):
0.005
Positive Predictive Value
Mastr(20):
0.790
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
RNASampler(seed):
0.063
Sensitivity
Mastr(20):
0.023
RNASampler(seed):
0.005
Positive Predictive Value
Mastr(20):
0.790
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs NanoFolder
Matthews Correlation Coefficient
Mastr(20):
0.134
NanoFolder:
0.052
Sensitivity
Mastr(20):
0.023
NanoFolder:
0.016
Positive Predictive Value
Mastr(20):
0.790
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Mastr(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Mastr(seed):
0.051
Sensitivity
Mastr(20):
0.023
Mastr(seed):
0.003
Positive Predictive Value
Mastr(20):
0.790
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Multilign(seed):
0.043
Sensitivity
Mastr(20):
0.023
Multilign(seed):
0.003
Positive Predictive Value
Mastr(20):
0.790
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Alterna |
1987
ContextFold vs Alterna
Matthews Correlation Coefficient
ContextFold:
0.680
Alterna:
0.129
Sensitivity
ContextFold:
0.644
Alterna:
0.030
Positive Predictive Value
ContextFold:
0.718
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Alterna
Matthews Correlation Coefficient
IPknot:
0.548
Alterna:
0.129
Sensitivity
IPknot:
0.475
Alterna:
0.030
Positive Predictive Value
IPknot:
0.632
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Alterna:
0.129
Sensitivity
PETfold_pre2.0(seed):
0.343
Alterna:
0.030
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Alterna
Matthews Correlation Coefficient
Contrafold:
0.508
Alterna:
0.129
Sensitivity
Contrafold:
0.481
Alterna:
0.030
Positive Predictive Value
Contrafold:
0.537
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Alterna
Matthews Correlation Coefficient
CentroidFold:
0.470
Alterna:
0.129
Sensitivity
CentroidFold:
0.369
Alterna:
0.030
Positive Predictive Value
CentroidFold:
0.600
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Alterna
Matthews Correlation Coefficient
Sfold:
0.469
Alterna:
0.129
Sensitivity
Sfold:
0.412
Alterna:
0.030
Positive Predictive Value
Sfold:
0.535
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Alterna
Matthews Correlation Coefficient
MaxExpect:
0.465
Alterna:
0.129
Sensitivity
MaxExpect:
0.443
Alterna:
0.030
Positive Predictive Value
MaxExpect:
0.488
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Alterna
Matthews Correlation Coefficient
ProbKnot:
0.460
Alterna:
0.129
Sensitivity
ProbKnot:
0.446
Alterna:
0.030
Positive Predictive Value
ProbKnot:
0.475
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Alterna
Matthews Correlation Coefficient
UNAFold:
0.445
Alterna:
0.129
Sensitivity
UNAFold:
0.445
Alterna:
0.030
Positive Predictive Value
UNAFold:
0.446
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Alterna:
0.129
Sensitivity
CentroidAlifold(seed):
0.213
Alterna:
0.030
Positive Predictive Value
CentroidAlifold(seed):
0.901
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Alterna
Matthews Correlation Coefficient
Fold:
0.437
Alterna:
0.129
Sensitivity
Fold:
0.432
Alterna:
0.030
Positive Predictive Value
Fold:
0.444
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Alterna
Matthews Correlation Coefficient
RNAfold:
0.429
Alterna:
0.129
Sensitivity
RNAfold:
0.422
Alterna:
0.030
Positive Predictive Value
RNAfold:
0.437
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Alterna
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Alterna:
0.129
Sensitivity
CentroidHomfold‑LAST:
0.216
Alterna:
0.030
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Alterna:
0.129
Sensitivity
PETfold_pre2.0(20):
0.211
Alterna:
0.030
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Alterna:
0.129
Sensitivity
CentroidAlifold(20):
0.190
Alterna:
0.030
Positive Predictive Value
CentroidAlifold(20):
0.891
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Alterna
Matthews Correlation Coefficient
PknotsRG:
0.386
Alterna:
0.129
Sensitivity
PknotsRG:
0.320
Alterna:
0.030
Positive Predictive Value
PknotsRG:
0.465
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Alterna
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Alterna:
0.129
Sensitivity
MXScarna(seed):
0.196
Alterna:
0.030
Positive Predictive Value
MXScarna(seed):
0.753
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Alterna
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Alterna:
0.129
Sensitivity
RNAalifold(20):
0.185
Alterna:
0.030
Positive Predictive Value
RNAalifold(20):
0.787
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Alterna
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Alterna:
0.129
Sensitivity
RNAalifold(seed):
0.169
Alterna:
0.030
Positive Predictive Value
RNAalifold(seed):
0.835
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Alterna
Matthews Correlation Coefficient
Afold:
0.358
Alterna:
0.129
Sensitivity
Afold:
0.295
Alterna:
0.030
Positive Predictive Value
Afold:
0.434
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Alterna
Matthews Correlation Coefficient
MXScarna(20):
0.359
Alterna:
0.129
Sensitivity
MXScarna(20):
0.181
Alterna:
0.030
Positive Predictive Value
MXScarna(20):
0.710
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Alterna
Matthews Correlation Coefficient
CRWrnafold:
0.346
Alterna:
0.129
Sensitivity
CRWrnafold:
0.275
Alterna:
0.030
Positive Predictive Value
CRWrnafold:
0.435
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Alterna
Matthews Correlation Coefficient
RNAsubopt:
0.336
Alterna:
0.129
Sensitivity
RNAsubopt:
0.217
Alterna:
0.030
Positive Predictive Value
RNAsubopt:
0.521
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Alterna
Matthews Correlation Coefficient
McQFold:
0.314
Alterna:
0.129
Sensitivity
McQFold:
0.205
Alterna:
0.030
Positive Predictive Value
McQFold:
0.480
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Alterna
Matthews Correlation Coefficient
TurboFold(20):
0.301
Alterna:
0.129
Sensitivity
TurboFold(20):
0.108
Alterna:
0.030
Positive Predictive Value
TurboFold(20):
0.837
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Alterna
Matthews Correlation Coefficient
PPfold(20):
0.298
Alterna:
0.129
Sensitivity
PPfold(20):
0.101
Alterna:
0.030
Positive Predictive Value
PPfold(20):
0.879
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Alterna
Matthews Correlation Coefficient
RNAshapes:
0.295
Alterna:
0.129
Sensitivity
RNAshapes:
0.180
Alterna:
0.030
Positive Predictive Value
RNAshapes:
0.484
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Alterna
Matthews Correlation Coefficient
Carnac(20):
0.288
Alterna:
0.129
Sensitivity
Carnac(20):
0.093
Alterna:
0.030
Positive Predictive Value
Carnac(20):
0.892
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Alterna
Matthews Correlation Coefficient
RNASLOpt:
0.281
Alterna:
0.129
Sensitivity
RNASLOpt:
0.158
Alterna:
0.030
Positive Predictive Value
RNASLOpt:
0.502
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Alterna
Matthews Correlation Coefficient
RSpredict(20):
0.270
Alterna:
0.129
Sensitivity
RSpredict(20):
0.124
Alterna:
0.030
Positive Predictive Value
RSpredict(20):
0.590
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Alterna
Matthews Correlation Coefficient
Multilign(20):
0.251
Alterna:
0.129
Sensitivity
Multilign(20):
0.083
Alterna:
0.030
Positive Predictive Value
Multilign(20):
0.762
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Alterna
Matthews Correlation Coefficient
RNAwolf:
0.251
Alterna:
0.129
Sensitivity
RNAwolf:
0.191
Alterna:
0.030
Positive Predictive Value
RNAwolf:
0.330
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Alterna
Matthews Correlation Coefficient
Murlet(20):
0.230
Alterna:
0.129
Sensitivity
Murlet(20):
0.067
Alterna:
0.030
Positive Predictive Value
Murlet(20):
0.787
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Alterna
Matthews Correlation Coefficient
Vsfold4:
0.217
Alterna:
0.129
Sensitivity
Vsfold4:
0.125
Alterna:
0.030
Positive Predictive Value
Vsfold4:
0.378
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Alterna
Matthews Correlation Coefficient
Vsfold5:
0.211
Alterna:
0.129
Sensitivity
Vsfold5:
0.125
Alterna:
0.030
Positive Predictive Value
Vsfold5:
0.359
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Alterna
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Alterna:
0.129
Sensitivity
RSpredict(seed):
0.075
Alterna:
0.030
Positive Predictive Value
RSpredict(seed):
0.597
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Alterna
Matthews Correlation Coefficient
CMfinder(20):
0.210
Alterna:
0.129
Sensitivity
CMfinder(20):
0.055
Alterna:
0.030
Positive Predictive Value
CMfinder(20):
0.794
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Alterna
Matthews Correlation Coefficient
HotKnots:
0.179
Alterna:
0.129
Sensitivity
HotKnots:
0.051
Alterna:
0.030
Positive Predictive Value
HotKnots:
0.624
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Alterna
Matthews Correlation Coefficient
RNASampler(20):
0.176
Alterna:
0.129
Sensitivity
RNASampler(20):
0.035
Alterna:
0.030
Positive Predictive Value
RNASampler(20):
0.877
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Alterna
Matthews Correlation Coefficient
Cylofold:
0.155
Alterna:
0.129
Sensitivity
Cylofold:
0.040
Alterna:
0.030
Positive Predictive Value
Cylofold:
0.606
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs Alterna
Matthews Correlation Coefficient
Pknots:
0.143
Alterna:
0.129
Sensitivity
Pknots:
0.033
Alterna:
0.030
Positive Predictive Value
Pknots:
0.624
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs Alterna
Matthews Correlation Coefficient
Mastr(20):
0.134
Alterna:
0.129
Sensitivity
Mastr(20):
0.023
Alterna:
0.030
Positive Predictive Value
Mastr(20):
0.790
Alterna:
0.566
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Alterna vs MCFold
Matthews Correlation Coefficient
Alterna:
0.129
MCFold:
0.125
Sensitivity
Alterna:
0.030
MCFold:
0.036
Positive Predictive Value
Alterna:
0.566
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs RDfolder
Matthews Correlation Coefficient
Alterna:
0.129
RDfolder:
0.122
Sensitivity
Alterna:
0.030
RDfolder:
0.025
Positive Predictive Value
Alterna:
0.566
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Murlet(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Murlet(seed):
0.079
Sensitivity
Alterna:
0.030
Murlet(seed):
0.008
Positive Predictive Value
Alterna:
0.566
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs CMfinder(seed)
Matthews Correlation Coefficient
Alterna:
0.129
CMfinder(seed):
0.080
Sensitivity
Alterna:
0.030
CMfinder(seed):
0.008
Positive Predictive Value
Alterna:
0.566
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs TurboFold(seed)
Matthews Correlation Coefficient
Alterna:
0.129
TurboFold(seed):
0.076
Sensitivity
Alterna:
0.030
TurboFold(seed):
0.007
Positive Predictive Value
Alterna:
0.566
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs PPfold(seed)
Matthews Correlation Coefficient
Alterna:
0.129
PPfold(seed):
0.069
Sensitivity
Alterna:
0.030
PPfold(seed):
0.006
Positive Predictive Value
Alterna:
0.566
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Carnac(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Carnac(seed):
0.066
Sensitivity
Alterna:
0.030
Carnac(seed):
0.005
Positive Predictive Value
Alterna:
0.566
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs RNASampler(seed)
Matthews Correlation Coefficient
Alterna:
0.129
RNASampler(seed):
0.063
Sensitivity
Alterna:
0.030
RNASampler(seed):
0.005
Positive Predictive Value
Alterna:
0.566
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs NanoFolder
Matthews Correlation Coefficient
Alterna:
0.129
NanoFolder:
0.052
Sensitivity
Alterna:
0.030
NanoFolder:
0.016
Positive Predictive Value
Alterna:
0.566
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Mastr(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Mastr(seed):
0.051
Sensitivity
Alterna:
0.030
Mastr(seed):
0.003
Positive Predictive Value
Alterna:
0.566
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Multilign(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Multilign(seed):
0.043
Sensitivity
Alterna:
0.030
Multilign(seed):
0.003
Positive Predictive Value
Alterna:
0.566
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MCFold |
1987
ContextFold vs MCFold
Matthews Correlation Coefficient
ContextFold:
0.680
MCFold:
0.125
Sensitivity
ContextFold:
0.644
MCFold:
0.036
Positive Predictive Value
ContextFold:
0.718
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs MCFold
Matthews Correlation Coefficient
IPknot:
0.548
MCFold:
0.125
Sensitivity
IPknot:
0.475
MCFold:
0.036
Positive Predictive Value
IPknot:
0.632
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
MCFold:
0.125
Sensitivity
PETfold_pre2.0(seed):
0.343
MCFold:
0.036
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs MCFold
Matthews Correlation Coefficient
Contrafold:
0.508
MCFold:
0.125
Sensitivity
Contrafold:
0.481
MCFold:
0.036
Positive Predictive Value
Contrafold:
0.537
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs MCFold
Matthews Correlation Coefficient
CentroidFold:
0.470
MCFold:
0.125
Sensitivity
CentroidFold:
0.369
MCFold:
0.036
Positive Predictive Value
CentroidFold:
0.600
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs MCFold
Matthews Correlation Coefficient
Sfold:
0.469
MCFold:
0.125
Sensitivity
Sfold:
0.412
MCFold:
0.036
Positive Predictive Value
Sfold:
0.535
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs MCFold
Matthews Correlation Coefficient
MaxExpect:
0.465
MCFold:
0.125
Sensitivity
MaxExpect:
0.443
MCFold:
0.036
Positive Predictive Value
MaxExpect:
0.488
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs MCFold
Matthews Correlation Coefficient
ProbKnot:
0.460
MCFold:
0.125
Sensitivity
ProbKnot:
0.446
MCFold:
0.036
Positive Predictive Value
ProbKnot:
0.475
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs MCFold
Matthews Correlation Coefficient
UNAFold:
0.445
MCFold:
0.125
Sensitivity
UNAFold:
0.445
MCFold:
0.036
Positive Predictive Value
UNAFold:
0.446
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
MCFold:
0.125
Sensitivity
CentroidAlifold(seed):
0.213
MCFold:
0.036
Positive Predictive Value
CentroidAlifold(seed):
0.901
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs MCFold
Matthews Correlation Coefficient
Fold:
0.437
MCFold:
0.125
Sensitivity
Fold:
0.432
MCFold:
0.036
Positive Predictive Value
Fold:
0.444
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs MCFold
Matthews Correlation Coefficient
RNAfold:
0.429
MCFold:
0.125
Sensitivity
RNAfold:
0.422
MCFold:
0.036
Positive Predictive Value
RNAfold:
0.437
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs MCFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
MCFold:
0.125
Sensitivity
CentroidHomfold‑LAST:
0.216
MCFold:
0.036
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
MCFold:
0.125
Sensitivity
PETfold_pre2.0(20):
0.211
MCFold:
0.036
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
MCFold:
0.125
Sensitivity
CentroidAlifold(20):
0.190
MCFold:
0.036
Positive Predictive Value
CentroidAlifold(20):
0.891
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs MCFold
Matthews Correlation Coefficient
PknotsRG:
0.386
MCFold:
0.125
Sensitivity
PknotsRG:
0.320
MCFold:
0.036
Positive Predictive Value
PknotsRG:
0.465
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs MCFold
Matthews Correlation Coefficient
MXScarna(seed):
0.385
MCFold:
0.125
Sensitivity
MXScarna(seed):
0.196
MCFold:
0.036
Positive Predictive Value
MXScarna(seed):
0.753
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs MCFold
Matthews Correlation Coefficient
RNAalifold(20):
0.381
MCFold:
0.125
Sensitivity
RNAalifold(20):
0.185
MCFold:
0.036
Positive Predictive Value
RNAalifold(20):
0.787
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs MCFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
MCFold:
0.125
Sensitivity
RNAalifold(seed):
0.169
MCFold:
0.036
Positive Predictive Value
RNAalifold(seed):
0.835
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs MCFold
Matthews Correlation Coefficient
Afold:
0.358
MCFold:
0.125
Sensitivity
Afold:
0.295
MCFold:
0.036
Positive Predictive Value
Afold:
0.434
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs MCFold
Matthews Correlation Coefficient
MXScarna(20):
0.359
MCFold:
0.125
Sensitivity
MXScarna(20):
0.181
MCFold:
0.036
Positive Predictive Value
MXScarna(20):
0.710
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs MCFold
Matthews Correlation Coefficient
CRWrnafold:
0.346
MCFold:
0.125
Sensitivity
CRWrnafold:
0.275
MCFold:
0.036
Positive Predictive Value
CRWrnafold:
0.435
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs MCFold
Matthews Correlation Coefficient
RNAsubopt:
0.336
MCFold:
0.125
Sensitivity
RNAsubopt:
0.217
MCFold:
0.036
Positive Predictive Value
RNAsubopt:
0.521
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs MCFold
Matthews Correlation Coefficient
McQFold:
0.314
MCFold:
0.125
Sensitivity
McQFold:
0.205
MCFold:
0.036
Positive Predictive Value
McQFold:
0.480
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs MCFold
Matthews Correlation Coefficient
TurboFold(20):
0.301
MCFold:
0.125
Sensitivity
TurboFold(20):
0.108
MCFold:
0.036
Positive Predictive Value
TurboFold(20):
0.837
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs MCFold
Matthews Correlation Coefficient
PPfold(20):
0.298
MCFold:
0.125
Sensitivity
PPfold(20):
0.101
MCFold:
0.036
Positive Predictive Value
PPfold(20):
0.879
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs MCFold
Matthews Correlation Coefficient
RNAshapes:
0.295
MCFold:
0.125
Sensitivity
RNAshapes:
0.180
MCFold:
0.036
Positive Predictive Value
RNAshapes:
0.484
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs MCFold
Matthews Correlation Coefficient
Carnac(20):
0.288
MCFold:
0.125
Sensitivity
Carnac(20):
0.093
MCFold:
0.036
Positive Predictive Value
Carnac(20):
0.892
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs MCFold
Matthews Correlation Coefficient
RNASLOpt:
0.281
MCFold:
0.125
Sensitivity
RNASLOpt:
0.158
MCFold:
0.036
Positive Predictive Value
RNASLOpt:
0.502
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs MCFold
Matthews Correlation Coefficient
RSpredict(20):
0.270
MCFold:
0.125
Sensitivity
RSpredict(20):
0.124
MCFold:
0.036
Positive Predictive Value
RSpredict(20):
0.590
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs MCFold
Matthews Correlation Coefficient
Multilign(20):
0.251
MCFold:
0.125
Sensitivity
Multilign(20):
0.083
MCFold:
0.036
Positive Predictive Value
Multilign(20):
0.762
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs MCFold
Matthews Correlation Coefficient
RNAwolf:
0.251
MCFold:
0.125
Sensitivity
RNAwolf:
0.191
MCFold:
0.036
Positive Predictive Value
RNAwolf:
0.330
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs MCFold
Matthews Correlation Coefficient
Murlet(20):
0.230
MCFold:
0.125
Sensitivity
Murlet(20):
0.067
MCFold:
0.036
Positive Predictive Value
Murlet(20):
0.787
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs MCFold
Matthews Correlation Coefficient
Vsfold4:
0.217
MCFold:
0.125
Sensitivity
Vsfold4:
0.125
MCFold:
0.036
Positive Predictive Value
Vsfold4:
0.378
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs MCFold
Matthews Correlation Coefficient
Vsfold5:
0.211
MCFold:
0.125
Sensitivity
Vsfold5:
0.125
MCFold:
0.036
Positive Predictive Value
Vsfold5:
0.359
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs MCFold
Matthews Correlation Coefficient
RSpredict(seed):
0.212
MCFold:
0.125
Sensitivity
RSpredict(seed):
0.075
MCFold:
0.036
Positive Predictive Value
RSpredict(seed):
0.597
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs MCFold
Matthews Correlation Coefficient
CMfinder(20):
0.210
MCFold:
0.125
Sensitivity
CMfinder(20):
0.055
MCFold:
0.036
Positive Predictive Value
CMfinder(20):
0.794
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs MCFold
Matthews Correlation Coefficient
HotKnots:
0.179
MCFold:
0.125
Sensitivity
HotKnots:
0.051
MCFold:
0.036
Positive Predictive Value
HotKnots:
0.624
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs MCFold
Matthews Correlation Coefficient
RNASampler(20):
0.176
MCFold:
0.125
Sensitivity
RNASampler(20):
0.035
MCFold:
0.036
Positive Predictive Value
RNASampler(20):
0.877
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs MCFold
Matthews Correlation Coefficient
Cylofold:
0.155
MCFold:
0.125
Sensitivity
Cylofold:
0.040
MCFold:
0.036
Positive Predictive Value
Cylofold:
0.606
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs MCFold
Matthews Correlation Coefficient
Pknots:
0.143
MCFold:
0.125
Sensitivity
Pknots:
0.033
MCFold:
0.036
Positive Predictive Value
Pknots:
0.624
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs MCFold
Matthews Correlation Coefficient
Mastr(20):
0.134
MCFold:
0.125
Sensitivity
Mastr(20):
0.023
MCFold:
0.036
Positive Predictive Value
Mastr(20):
0.790
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs MCFold
Matthews Correlation Coefficient
Alterna:
0.129
MCFold:
0.125
Sensitivity
Alterna:
0.030
MCFold:
0.036
Positive Predictive Value
Alterna:
0.566
MCFold:
0.435
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
MCFold vs RDfolder
Matthews Correlation Coefficient
MCFold:
0.125
RDfolder:
0.122
Sensitivity
MCFold:
0.036
RDfolder:
0.025
Positive Predictive Value
MCFold:
0.435
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Murlet(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Murlet(seed):
0.079
Sensitivity
MCFold:
0.036
Murlet(seed):
0.008
Positive Predictive Value
MCFold:
0.435
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs CMfinder(seed)
Matthews Correlation Coefficient
MCFold:
0.125
CMfinder(seed):
0.080
Sensitivity
MCFold:
0.036
CMfinder(seed):
0.008
Positive Predictive Value
MCFold:
0.435
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs TurboFold(seed)
Matthews Correlation Coefficient
MCFold:
0.125
TurboFold(seed):
0.076
Sensitivity
MCFold:
0.036
TurboFold(seed):
0.007
Positive Predictive Value
MCFold:
0.435
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs PPfold(seed)
Matthews Correlation Coefficient
MCFold:
0.125
PPfold(seed):
0.069
Sensitivity
MCFold:
0.036
PPfold(seed):
0.006
Positive Predictive Value
MCFold:
0.435
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Carnac(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Carnac(seed):
0.066
Sensitivity
MCFold:
0.036
Carnac(seed):
0.005
Positive Predictive Value
MCFold:
0.435
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs RNASampler(seed)
Matthews Correlation Coefficient
MCFold:
0.125
RNASampler(seed):
0.063
Sensitivity
MCFold:
0.036
RNASampler(seed):
0.005
Positive Predictive Value
MCFold:
0.435
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs NanoFolder
Matthews Correlation Coefficient
MCFold:
0.125
NanoFolder:
0.052
Sensitivity
MCFold:
0.036
NanoFolder:
0.016
Positive Predictive Value
MCFold:
0.435
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Mastr(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Mastr(seed):
0.051
Sensitivity
MCFold:
0.036
Mastr(seed):
0.003
Positive Predictive Value
MCFold:
0.435
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Multilign(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Multilign(seed):
0.043
Sensitivity
MCFold:
0.036
Multilign(seed):
0.003
Positive Predictive Value
MCFold:
0.435
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RDfolder |
1987
ContextFold vs RDfolder
Matthews Correlation Coefficient
ContextFold:
0.680
RDfolder:
0.122
Sensitivity
ContextFold:
0.644
RDfolder:
0.025
Positive Predictive Value
ContextFold:
0.718
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RDfolder
Matthews Correlation Coefficient
IPknot:
0.548
RDfolder:
0.122
Sensitivity
IPknot:
0.475
RDfolder:
0.025
Positive Predictive Value
IPknot:
0.632
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RDfolder:
0.122
Sensitivity
PETfold_pre2.0(seed):
0.343
RDfolder:
0.025
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RDfolder
Matthews Correlation Coefficient
Contrafold:
0.508
RDfolder:
0.122
Sensitivity
Contrafold:
0.481
RDfolder:
0.025
Positive Predictive Value
Contrafold:
0.537
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RDfolder
Matthews Correlation Coefficient
CentroidFold:
0.470
RDfolder:
0.122
Sensitivity
CentroidFold:
0.369
RDfolder:
0.025
Positive Predictive Value
CentroidFold:
0.600
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RDfolder
Matthews Correlation Coefficient
Sfold:
0.469
RDfolder:
0.122
Sensitivity
Sfold:
0.412
RDfolder:
0.025
Positive Predictive Value
Sfold:
0.535
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RDfolder
Matthews Correlation Coefficient
MaxExpect:
0.465
RDfolder:
0.122
Sensitivity
MaxExpect:
0.443
RDfolder:
0.025
Positive Predictive Value
MaxExpect:
0.488
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RDfolder
Matthews Correlation Coefficient
ProbKnot:
0.460
RDfolder:
0.122
Sensitivity
ProbKnot:
0.446
RDfolder:
0.025
Positive Predictive Value
ProbKnot:
0.475
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RDfolder
Matthews Correlation Coefficient
UNAFold:
0.445
RDfolder:
0.122
Sensitivity
UNAFold:
0.445
RDfolder:
0.025
Positive Predictive Value
UNAFold:
0.446
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RDfolder:
0.122
Sensitivity
CentroidAlifold(seed):
0.213
RDfolder:
0.025
Positive Predictive Value
CentroidAlifold(seed):
0.901
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RDfolder
Matthews Correlation Coefficient
Fold:
0.437
RDfolder:
0.122
Sensitivity
Fold:
0.432
RDfolder:
0.025
Positive Predictive Value
Fold:
0.444
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RDfolder
Matthews Correlation Coefficient
RNAfold:
0.429
RDfolder:
0.122
Sensitivity
RNAfold:
0.422
RDfolder:
0.025
Positive Predictive Value
RNAfold:
0.437
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RDfolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RDfolder:
0.122
Sensitivity
CentroidHomfold‑LAST:
0.216
RDfolder:
0.025
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RDfolder:
0.122
Sensitivity
PETfold_pre2.0(20):
0.211
RDfolder:
0.025
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RDfolder:
0.122
Sensitivity
CentroidAlifold(20):
0.190
RDfolder:
0.025
Positive Predictive Value
CentroidAlifold(20):
0.891
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RDfolder
Matthews Correlation Coefficient
PknotsRG:
0.386
RDfolder:
0.122
Sensitivity
PknotsRG:
0.320
RDfolder:
0.025
Positive Predictive Value
PknotsRG:
0.465
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RDfolder
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RDfolder:
0.122
Sensitivity
MXScarna(seed):
0.196
RDfolder:
0.025
Positive Predictive Value
MXScarna(seed):
0.753
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RDfolder:
0.122
Sensitivity
RNAalifold(20):
0.185
RDfolder:
0.025
Positive Predictive Value
RNAalifold(20):
0.787
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RDfolder:
0.122
Sensitivity
RNAalifold(seed):
0.169
RDfolder:
0.025
Positive Predictive Value
RNAalifold(seed):
0.835
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RDfolder
Matthews Correlation Coefficient
Afold:
0.358
RDfolder:
0.122
Sensitivity
Afold:
0.295
RDfolder:
0.025
Positive Predictive Value
Afold:
0.434
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RDfolder
Matthews Correlation Coefficient
MXScarna(20):
0.359
RDfolder:
0.122
Sensitivity
MXScarna(20):
0.181
RDfolder:
0.025
Positive Predictive Value
MXScarna(20):
0.710
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RDfolder
Matthews Correlation Coefficient
CRWrnafold:
0.346
RDfolder:
0.122
Sensitivity
CRWrnafold:
0.275
RDfolder:
0.025
Positive Predictive Value
CRWrnafold:
0.435
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RDfolder
Matthews Correlation Coefficient
RNAsubopt:
0.336
RDfolder:
0.122
Sensitivity
RNAsubopt:
0.217
RDfolder:
0.025
Positive Predictive Value
RNAsubopt:
0.521
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RDfolder
Matthews Correlation Coefficient
McQFold:
0.314
RDfolder:
0.122
Sensitivity
McQFold:
0.205
RDfolder:
0.025
Positive Predictive Value
McQFold:
0.480
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RDfolder
Matthews Correlation Coefficient
TurboFold(20):
0.301
RDfolder:
0.122
Sensitivity
TurboFold(20):
0.108
RDfolder:
0.025
Positive Predictive Value
TurboFold(20):
0.837
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RDfolder
Matthews Correlation Coefficient
PPfold(20):
0.298
RDfolder:
0.122
Sensitivity
PPfold(20):
0.101
RDfolder:
0.025
Positive Predictive Value
PPfold(20):
0.879
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RDfolder
Matthews Correlation Coefficient
RNAshapes:
0.295
RDfolder:
0.122
Sensitivity
RNAshapes:
0.180
RDfolder:
0.025
Positive Predictive Value
RNAshapes:
0.484
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RDfolder
Matthews Correlation Coefficient
Carnac(20):
0.288
RDfolder:
0.122
Sensitivity
Carnac(20):
0.093
RDfolder:
0.025
Positive Predictive Value
Carnac(20):
0.892
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs RDfolder
Matthews Correlation Coefficient
RNASLOpt:
0.281
RDfolder:
0.122
Sensitivity
RNASLOpt:
0.158
RDfolder:
0.025
Positive Predictive Value
RNASLOpt:
0.502
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs RDfolder
Matthews Correlation Coefficient
RSpredict(20):
0.270
RDfolder:
0.122
Sensitivity
RSpredict(20):
0.124
RDfolder:
0.025
Positive Predictive Value
RSpredict(20):
0.590
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs RDfolder
Matthews Correlation Coefficient
Multilign(20):
0.251
RDfolder:
0.122
Sensitivity
Multilign(20):
0.083
RDfolder:
0.025
Positive Predictive Value
Multilign(20):
0.762
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs RDfolder
Matthews Correlation Coefficient
RNAwolf:
0.251
RDfolder:
0.122
Sensitivity
RNAwolf:
0.191
RDfolder:
0.025
Positive Predictive Value
RNAwolf:
0.330
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs RDfolder
Matthews Correlation Coefficient
Murlet(20):
0.230
RDfolder:
0.122
Sensitivity
Murlet(20):
0.067
RDfolder:
0.025
Positive Predictive Value
Murlet(20):
0.787
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs RDfolder
Matthews Correlation Coefficient
Vsfold4:
0.217
RDfolder:
0.122
Sensitivity
Vsfold4:
0.125
RDfolder:
0.025
Positive Predictive Value
Vsfold4:
0.378
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs RDfolder
Matthews Correlation Coefficient
Vsfold5:
0.211
RDfolder:
0.122
Sensitivity
Vsfold5:
0.125
RDfolder:
0.025
Positive Predictive Value
Vsfold5:
0.359
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs RDfolder
Matthews Correlation Coefficient
RSpredict(seed):
0.212
RDfolder:
0.122
Sensitivity
RSpredict(seed):
0.075
RDfolder:
0.025
Positive Predictive Value
RSpredict(seed):
0.597
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs RDfolder
Matthews Correlation Coefficient
CMfinder(20):
0.210
RDfolder:
0.122
Sensitivity
CMfinder(20):
0.055
RDfolder:
0.025
Positive Predictive Value
CMfinder(20):
0.794
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs RDfolder
Matthews Correlation Coefficient
HotKnots:
0.179
RDfolder:
0.122
Sensitivity
HotKnots:
0.051
RDfolder:
0.025
Positive Predictive Value
HotKnots:
0.624
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs RDfolder
Matthews Correlation Coefficient
RNASampler(20):
0.176
RDfolder:
0.122
Sensitivity
RNASampler(20):
0.035
RDfolder:
0.025
Positive Predictive Value
RNASampler(20):
0.877
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs RDfolder
Matthews Correlation Coefficient
Cylofold:
0.155
RDfolder:
0.122
Sensitivity
Cylofold:
0.040
RDfolder:
0.025
Positive Predictive Value
Cylofold:
0.606
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs RDfolder
Matthews Correlation Coefficient
Pknots:
0.143
RDfolder:
0.122
Sensitivity
Pknots:
0.033
RDfolder:
0.025
Positive Predictive Value
Pknots:
0.624
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs RDfolder
Matthews Correlation Coefficient
Mastr(20):
0.134
RDfolder:
0.122
Sensitivity
Mastr(20):
0.023
RDfolder:
0.025
Positive Predictive Value
Mastr(20):
0.790
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs RDfolder
Matthews Correlation Coefficient
Alterna:
0.129
RDfolder:
0.122
Sensitivity
Alterna:
0.030
RDfolder:
0.025
Positive Predictive Value
Alterna:
0.566
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs RDfolder
Matthews Correlation Coefficient
MCFold:
0.125
RDfolder:
0.122
Sensitivity
MCFold:
0.036
RDfolder:
0.025
Positive Predictive Value
MCFold:
0.435
RDfolder:
0.592
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RDfolder vs Murlet(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Murlet(seed):
0.079
Sensitivity
RDfolder:
0.025
Murlet(seed):
0.008
Positive Predictive Value
RDfolder:
0.592
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs CMfinder(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
CMfinder(seed):
0.080
Sensitivity
RDfolder:
0.025
CMfinder(seed):
0.008
Positive Predictive Value
RDfolder:
0.592
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs TurboFold(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
TurboFold(seed):
0.076
Sensitivity
RDfolder:
0.025
TurboFold(seed):
0.007
Positive Predictive Value
RDfolder:
0.592
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs PPfold(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
PPfold(seed):
0.069
Sensitivity
RDfolder:
0.025
PPfold(seed):
0.006
Positive Predictive Value
RDfolder:
0.592
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs Carnac(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Carnac(seed):
0.066
Sensitivity
RDfolder:
0.025
Carnac(seed):
0.005
Positive Predictive Value
RDfolder:
0.592
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs RNASampler(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
RNASampler(seed):
0.063
Sensitivity
RDfolder:
0.025
RNASampler(seed):
0.005
Positive Predictive Value
RDfolder:
0.592
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs NanoFolder
Matthews Correlation Coefficient
RDfolder:
0.122
NanoFolder:
0.052
Sensitivity
RDfolder:
0.025
NanoFolder:
0.016
Positive Predictive Value
RDfolder:
0.592
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs Mastr(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Mastr(seed):
0.051
Sensitivity
RDfolder:
0.025
Mastr(seed):
0.003
Positive Predictive Value
RDfolder:
0.592
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs Multilign(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Multilign(seed):
0.043
Sensitivity
RDfolder:
0.025
Multilign(seed):
0.003
Positive Predictive Value
RDfolder:
0.592
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Murlet(seed) |
1987
ContextFold vs Murlet(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Murlet(seed):
0.079
Sensitivity
ContextFold:
0.644
Murlet(seed):
0.008
Positive Predictive Value
ContextFold:
0.718
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Murlet(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Murlet(seed):
0.079
Sensitivity
IPknot:
0.475
Murlet(seed):
0.008
Positive Predictive Value
IPknot:
0.632
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Murlet(seed):
0.079
Sensitivity
PETfold_pre2.0(seed):
0.343
Murlet(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Murlet(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Murlet(seed):
0.079
Sensitivity
Contrafold:
0.481
Murlet(seed):
0.008
Positive Predictive Value
Contrafold:
0.537
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Murlet(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Murlet(seed):
0.079
Sensitivity
CentroidFold:
0.369
Murlet(seed):
0.008
Positive Predictive Value
CentroidFold:
0.600
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Murlet(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Murlet(seed):
0.079
Sensitivity
Sfold:
0.412
Murlet(seed):
0.008
Positive Predictive Value
Sfold:
0.535
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Murlet(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Murlet(seed):
0.079
Sensitivity
MaxExpect:
0.443
Murlet(seed):
0.008
Positive Predictive Value
MaxExpect:
0.488
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Murlet(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Murlet(seed):
0.079
Sensitivity
ProbKnot:
0.446
Murlet(seed):
0.008
Positive Predictive Value
ProbKnot:
0.475
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Murlet(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Murlet(seed):
0.079
Sensitivity
UNAFold:
0.445
Murlet(seed):
0.008
Positive Predictive Value
UNAFold:
0.446
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Murlet(seed):
0.079
Sensitivity
CentroidAlifold(seed):
0.213
Murlet(seed):
0.008
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Murlet(seed)
Matthews Correlation Coefficient
Fold:
0.437
Murlet(seed):
0.079
Sensitivity
Fold:
0.432
Murlet(seed):
0.008
Positive Predictive Value
Fold:
0.444
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Murlet(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Murlet(seed):
0.079
Sensitivity
RNAfold:
0.422
Murlet(seed):
0.008
Positive Predictive Value
RNAfold:
0.437
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Murlet(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Murlet(seed):
0.079
Sensitivity
CentroidHomfold‑LAST:
0.216
Murlet(seed):
0.008
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Murlet(seed):
0.079
Sensitivity
PETfold_pre2.0(20):
0.211
Murlet(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Murlet(seed):
0.079
Sensitivity
CentroidAlifold(20):
0.190
Murlet(seed):
0.008
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Murlet(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Murlet(seed):
0.079
Sensitivity
PknotsRG:
0.320
Murlet(seed):
0.008
Positive Predictive Value
PknotsRG:
0.465
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Murlet(seed):
0.079
Sensitivity
MXScarna(seed):
0.196
Murlet(seed):
0.008
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Murlet(seed):
0.079
Sensitivity
RNAalifold(20):
0.185
Murlet(seed):
0.008
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Murlet(seed):
0.079
Sensitivity
RNAalifold(seed):
0.169
Murlet(seed):
0.008
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Murlet(seed)
Matthews Correlation Coefficient
Afold:
0.358
Murlet(seed):
0.079
Sensitivity
Afold:
0.295
Murlet(seed):
0.008
Positive Predictive Value
Afold:
0.434
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Murlet(seed):
0.079
Sensitivity
MXScarna(20):
0.181
Murlet(seed):
0.008
Positive Predictive Value
MXScarna(20):
0.710
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Murlet(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Murlet(seed):
0.079
Sensitivity
CRWrnafold:
0.275
Murlet(seed):
0.008
Positive Predictive Value
CRWrnafold:
0.435
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Murlet(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Murlet(seed):
0.079
Sensitivity
RNAsubopt:
0.217
Murlet(seed):
0.008
Positive Predictive Value
RNAsubopt:
0.521
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Murlet(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Murlet(seed):
0.079
Sensitivity
McQFold:
0.205
Murlet(seed):
0.008
Positive Predictive Value
McQFold:
0.480
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Murlet(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Murlet(seed):
0.079
Sensitivity
TurboFold(20):
0.108
Murlet(seed):
0.008
Positive Predictive Value
TurboFold(20):
0.837
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Murlet(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Murlet(seed):
0.079
Sensitivity
PPfold(20):
0.101
Murlet(seed):
0.008
Positive Predictive Value
PPfold(20):
0.879
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Murlet(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Murlet(seed):
0.079
Sensitivity
RNAshapes:
0.180
Murlet(seed):
0.008
Positive Predictive Value
RNAshapes:
0.484
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Murlet(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Murlet(seed):
0.079
Sensitivity
Carnac(20):
0.093
Murlet(seed):
0.008
Positive Predictive Value
Carnac(20):
0.892
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Murlet(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Murlet(seed):
0.079
Sensitivity
RNASLOpt:
0.158
Murlet(seed):
0.008
Positive Predictive Value
RNASLOpt:
0.502
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Murlet(seed):
0.079
Sensitivity
RSpredict(20):
0.124
Murlet(seed):
0.008
Positive Predictive Value
RSpredict(20):
0.590
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Murlet(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Murlet(seed):
0.079
Sensitivity
Multilign(20):
0.083
Murlet(seed):
0.008
Positive Predictive Value
Multilign(20):
0.762
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Murlet(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Murlet(seed):
0.079
Sensitivity
RNAwolf:
0.191
Murlet(seed):
0.008
Positive Predictive Value
RNAwolf:
0.330
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Murlet(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Murlet(seed):
0.079
Sensitivity
Murlet(20):
0.067
Murlet(seed):
0.008
Positive Predictive Value
Murlet(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Murlet(seed):
0.079
Sensitivity
Vsfold4:
0.125
Murlet(seed):
0.008
Positive Predictive Value
Vsfold4:
0.378
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Murlet(seed):
0.079
Sensitivity
Vsfold5:
0.125
Murlet(seed):
0.008
Positive Predictive Value
Vsfold5:
0.359
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Murlet(seed):
0.079
Sensitivity
RSpredict(seed):
0.075
Murlet(seed):
0.008
Positive Predictive Value
RSpredict(seed):
0.597
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Murlet(seed):
0.079
Sensitivity
CMfinder(20):
0.055
Murlet(seed):
0.008
Positive Predictive Value
CMfinder(20):
0.794
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Murlet(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Murlet(seed):
0.079
Sensitivity
HotKnots:
0.051
Murlet(seed):
0.008
Positive Predictive Value
HotKnots:
0.624
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Murlet(seed):
0.079
Sensitivity
RNASampler(20):
0.035
Murlet(seed):
0.008
Positive Predictive Value
RNASampler(20):
0.877
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Murlet(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Murlet(seed):
0.079
Sensitivity
Cylofold:
0.040
Murlet(seed):
0.008
Positive Predictive Value
Cylofold:
0.606
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs Murlet(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Murlet(seed):
0.079
Sensitivity
Pknots:
0.033
Murlet(seed):
0.008
Positive Predictive Value
Pknots:
0.624
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs Murlet(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Murlet(seed):
0.079
Sensitivity
Mastr(20):
0.023
Murlet(seed):
0.008
Positive Predictive Value
Mastr(20):
0.790
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs Murlet(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Murlet(seed):
0.079
Sensitivity
Alterna:
0.030
Murlet(seed):
0.008
Positive Predictive Value
Alterna:
0.566
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs Murlet(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Murlet(seed):
0.079
Sensitivity
MCFold:
0.036
Murlet(seed):
0.008
Positive Predictive Value
MCFold:
0.435
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs Murlet(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Murlet(seed):
0.079
Sensitivity
RDfolder:
0.025
Murlet(seed):
0.008
Positive Predictive Value
RDfolder:
0.592
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
CMfinder(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Murlet(seed):
0.079
Sensitivity
CMfinder(seed):
0.008
Murlet(seed):
0.008
Positive Predictive Value
CMfinder(seed):
0.757
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.00238364898467
|
+
Murlet(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
TurboFold(seed):
0.076
Sensitivity
Murlet(seed):
0.008
TurboFold(seed):
0.007
Positive Predictive Value
Murlet(seed):
0.811
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 5.5212650266e-05
|
+
Murlet(seed) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
PPfold(seed):
0.069
Sensitivity
Murlet(seed):
0.008
PPfold(seed):
0.006
Positive Predictive Value
Murlet(seed):
0.811
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
Carnac(seed):
0.066
Sensitivity
Murlet(seed):
0.008
Carnac(seed):
0.005
Positive Predictive Value
Murlet(seed):
0.811
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
RNASampler(seed):
0.063
Sensitivity
Murlet(seed):
0.008
RNASampler(seed):
0.005
Positive Predictive Value
Murlet(seed):
0.811
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs NanoFolder
Matthews Correlation Coefficient
Murlet(seed):
0.079
NanoFolder:
0.052
Sensitivity
Murlet(seed):
0.008
NanoFolder:
0.016
Positive Predictive Value
Murlet(seed):
0.811
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
Mastr(seed):
0.051
Sensitivity
Murlet(seed):
0.008
Mastr(seed):
0.003
Positive Predictive Value
Murlet(seed):
0.811
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
Multilign(seed):
0.043
Sensitivity
Murlet(seed):
0.008
Multilign(seed):
0.003
Positive Predictive Value
Murlet(seed):
0.811
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CMfinder(seed) |
1987
ContextFold vs CMfinder(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
CMfinder(seed):
0.080
Sensitivity
ContextFold:
0.644
CMfinder(seed):
0.008
Positive Predictive Value
ContextFold:
0.718
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs CMfinder(seed)
Matthews Correlation Coefficient
IPknot:
0.548
CMfinder(seed):
0.080
Sensitivity
IPknot:
0.475
CMfinder(seed):
0.008
Positive Predictive Value
IPknot:
0.632
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
CMfinder(seed):
0.080
Sensitivity
PETfold_pre2.0(seed):
0.343
CMfinder(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs CMfinder(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
CMfinder(seed):
0.080
Sensitivity
Contrafold:
0.481
CMfinder(seed):
0.008
Positive Predictive Value
Contrafold:
0.537
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
CMfinder(seed):
0.080
Sensitivity
CentroidFold:
0.369
CMfinder(seed):
0.008
Positive Predictive Value
CentroidFold:
0.600
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs CMfinder(seed)
Matthews Correlation Coefficient
Sfold:
0.469
CMfinder(seed):
0.080
Sensitivity
Sfold:
0.412
CMfinder(seed):
0.008
Positive Predictive Value
Sfold:
0.535
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs CMfinder(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
CMfinder(seed):
0.080
Sensitivity
MaxExpect:
0.443
CMfinder(seed):
0.008
Positive Predictive Value
MaxExpect:
0.488
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs CMfinder(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
CMfinder(seed):
0.080
Sensitivity
ProbKnot:
0.446
CMfinder(seed):
0.008
Positive Predictive Value
ProbKnot:
0.475
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs CMfinder(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
CMfinder(seed):
0.080
Sensitivity
UNAFold:
0.445
CMfinder(seed):
0.008
Positive Predictive Value
UNAFold:
0.446
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
CMfinder(seed):
0.080
Sensitivity
CentroidAlifold(seed):
0.213
CMfinder(seed):
0.008
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs CMfinder(seed)
Matthews Correlation Coefficient
Fold:
0.437
CMfinder(seed):
0.080
Sensitivity
Fold:
0.432
CMfinder(seed):
0.008
Positive Predictive Value
Fold:
0.444
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs CMfinder(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
CMfinder(seed):
0.080
Sensitivity
RNAfold:
0.422
CMfinder(seed):
0.008
Positive Predictive Value
RNAfold:
0.437
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
CMfinder(seed):
0.080
Sensitivity
CentroidHomfold‑LAST:
0.216
CMfinder(seed):
0.008
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
CMfinder(seed):
0.080
Sensitivity
PETfold_pre2.0(20):
0.211
CMfinder(seed):
0.008
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
CMfinder(seed):
0.080
Sensitivity
CentroidAlifold(20):
0.190
CMfinder(seed):
0.008
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs CMfinder(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
CMfinder(seed):
0.080
Sensitivity
PknotsRG:
0.320
CMfinder(seed):
0.008
Positive Predictive Value
PknotsRG:
0.465
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
CMfinder(seed):
0.080
Sensitivity
MXScarna(seed):
0.196
CMfinder(seed):
0.008
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
CMfinder(seed):
0.080
Sensitivity
RNAalifold(20):
0.185
CMfinder(seed):
0.008
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
CMfinder(seed):
0.080
Sensitivity
RNAalifold(seed):
0.169
CMfinder(seed):
0.008
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs CMfinder(seed)
Matthews Correlation Coefficient
Afold:
0.358
CMfinder(seed):
0.080
Sensitivity
Afold:
0.295
CMfinder(seed):
0.008
Positive Predictive Value
Afold:
0.434
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
CMfinder(seed):
0.080
Sensitivity
MXScarna(20):
0.181
CMfinder(seed):
0.008
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs CMfinder(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
CMfinder(seed):
0.080
Sensitivity
CRWrnafold:
0.275
CMfinder(seed):
0.008
Positive Predictive Value
CRWrnafold:
0.435
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs CMfinder(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
CMfinder(seed):
0.080
Sensitivity
RNAsubopt:
0.217
CMfinder(seed):
0.008
Positive Predictive Value
RNAsubopt:
0.521
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs CMfinder(seed)
Matthews Correlation Coefficient
McQFold:
0.314
CMfinder(seed):
0.080
Sensitivity
McQFold:
0.205
CMfinder(seed):
0.008
Positive Predictive Value
McQFold:
0.480
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
CMfinder(seed):
0.080
Sensitivity
TurboFold(20):
0.108
CMfinder(seed):
0.008
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
CMfinder(seed):
0.080
Sensitivity
PPfold(20):
0.101
CMfinder(seed):
0.008
Positive Predictive Value
PPfold(20):
0.879
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs CMfinder(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
CMfinder(seed):
0.080
Sensitivity
RNAshapes:
0.180
CMfinder(seed):
0.008
Positive Predictive Value
RNAshapes:
0.484
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
CMfinder(seed):
0.080
Sensitivity
Carnac(20):
0.093
CMfinder(seed):
0.008
Positive Predictive Value
Carnac(20):
0.892
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs CMfinder(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
CMfinder(seed):
0.080
Sensitivity
RNASLOpt:
0.158
CMfinder(seed):
0.008
Positive Predictive Value
RNASLOpt:
0.502
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
CMfinder(seed):
0.080
Sensitivity
RSpredict(20):
0.124
CMfinder(seed):
0.008
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
CMfinder(seed):
0.080
Sensitivity
Multilign(20):
0.083
CMfinder(seed):
0.008
Positive Predictive Value
Multilign(20):
0.762
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs CMfinder(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
CMfinder(seed):
0.080
Sensitivity
RNAwolf:
0.191
CMfinder(seed):
0.008
Positive Predictive Value
RNAwolf:
0.330
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
CMfinder(seed):
0.080
Sensitivity
Murlet(20):
0.067
CMfinder(seed):
0.008
Positive Predictive Value
Murlet(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
CMfinder(seed):
0.080
Sensitivity
Vsfold4:
0.125
CMfinder(seed):
0.008
Positive Predictive Value
Vsfold4:
0.378
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
CMfinder(seed):
0.080
Sensitivity
Vsfold5:
0.125
CMfinder(seed):
0.008
Positive Predictive Value
Vsfold5:
0.359
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
CMfinder(seed):
0.080
Sensitivity
RSpredict(seed):
0.075
CMfinder(seed):
0.008
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
CMfinder(seed):
0.080
Sensitivity
CMfinder(20):
0.055
CMfinder(seed):
0.008
Positive Predictive Value
CMfinder(20):
0.794
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs CMfinder(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
CMfinder(seed):
0.080
Sensitivity
HotKnots:
0.051
CMfinder(seed):
0.008
Positive Predictive Value
HotKnots:
0.624
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
CMfinder(seed):
0.080
Sensitivity
RNASampler(20):
0.035
CMfinder(seed):
0.008
Positive Predictive Value
RNASampler(20):
0.877
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs CMfinder(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
CMfinder(seed):
0.080
Sensitivity
Cylofold:
0.040
CMfinder(seed):
0.008
Positive Predictive Value
Cylofold:
0.606
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs CMfinder(seed)
Matthews Correlation Coefficient
Pknots:
0.143
CMfinder(seed):
0.080
Sensitivity
Pknots:
0.033
CMfinder(seed):
0.008
Positive Predictive Value
Pknots:
0.624
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
CMfinder(seed):
0.080
Sensitivity
Mastr(20):
0.023
CMfinder(seed):
0.008
Positive Predictive Value
Mastr(20):
0.790
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs CMfinder(seed)
Matthews Correlation Coefficient
Alterna:
0.129
CMfinder(seed):
0.080
Sensitivity
Alterna:
0.030
CMfinder(seed):
0.008
Positive Predictive Value
Alterna:
0.566
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs CMfinder(seed)
Matthews Correlation Coefficient
MCFold:
0.125
CMfinder(seed):
0.080
Sensitivity
MCFold:
0.036
CMfinder(seed):
0.008
Positive Predictive Value
MCFold:
0.435
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs CMfinder(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
CMfinder(seed):
0.080
Sensitivity
RDfolder:
0.025
CMfinder(seed):
0.008
Positive Predictive Value
RDfolder:
0.592
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Murlet(seed):
0.079
Sensitivity
CMfinder(seed):
0.008
Murlet(seed):
0.008
Positive Predictive Value
CMfinder(seed):
0.757
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.00238364898467
|
|
+
CMfinder(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
TurboFold(seed):
0.076
Sensitivity
CMfinder(seed):
0.008
TurboFold(seed):
0.007
Positive Predictive Value
CMfinder(seed):
0.757
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 1.83595875043e-07
|
+
CMfinder(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
PPfold(seed):
0.069
Sensitivity
CMfinder(seed):
0.008
PPfold(seed):
0.006
Positive Predictive Value
CMfinder(seed):
0.757
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Carnac(seed):
0.066
Sensitivity
CMfinder(seed):
0.008
Carnac(seed):
0.005
Positive Predictive Value
CMfinder(seed):
0.757
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
RNASampler(seed):
0.063
Sensitivity
CMfinder(seed):
0.008
RNASampler(seed):
0.005
Positive Predictive Value
CMfinder(seed):
0.757
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(seed):
0.080
NanoFolder:
0.052
Sensitivity
CMfinder(seed):
0.008
NanoFolder:
0.016
Positive Predictive Value
CMfinder(seed):
0.757
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Mastr(seed):
0.051
Sensitivity
CMfinder(seed):
0.008
Mastr(seed):
0.003
Positive Predictive Value
CMfinder(seed):
0.757
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Multilign(seed):
0.043
Sensitivity
CMfinder(seed):
0.008
Multilign(seed):
0.003
Positive Predictive Value
CMfinder(seed):
0.757
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| TurboFold(seed) |
1987
ContextFold vs TurboFold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
TurboFold(seed):
0.076
Sensitivity
ContextFold:
0.644
TurboFold(seed):
0.007
Positive Predictive Value
ContextFold:
0.718
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs TurboFold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
TurboFold(seed):
0.076
Sensitivity
IPknot:
0.475
TurboFold(seed):
0.007
Positive Predictive Value
IPknot:
0.632
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
TurboFold(seed):
0.076
Sensitivity
PETfold_pre2.0(seed):
0.343
TurboFold(seed):
0.007
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs TurboFold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
TurboFold(seed):
0.076
Sensitivity
Contrafold:
0.481
TurboFold(seed):
0.007
Positive Predictive Value
Contrafold:
0.537
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
TurboFold(seed):
0.076
Sensitivity
CentroidFold:
0.369
TurboFold(seed):
0.007
Positive Predictive Value
CentroidFold:
0.600
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs TurboFold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
TurboFold(seed):
0.076
Sensitivity
Sfold:
0.412
TurboFold(seed):
0.007
Positive Predictive Value
Sfold:
0.535
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs TurboFold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
TurboFold(seed):
0.076
Sensitivity
MaxExpect:
0.443
TurboFold(seed):
0.007
Positive Predictive Value
MaxExpect:
0.488
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs TurboFold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
TurboFold(seed):
0.076
Sensitivity
ProbKnot:
0.446
TurboFold(seed):
0.007
Positive Predictive Value
ProbKnot:
0.475
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs TurboFold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
TurboFold(seed):
0.076
Sensitivity
UNAFold:
0.445
TurboFold(seed):
0.007
Positive Predictive Value
UNAFold:
0.446
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
TurboFold(seed):
0.076
Sensitivity
CentroidAlifold(seed):
0.213
TurboFold(seed):
0.007
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs TurboFold(seed)
Matthews Correlation Coefficient
Fold:
0.437
TurboFold(seed):
0.076
Sensitivity
Fold:
0.432
TurboFold(seed):
0.007
Positive Predictive Value
Fold:
0.444
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs TurboFold(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
TurboFold(seed):
0.076
Sensitivity
RNAfold:
0.422
TurboFold(seed):
0.007
Positive Predictive Value
RNAfold:
0.437
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
TurboFold(seed):
0.076
Sensitivity
CentroidHomfold‑LAST:
0.216
TurboFold(seed):
0.007
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
TurboFold(seed):
0.076
Sensitivity
PETfold_pre2.0(20):
0.211
TurboFold(seed):
0.007
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
TurboFold(seed):
0.076
Sensitivity
CentroidAlifold(20):
0.190
TurboFold(seed):
0.007
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs TurboFold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
TurboFold(seed):
0.076
Sensitivity
PknotsRG:
0.320
TurboFold(seed):
0.007
Positive Predictive Value
PknotsRG:
0.465
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
TurboFold(seed):
0.076
Sensitivity
MXScarna(seed):
0.196
TurboFold(seed):
0.007
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
TurboFold(seed):
0.076
Sensitivity
RNAalifold(20):
0.185
TurboFold(seed):
0.007
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
TurboFold(seed):
0.076
Sensitivity
RNAalifold(seed):
0.169
TurboFold(seed):
0.007
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs TurboFold(seed)
Matthews Correlation Coefficient
Afold:
0.358
TurboFold(seed):
0.076
Sensitivity
Afold:
0.295
TurboFold(seed):
0.007
Positive Predictive Value
Afold:
0.434
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
TurboFold(seed):
0.076
Sensitivity
MXScarna(20):
0.181
TurboFold(seed):
0.007
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs TurboFold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
TurboFold(seed):
0.076
Sensitivity
CRWrnafold:
0.275
TurboFold(seed):
0.007
Positive Predictive Value
CRWrnafold:
0.435
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs TurboFold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
TurboFold(seed):
0.076
Sensitivity
RNAsubopt:
0.217
TurboFold(seed):
0.007
Positive Predictive Value
RNAsubopt:
0.521
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs TurboFold(seed)
Matthews Correlation Coefficient
McQFold:
0.314
TurboFold(seed):
0.076
Sensitivity
McQFold:
0.205
TurboFold(seed):
0.007
Positive Predictive Value
McQFold:
0.480
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
TurboFold(seed):
0.076
Sensitivity
TurboFold(20):
0.108
TurboFold(seed):
0.007
Positive Predictive Value
TurboFold(20):
0.837
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
TurboFold(seed):
0.076
Sensitivity
PPfold(20):
0.101
TurboFold(seed):
0.007
Positive Predictive Value
PPfold(20):
0.879
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs TurboFold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
TurboFold(seed):
0.076
Sensitivity
RNAshapes:
0.180
TurboFold(seed):
0.007
Positive Predictive Value
RNAshapes:
0.484
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
TurboFold(seed):
0.076
Sensitivity
Carnac(20):
0.093
TurboFold(seed):
0.007
Positive Predictive Value
Carnac(20):
0.892
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs TurboFold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
TurboFold(seed):
0.076
Sensitivity
RNASLOpt:
0.158
TurboFold(seed):
0.007
Positive Predictive Value
RNASLOpt:
0.502
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
TurboFold(seed):
0.076
Sensitivity
RSpredict(20):
0.124
TurboFold(seed):
0.007
Positive Predictive Value
RSpredict(20):
0.590
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
TurboFold(seed):
0.076
Sensitivity
Multilign(20):
0.083
TurboFold(seed):
0.007
Positive Predictive Value
Multilign(20):
0.762
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs TurboFold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
TurboFold(seed):
0.076
Sensitivity
RNAwolf:
0.191
TurboFold(seed):
0.007
Positive Predictive Value
RNAwolf:
0.330
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
TurboFold(seed):
0.076
Sensitivity
Murlet(20):
0.067
TurboFold(seed):
0.007
Positive Predictive Value
Murlet(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
TurboFold(seed):
0.076
Sensitivity
Vsfold4:
0.125
TurboFold(seed):
0.007
Positive Predictive Value
Vsfold4:
0.378
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
TurboFold(seed):
0.076
Sensitivity
Vsfold5:
0.125
TurboFold(seed):
0.007
Positive Predictive Value
Vsfold5:
0.359
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
TurboFold(seed):
0.076
Sensitivity
RSpredict(seed):
0.075
TurboFold(seed):
0.007
Positive Predictive Value
RSpredict(seed):
0.597
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
TurboFold(seed):
0.076
Sensitivity
CMfinder(20):
0.055
TurboFold(seed):
0.007
Positive Predictive Value
CMfinder(20):
0.794
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs TurboFold(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
TurboFold(seed):
0.076
Sensitivity
HotKnots:
0.051
TurboFold(seed):
0.007
Positive Predictive Value
HotKnots:
0.624
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
TurboFold(seed):
0.076
Sensitivity
RNASampler(20):
0.035
TurboFold(seed):
0.007
Positive Predictive Value
RNASampler(20):
0.877
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs TurboFold(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
TurboFold(seed):
0.076
Sensitivity
Cylofold:
0.040
TurboFold(seed):
0.007
Positive Predictive Value
Cylofold:
0.606
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs TurboFold(seed)
Matthews Correlation Coefficient
Pknots:
0.143
TurboFold(seed):
0.076
Sensitivity
Pknots:
0.033
TurboFold(seed):
0.007
Positive Predictive Value
Pknots:
0.624
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
TurboFold(seed):
0.076
Sensitivity
Mastr(20):
0.023
TurboFold(seed):
0.007
Positive Predictive Value
Mastr(20):
0.790
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs TurboFold(seed)
Matthews Correlation Coefficient
Alterna:
0.129
TurboFold(seed):
0.076
Sensitivity
Alterna:
0.030
TurboFold(seed):
0.007
Positive Predictive Value
Alterna:
0.566
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs TurboFold(seed)
Matthews Correlation Coefficient
MCFold:
0.125
TurboFold(seed):
0.076
Sensitivity
MCFold:
0.036
TurboFold(seed):
0.007
Positive Predictive Value
MCFold:
0.435
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs TurboFold(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
TurboFold(seed):
0.076
Sensitivity
RDfolder:
0.025
TurboFold(seed):
0.007
Positive Predictive Value
RDfolder:
0.592
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
TurboFold(seed):
0.076
Sensitivity
Murlet(seed):
0.008
TurboFold(seed):
0.007
Positive Predictive Value
Murlet(seed):
0.811
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 5.5212650266e-05
|
1987
CMfinder(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
TurboFold(seed):
0.076
Sensitivity
CMfinder(seed):
0.008
TurboFold(seed):
0.007
Positive Predictive Value
CMfinder(seed):
0.757
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 1.83595875043e-07
|
|
+
TurboFold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
PPfold(seed):
0.069
Sensitivity
TurboFold(seed):
0.007
PPfold(seed):
0.006
Positive Predictive Value
TurboFold(seed):
0.822
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
Carnac(seed):
0.066
Sensitivity
TurboFold(seed):
0.007
Carnac(seed):
0.005
Positive Predictive Value
TurboFold(seed):
0.822
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
RNASampler(seed):
0.063
Sensitivity
TurboFold(seed):
0.007
RNASampler(seed):
0.005
Positive Predictive Value
TurboFold(seed):
0.822
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(seed):
0.076
NanoFolder:
0.052
Sensitivity
TurboFold(seed):
0.007
NanoFolder:
0.016
Positive Predictive Value
TurboFold(seed):
0.822
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
Mastr(seed):
0.051
Sensitivity
TurboFold(seed):
0.007
Mastr(seed):
0.003
Positive Predictive Value
TurboFold(seed):
0.822
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
Multilign(seed):
0.043
Sensitivity
TurboFold(seed):
0.007
Multilign(seed):
0.003
Positive Predictive Value
TurboFold(seed):
0.822
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PPfold(seed) |
1987
ContextFold vs PPfold(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
PPfold(seed):
0.069
Sensitivity
ContextFold:
0.644
PPfold(seed):
0.006
Positive Predictive Value
ContextFold:
0.718
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs PPfold(seed)
Matthews Correlation Coefficient
IPknot:
0.548
PPfold(seed):
0.069
Sensitivity
IPknot:
0.475
PPfold(seed):
0.006
Positive Predictive Value
IPknot:
0.632
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
PPfold(seed):
0.069
Sensitivity
PETfold_pre2.0(seed):
0.343
PPfold(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs PPfold(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
PPfold(seed):
0.069
Sensitivity
Contrafold:
0.481
PPfold(seed):
0.006
Positive Predictive Value
Contrafold:
0.537
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs PPfold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
PPfold(seed):
0.069
Sensitivity
CentroidFold:
0.369
PPfold(seed):
0.006
Positive Predictive Value
CentroidFold:
0.600
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs PPfold(seed)
Matthews Correlation Coefficient
Sfold:
0.469
PPfold(seed):
0.069
Sensitivity
Sfold:
0.412
PPfold(seed):
0.006
Positive Predictive Value
Sfold:
0.535
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs PPfold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
PPfold(seed):
0.069
Sensitivity
MaxExpect:
0.443
PPfold(seed):
0.006
Positive Predictive Value
MaxExpect:
0.488
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs PPfold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
PPfold(seed):
0.069
Sensitivity
ProbKnot:
0.446
PPfold(seed):
0.006
Positive Predictive Value
ProbKnot:
0.475
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs PPfold(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
PPfold(seed):
0.069
Sensitivity
UNAFold:
0.445
PPfold(seed):
0.006
Positive Predictive Value
UNAFold:
0.446
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
PPfold(seed):
0.069
Sensitivity
CentroidAlifold(seed):
0.213
PPfold(seed):
0.006
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs PPfold(seed)
Matthews Correlation Coefficient
Fold:
0.437
PPfold(seed):
0.069
Sensitivity
Fold:
0.432
PPfold(seed):
0.006
Positive Predictive Value
Fold:
0.444
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs PPfold(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
PPfold(seed):
0.069
Sensitivity
RNAfold:
0.422
PPfold(seed):
0.006
Positive Predictive Value
RNAfold:
0.437
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs PPfold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
PPfold(seed):
0.069
Sensitivity
CentroidHomfold‑LAST:
0.216
PPfold(seed):
0.006
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
PPfold(seed):
0.069
Sensitivity
PETfold_pre2.0(20):
0.211
PPfold(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
PPfold(seed):
0.069
Sensitivity
CentroidAlifold(20):
0.190
PPfold(seed):
0.006
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs PPfold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
PPfold(seed):
0.069
Sensitivity
PknotsRG:
0.320
PPfold(seed):
0.006
Positive Predictive Value
PknotsRG:
0.465
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
PPfold(seed):
0.069
Sensitivity
MXScarna(seed):
0.196
PPfold(seed):
0.006
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
PPfold(seed):
0.069
Sensitivity
RNAalifold(20):
0.185
PPfold(seed):
0.006
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
PPfold(seed):
0.069
Sensitivity
RNAalifold(seed):
0.169
PPfold(seed):
0.006
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs PPfold(seed)
Matthews Correlation Coefficient
Afold:
0.358
PPfold(seed):
0.069
Sensitivity
Afold:
0.295
PPfold(seed):
0.006
Positive Predictive Value
Afold:
0.434
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
PPfold(seed):
0.069
Sensitivity
MXScarna(20):
0.181
PPfold(seed):
0.006
Positive Predictive Value
MXScarna(20):
0.710
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs PPfold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
PPfold(seed):
0.069
Sensitivity
CRWrnafold:
0.275
PPfold(seed):
0.006
Positive Predictive Value
CRWrnafold:
0.435
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs PPfold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
PPfold(seed):
0.069
Sensitivity
RNAsubopt:
0.217
PPfold(seed):
0.006
Positive Predictive Value
RNAsubopt:
0.521
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs PPfold(seed)
Matthews Correlation Coefficient
McQFold:
0.314
PPfold(seed):
0.069
Sensitivity
McQFold:
0.205
PPfold(seed):
0.006
Positive Predictive Value
McQFold:
0.480
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
PPfold(seed):
0.069
Sensitivity
TurboFold(20):
0.108
PPfold(seed):
0.006
Positive Predictive Value
TurboFold(20):
0.837
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs PPfold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
PPfold(seed):
0.069
Sensitivity
PPfold(20):
0.101
PPfold(seed):
0.006
Positive Predictive Value
PPfold(20):
0.879
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs PPfold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
PPfold(seed):
0.069
Sensitivity
RNAshapes:
0.180
PPfold(seed):
0.006
Positive Predictive Value
RNAshapes:
0.484
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs PPfold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
PPfold(seed):
0.069
Sensitivity
Carnac(20):
0.093
PPfold(seed):
0.006
Positive Predictive Value
Carnac(20):
0.892
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs PPfold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
PPfold(seed):
0.069
Sensitivity
RNASLOpt:
0.158
PPfold(seed):
0.006
Positive Predictive Value
RNASLOpt:
0.502
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
PPfold(seed):
0.069
Sensitivity
RSpredict(20):
0.124
PPfold(seed):
0.006
Positive Predictive Value
RSpredict(20):
0.590
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs PPfold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
PPfold(seed):
0.069
Sensitivity
Multilign(20):
0.083
PPfold(seed):
0.006
Positive Predictive Value
Multilign(20):
0.762
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs PPfold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
PPfold(seed):
0.069
Sensitivity
RNAwolf:
0.191
PPfold(seed):
0.006
Positive Predictive Value
RNAwolf:
0.330
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
PPfold(seed):
0.069
Sensitivity
Murlet(20):
0.067
PPfold(seed):
0.006
Positive Predictive Value
Murlet(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
PPfold(seed):
0.069
Sensitivity
Vsfold4:
0.125
PPfold(seed):
0.006
Positive Predictive Value
Vsfold4:
0.378
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
PPfold(seed):
0.069
Sensitivity
Vsfold5:
0.125
PPfold(seed):
0.006
Positive Predictive Value
Vsfold5:
0.359
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
PPfold(seed):
0.069
Sensitivity
RSpredict(seed):
0.075
PPfold(seed):
0.006
Positive Predictive Value
RSpredict(seed):
0.597
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
PPfold(seed):
0.069
Sensitivity
CMfinder(20):
0.055
PPfold(seed):
0.006
Positive Predictive Value
CMfinder(20):
0.794
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs PPfold(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
PPfold(seed):
0.069
Sensitivity
HotKnots:
0.051
PPfold(seed):
0.006
Positive Predictive Value
HotKnots:
0.624
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
PPfold(seed):
0.069
Sensitivity
RNASampler(20):
0.035
PPfold(seed):
0.006
Positive Predictive Value
RNASampler(20):
0.877
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs PPfold(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
PPfold(seed):
0.069
Sensitivity
Cylofold:
0.040
PPfold(seed):
0.006
Positive Predictive Value
Cylofold:
0.606
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs PPfold(seed)
Matthews Correlation Coefficient
Pknots:
0.143
PPfold(seed):
0.069
Sensitivity
Pknots:
0.033
PPfold(seed):
0.006
Positive Predictive Value
Pknots:
0.624
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs PPfold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
PPfold(seed):
0.069
Sensitivity
Mastr(20):
0.023
PPfold(seed):
0.006
Positive Predictive Value
Mastr(20):
0.790
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs PPfold(seed)
Matthews Correlation Coefficient
Alterna:
0.129
PPfold(seed):
0.069
Sensitivity
Alterna:
0.030
PPfold(seed):
0.006
Positive Predictive Value
Alterna:
0.566
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs PPfold(seed)
Matthews Correlation Coefficient
MCFold:
0.125
PPfold(seed):
0.069
Sensitivity
MCFold:
0.036
PPfold(seed):
0.006
Positive Predictive Value
MCFold:
0.435
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs PPfold(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
PPfold(seed):
0.069
Sensitivity
RDfolder:
0.025
PPfold(seed):
0.006
Positive Predictive Value
RDfolder:
0.592
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
PPfold(seed):
0.069
Sensitivity
Murlet(seed):
0.008
PPfold(seed):
0.006
Positive Predictive Value
Murlet(seed):
0.811
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
PPfold(seed):
0.069
Sensitivity
CMfinder(seed):
0.008
PPfold(seed):
0.006
Positive Predictive Value
CMfinder(seed):
0.757
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
PPfold(seed):
0.069
Sensitivity
TurboFold(seed):
0.007
PPfold(seed):
0.006
Positive Predictive Value
TurboFold(seed):
0.822
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PPfold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
Carnac(seed):
0.066
Sensitivity
PPfold(seed):
0.006
Carnac(seed):
0.005
Positive Predictive Value
PPfold(seed):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
PPfold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
RNASampler(seed):
0.063
Sensitivity
PPfold(seed):
0.006
RNASampler(seed):
0.005
Positive Predictive Value
PPfold(seed):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(seed) vs NanoFolder
Matthews Correlation Coefficient
PPfold(seed):
0.069
NanoFolder:
0.052
Sensitivity
PPfold(seed):
0.006
NanoFolder:
0.016
Positive Predictive Value
PPfold(seed):
0.794
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
Mastr(seed):
0.051
Sensitivity
PPfold(seed):
0.006
Mastr(seed):
0.003
Positive Predictive Value
PPfold(seed):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
Multilign(seed):
0.043
Sensitivity
PPfold(seed):
0.006
Multilign(seed):
0.003
Positive Predictive Value
PPfold(seed):
0.794
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Carnac(seed) |
1987
ContextFold vs Carnac(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Carnac(seed):
0.066
Sensitivity
ContextFold:
0.644
Carnac(seed):
0.005
Positive Predictive Value
ContextFold:
0.718
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Carnac(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Carnac(seed):
0.066
Sensitivity
IPknot:
0.475
Carnac(seed):
0.005
Positive Predictive Value
IPknot:
0.632
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Carnac(seed):
0.066
Sensitivity
PETfold_pre2.0(seed):
0.343
Carnac(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Carnac(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Carnac(seed):
0.066
Sensitivity
Contrafold:
0.481
Carnac(seed):
0.005
Positive Predictive Value
Contrafold:
0.537
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Carnac(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Carnac(seed):
0.066
Sensitivity
CentroidFold:
0.369
Carnac(seed):
0.005
Positive Predictive Value
CentroidFold:
0.600
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Carnac(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Carnac(seed):
0.066
Sensitivity
Sfold:
0.412
Carnac(seed):
0.005
Positive Predictive Value
Sfold:
0.535
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Carnac(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Carnac(seed):
0.066
Sensitivity
MaxExpect:
0.443
Carnac(seed):
0.005
Positive Predictive Value
MaxExpect:
0.488
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Carnac(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Carnac(seed):
0.066
Sensitivity
ProbKnot:
0.446
Carnac(seed):
0.005
Positive Predictive Value
ProbKnot:
0.475
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Carnac(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Carnac(seed):
0.066
Sensitivity
UNAFold:
0.445
Carnac(seed):
0.005
Positive Predictive Value
UNAFold:
0.446
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Carnac(seed):
0.066
Sensitivity
CentroidAlifold(seed):
0.213
Carnac(seed):
0.005
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Carnac(seed)
Matthews Correlation Coefficient
Fold:
0.437
Carnac(seed):
0.066
Sensitivity
Fold:
0.432
Carnac(seed):
0.005
Positive Predictive Value
Fold:
0.444
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Carnac(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Carnac(seed):
0.066
Sensitivity
RNAfold:
0.422
Carnac(seed):
0.005
Positive Predictive Value
RNAfold:
0.437
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Carnac(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Carnac(seed):
0.066
Sensitivity
CentroidHomfold‑LAST:
0.216
Carnac(seed):
0.005
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Carnac(seed):
0.066
Sensitivity
PETfold_pre2.0(20):
0.211
Carnac(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Carnac(seed):
0.066
Sensitivity
CentroidAlifold(20):
0.190
Carnac(seed):
0.005
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Carnac(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Carnac(seed):
0.066
Sensitivity
PknotsRG:
0.320
Carnac(seed):
0.005
Positive Predictive Value
PknotsRG:
0.465
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Carnac(seed):
0.066
Sensitivity
MXScarna(seed):
0.196
Carnac(seed):
0.005
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Carnac(seed):
0.066
Sensitivity
RNAalifold(20):
0.185
Carnac(seed):
0.005
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Carnac(seed):
0.066
Sensitivity
RNAalifold(seed):
0.169
Carnac(seed):
0.005
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Carnac(seed)
Matthews Correlation Coefficient
Afold:
0.358
Carnac(seed):
0.066
Sensitivity
Afold:
0.295
Carnac(seed):
0.005
Positive Predictive Value
Afold:
0.434
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Carnac(seed):
0.066
Sensitivity
MXScarna(20):
0.181
Carnac(seed):
0.005
Positive Predictive Value
MXScarna(20):
0.710
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Carnac(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Carnac(seed):
0.066
Sensitivity
CRWrnafold:
0.275
Carnac(seed):
0.005
Positive Predictive Value
CRWrnafold:
0.435
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Carnac(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Carnac(seed):
0.066
Sensitivity
RNAsubopt:
0.217
Carnac(seed):
0.005
Positive Predictive Value
RNAsubopt:
0.521
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Carnac(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Carnac(seed):
0.066
Sensitivity
McQFold:
0.205
Carnac(seed):
0.005
Positive Predictive Value
McQFold:
0.480
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Carnac(seed):
0.066
Sensitivity
TurboFold(20):
0.108
Carnac(seed):
0.005
Positive Predictive Value
TurboFold(20):
0.837
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Carnac(seed):
0.066
Sensitivity
PPfold(20):
0.101
Carnac(seed):
0.005
Positive Predictive Value
PPfold(20):
0.879
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Carnac(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Carnac(seed):
0.066
Sensitivity
RNAshapes:
0.180
Carnac(seed):
0.005
Positive Predictive Value
RNAshapes:
0.484
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Carnac(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Carnac(seed):
0.066
Sensitivity
Carnac(20):
0.093
Carnac(seed):
0.005
Positive Predictive Value
Carnac(20):
0.892
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Carnac(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Carnac(seed):
0.066
Sensitivity
RNASLOpt:
0.158
Carnac(seed):
0.005
Positive Predictive Value
RNASLOpt:
0.502
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Carnac(seed):
0.066
Sensitivity
RSpredict(20):
0.124
Carnac(seed):
0.005
Positive Predictive Value
RSpredict(20):
0.590
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Carnac(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Carnac(seed):
0.066
Sensitivity
Multilign(20):
0.083
Carnac(seed):
0.005
Positive Predictive Value
Multilign(20):
0.762
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Carnac(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Carnac(seed):
0.066
Sensitivity
RNAwolf:
0.191
Carnac(seed):
0.005
Positive Predictive Value
RNAwolf:
0.330
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Carnac(seed):
0.066
Sensitivity
Murlet(20):
0.067
Carnac(seed):
0.005
Positive Predictive Value
Murlet(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Carnac(seed):
0.066
Sensitivity
Vsfold4:
0.125
Carnac(seed):
0.005
Positive Predictive Value
Vsfold4:
0.378
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Carnac(seed):
0.066
Sensitivity
Vsfold5:
0.125
Carnac(seed):
0.005
Positive Predictive Value
Vsfold5:
0.359
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Carnac(seed):
0.066
Sensitivity
RSpredict(seed):
0.075
Carnac(seed):
0.005
Positive Predictive Value
RSpredict(seed):
0.597
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Carnac(seed):
0.066
Sensitivity
CMfinder(20):
0.055
Carnac(seed):
0.005
Positive Predictive Value
CMfinder(20):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Carnac(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Carnac(seed):
0.066
Sensitivity
HotKnots:
0.051
Carnac(seed):
0.005
Positive Predictive Value
HotKnots:
0.624
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Carnac(seed):
0.066
Sensitivity
RNASampler(20):
0.035
Carnac(seed):
0.005
Positive Predictive Value
RNASampler(20):
0.877
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Carnac(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Carnac(seed):
0.066
Sensitivity
Cylofold:
0.040
Carnac(seed):
0.005
Positive Predictive Value
Cylofold:
0.606
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs Carnac(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Carnac(seed):
0.066
Sensitivity
Pknots:
0.033
Carnac(seed):
0.005
Positive Predictive Value
Pknots:
0.624
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs Carnac(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Carnac(seed):
0.066
Sensitivity
Mastr(20):
0.023
Carnac(seed):
0.005
Positive Predictive Value
Mastr(20):
0.790
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs Carnac(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Carnac(seed):
0.066
Sensitivity
Alterna:
0.030
Carnac(seed):
0.005
Positive Predictive Value
Alterna:
0.566
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs Carnac(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Carnac(seed):
0.066
Sensitivity
MCFold:
0.036
Carnac(seed):
0.005
Positive Predictive Value
MCFold:
0.435
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs Carnac(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Carnac(seed):
0.066
Sensitivity
RDfolder:
0.025
Carnac(seed):
0.005
Positive Predictive Value
RDfolder:
0.592
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
Carnac(seed):
0.066
Sensitivity
Murlet(seed):
0.008
Carnac(seed):
0.005
Positive Predictive Value
Murlet(seed):
0.811
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Carnac(seed):
0.066
Sensitivity
CMfinder(seed):
0.008
Carnac(seed):
0.005
Positive Predictive Value
CMfinder(seed):
0.757
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
Carnac(seed):
0.066
Sensitivity
TurboFold(seed):
0.007
Carnac(seed):
0.005
Positive Predictive Value
TurboFold(seed):
0.822
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
Carnac(seed):
0.066
Sensitivity
PPfold(seed):
0.006
Carnac(seed):
0.005
Positive Predictive Value
PPfold(seed):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
|
+
Carnac(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.066
RNASampler(seed):
0.063
Sensitivity
Carnac(seed):
0.005
RNASampler(seed):
0.005
Positive Predictive Value
Carnac(seed):
0.867
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.000857786362802
|
+
Carnac(seed) vs NanoFolder
Matthews Correlation Coefficient
Carnac(seed):
0.066
NanoFolder:
0.052
Sensitivity
Carnac(seed):
0.005
NanoFolder:
0.016
Positive Predictive Value
Carnac(seed):
0.867
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.066
Mastr(seed):
0.051
Sensitivity
Carnac(seed):
0.005
Mastr(seed):
0.003
Positive Predictive Value
Carnac(seed):
0.867
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.066
Multilign(seed):
0.043
Sensitivity
Carnac(seed):
0.005
Multilign(seed):
0.003
Positive Predictive Value
Carnac(seed):
0.867
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNASampler(seed) |
1987
ContextFold vs RNASampler(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
RNASampler(seed):
0.063
Sensitivity
ContextFold:
0.644
RNASampler(seed):
0.005
Positive Predictive Value
ContextFold:
0.718
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs RNASampler(seed)
Matthews Correlation Coefficient
IPknot:
0.548
RNASampler(seed):
0.063
Sensitivity
IPknot:
0.475
RNASampler(seed):
0.005
Positive Predictive Value
IPknot:
0.632
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
RNASampler(seed):
0.063
Sensitivity
PETfold_pre2.0(seed):
0.343
RNASampler(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs RNASampler(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
RNASampler(seed):
0.063
Sensitivity
Contrafold:
0.481
RNASampler(seed):
0.005
Positive Predictive Value
Contrafold:
0.537
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
RNASampler(seed):
0.063
Sensitivity
CentroidFold:
0.369
RNASampler(seed):
0.005
Positive Predictive Value
CentroidFold:
0.600
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs RNASampler(seed)
Matthews Correlation Coefficient
Sfold:
0.469
RNASampler(seed):
0.063
Sensitivity
Sfold:
0.412
RNASampler(seed):
0.005
Positive Predictive Value
Sfold:
0.535
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs RNASampler(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
RNASampler(seed):
0.063
Sensitivity
MaxExpect:
0.443
RNASampler(seed):
0.005
Positive Predictive Value
MaxExpect:
0.488
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs RNASampler(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
RNASampler(seed):
0.063
Sensitivity
ProbKnot:
0.446
RNASampler(seed):
0.005
Positive Predictive Value
ProbKnot:
0.475
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs RNASampler(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
RNASampler(seed):
0.063
Sensitivity
UNAFold:
0.445
RNASampler(seed):
0.005
Positive Predictive Value
UNAFold:
0.446
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
RNASampler(seed):
0.063
Sensitivity
CentroidAlifold(seed):
0.213
RNASampler(seed):
0.005
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs RNASampler(seed)
Matthews Correlation Coefficient
Fold:
0.437
RNASampler(seed):
0.063
Sensitivity
Fold:
0.432
RNASampler(seed):
0.005
Positive Predictive Value
Fold:
0.444
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs RNASampler(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
RNASampler(seed):
0.063
Sensitivity
RNAfold:
0.422
RNASampler(seed):
0.005
Positive Predictive Value
RNAfold:
0.437
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
RNASampler(seed):
0.063
Sensitivity
CentroidHomfold‑LAST:
0.216
RNASampler(seed):
0.005
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
RNASampler(seed):
0.063
Sensitivity
PETfold_pre2.0(20):
0.211
RNASampler(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
RNASampler(seed):
0.063
Sensitivity
CentroidAlifold(20):
0.190
RNASampler(seed):
0.005
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs RNASampler(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
RNASampler(seed):
0.063
Sensitivity
PknotsRG:
0.320
RNASampler(seed):
0.005
Positive Predictive Value
PknotsRG:
0.465
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
RNASampler(seed):
0.063
Sensitivity
MXScarna(seed):
0.196
RNASampler(seed):
0.005
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
RNASampler(seed):
0.063
Sensitivity
RNAalifold(20):
0.185
RNASampler(seed):
0.005
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
RNASampler(seed):
0.063
Sensitivity
RNAalifold(seed):
0.169
RNASampler(seed):
0.005
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs RNASampler(seed)
Matthews Correlation Coefficient
Afold:
0.358
RNASampler(seed):
0.063
Sensitivity
Afold:
0.295
RNASampler(seed):
0.005
Positive Predictive Value
Afold:
0.434
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
RNASampler(seed):
0.063
Sensitivity
MXScarna(20):
0.181
RNASampler(seed):
0.005
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs RNASampler(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
RNASampler(seed):
0.063
Sensitivity
CRWrnafold:
0.275
RNASampler(seed):
0.005
Positive Predictive Value
CRWrnafold:
0.435
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs RNASampler(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
RNASampler(seed):
0.063
Sensitivity
RNAsubopt:
0.217
RNASampler(seed):
0.005
Positive Predictive Value
RNAsubopt:
0.521
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs RNASampler(seed)
Matthews Correlation Coefficient
McQFold:
0.314
RNASampler(seed):
0.063
Sensitivity
McQFold:
0.205
RNASampler(seed):
0.005
Positive Predictive Value
McQFold:
0.480
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
RNASampler(seed):
0.063
Sensitivity
TurboFold(20):
0.108
RNASampler(seed):
0.005
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
RNASampler(seed):
0.063
Sensitivity
PPfold(20):
0.101
RNASampler(seed):
0.005
Positive Predictive Value
PPfold(20):
0.879
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs RNASampler(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
RNASampler(seed):
0.063
Sensitivity
RNAshapes:
0.180
RNASampler(seed):
0.005
Positive Predictive Value
RNAshapes:
0.484
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
RNASampler(seed):
0.063
Sensitivity
Carnac(20):
0.093
RNASampler(seed):
0.005
Positive Predictive Value
Carnac(20):
0.892
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs RNASampler(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
RNASampler(seed):
0.063
Sensitivity
RNASLOpt:
0.158
RNASampler(seed):
0.005
Positive Predictive Value
RNASLOpt:
0.502
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
RNASampler(seed):
0.063
Sensitivity
RSpredict(20):
0.124
RNASampler(seed):
0.005
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
RNASampler(seed):
0.063
Sensitivity
Multilign(20):
0.083
RNASampler(seed):
0.005
Positive Predictive Value
Multilign(20):
0.762
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs RNASampler(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
RNASampler(seed):
0.063
Sensitivity
RNAwolf:
0.191
RNASampler(seed):
0.005
Positive Predictive Value
RNAwolf:
0.330
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
RNASampler(seed):
0.063
Sensitivity
Murlet(20):
0.067
RNASampler(seed):
0.005
Positive Predictive Value
Murlet(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
RNASampler(seed):
0.063
Sensitivity
Vsfold4:
0.125
RNASampler(seed):
0.005
Positive Predictive Value
Vsfold4:
0.378
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
RNASampler(seed):
0.063
Sensitivity
Vsfold5:
0.125
RNASampler(seed):
0.005
Positive Predictive Value
Vsfold5:
0.359
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
RNASampler(seed):
0.063
Sensitivity
RSpredict(seed):
0.075
RNASampler(seed):
0.005
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
RNASampler(seed):
0.063
Sensitivity
CMfinder(20):
0.055
RNASampler(seed):
0.005
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs RNASampler(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
RNASampler(seed):
0.063
Sensitivity
HotKnots:
0.051
RNASampler(seed):
0.005
Positive Predictive Value
HotKnots:
0.624
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
RNASampler(seed):
0.063
Sensitivity
RNASampler(20):
0.035
RNASampler(seed):
0.005
Positive Predictive Value
RNASampler(20):
0.877
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs RNASampler(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
RNASampler(seed):
0.063
Sensitivity
Cylofold:
0.040
RNASampler(seed):
0.005
Positive Predictive Value
Cylofold:
0.606
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs RNASampler(seed)
Matthews Correlation Coefficient
Pknots:
0.143
RNASampler(seed):
0.063
Sensitivity
Pknots:
0.033
RNASampler(seed):
0.005
Positive Predictive Value
Pknots:
0.624
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
RNASampler(seed):
0.063
Sensitivity
Mastr(20):
0.023
RNASampler(seed):
0.005
Positive Predictive Value
Mastr(20):
0.790
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs RNASampler(seed)
Matthews Correlation Coefficient
Alterna:
0.129
RNASampler(seed):
0.063
Sensitivity
Alterna:
0.030
RNASampler(seed):
0.005
Positive Predictive Value
Alterna:
0.566
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs RNASampler(seed)
Matthews Correlation Coefficient
MCFold:
0.125
RNASampler(seed):
0.063
Sensitivity
MCFold:
0.036
RNASampler(seed):
0.005
Positive Predictive Value
MCFold:
0.435
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs RNASampler(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
RNASampler(seed):
0.063
Sensitivity
RDfolder:
0.025
RNASampler(seed):
0.005
Positive Predictive Value
RDfolder:
0.592
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
RNASampler(seed):
0.063
Sensitivity
Murlet(seed):
0.008
RNASampler(seed):
0.005
Positive Predictive Value
Murlet(seed):
0.811
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
RNASampler(seed):
0.063
Sensitivity
CMfinder(seed):
0.008
RNASampler(seed):
0.005
Positive Predictive Value
CMfinder(seed):
0.757
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
RNASampler(seed):
0.063
Sensitivity
TurboFold(seed):
0.007
RNASampler(seed):
0.005
Positive Predictive Value
TurboFold(seed):
0.822
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
RNASampler(seed):
0.063
Sensitivity
PPfold(seed):
0.006
RNASampler(seed):
0.005
Positive Predictive Value
PPfold(seed):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.066
RNASampler(seed):
0.063
Sensitivity
Carnac(seed):
0.005
RNASampler(seed):
0.005
Positive Predictive Value
Carnac(seed):
0.867
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.000857786362802
|
|
+
RNASampler(seed) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(seed):
0.063
NanoFolder:
0.052
Sensitivity
RNASampler(seed):
0.005
NanoFolder:
0.016
Positive Predictive Value
RNASampler(seed):
0.847
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.063
Mastr(seed):
0.051
Sensitivity
RNASampler(seed):
0.005
Mastr(seed):
0.003
Positive Predictive Value
RNASampler(seed):
0.847
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.063
Multilign(seed):
0.043
Sensitivity
RNASampler(seed):
0.005
Multilign(seed):
0.003
Positive Predictive Value
RNASampler(seed):
0.847
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| NanoFolder |
1987
ContextFold vs NanoFolder
Matthews Correlation Coefficient
ContextFold:
0.680
NanoFolder:
0.052
Sensitivity
ContextFold:
0.644
NanoFolder:
0.016
Positive Predictive Value
ContextFold:
0.718
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs NanoFolder
Matthews Correlation Coefficient
IPknot:
0.548
NanoFolder:
0.052
Sensitivity
IPknot:
0.475
NanoFolder:
0.016
Positive Predictive Value
IPknot:
0.632
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
NanoFolder:
0.052
Sensitivity
PETfold_pre2.0(seed):
0.343
NanoFolder:
0.016
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs NanoFolder
Matthews Correlation Coefficient
Contrafold:
0.508
NanoFolder:
0.052
Sensitivity
Contrafold:
0.481
NanoFolder:
0.016
Positive Predictive Value
Contrafold:
0.537
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs NanoFolder
Matthews Correlation Coefficient
CentroidFold:
0.470
NanoFolder:
0.052
Sensitivity
CentroidFold:
0.369
NanoFolder:
0.016
Positive Predictive Value
CentroidFold:
0.600
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs NanoFolder
Matthews Correlation Coefficient
Sfold:
0.469
NanoFolder:
0.052
Sensitivity
Sfold:
0.412
NanoFolder:
0.016
Positive Predictive Value
Sfold:
0.535
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs NanoFolder
Matthews Correlation Coefficient
MaxExpect:
0.465
NanoFolder:
0.052
Sensitivity
MaxExpect:
0.443
NanoFolder:
0.016
Positive Predictive Value
MaxExpect:
0.488
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs NanoFolder
Matthews Correlation Coefficient
ProbKnot:
0.460
NanoFolder:
0.052
Sensitivity
ProbKnot:
0.446
NanoFolder:
0.016
Positive Predictive Value
ProbKnot:
0.475
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs NanoFolder
Matthews Correlation Coefficient
UNAFold:
0.445
NanoFolder:
0.052
Sensitivity
UNAFold:
0.445
NanoFolder:
0.016
Positive Predictive Value
UNAFold:
0.446
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
NanoFolder:
0.052
Sensitivity
CentroidAlifold(seed):
0.213
NanoFolder:
0.016
Positive Predictive Value
CentroidAlifold(seed):
0.901
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs NanoFolder
Matthews Correlation Coefficient
Fold:
0.437
NanoFolder:
0.052
Sensitivity
Fold:
0.432
NanoFolder:
0.016
Positive Predictive Value
Fold:
0.444
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs NanoFolder
Matthews Correlation Coefficient
RNAfold:
0.429
NanoFolder:
0.052
Sensitivity
RNAfold:
0.422
NanoFolder:
0.016
Positive Predictive Value
RNAfold:
0.437
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs NanoFolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
NanoFolder:
0.052
Sensitivity
CentroidHomfold‑LAST:
0.216
NanoFolder:
0.016
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
NanoFolder:
0.052
Sensitivity
PETfold_pre2.0(20):
0.211
NanoFolder:
0.016
Positive Predictive Value
PETfold_pre2.0(20):
0.807
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
NanoFolder:
0.052
Sensitivity
CentroidAlifold(20):
0.190
NanoFolder:
0.016
Positive Predictive Value
CentroidAlifold(20):
0.891
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs NanoFolder
Matthews Correlation Coefficient
PknotsRG:
0.386
NanoFolder:
0.052
Sensitivity
PknotsRG:
0.320
NanoFolder:
0.016
Positive Predictive Value
PknotsRG:
0.465
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(seed):
0.385
NanoFolder:
0.052
Sensitivity
MXScarna(seed):
0.196
NanoFolder:
0.016
Positive Predictive Value
MXScarna(seed):
0.753
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(20):
0.381
NanoFolder:
0.052
Sensitivity
RNAalifold(20):
0.185
NanoFolder:
0.016
Positive Predictive Value
RNAalifold(20):
0.787
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
NanoFolder:
0.052
Sensitivity
RNAalifold(seed):
0.169
NanoFolder:
0.016
Positive Predictive Value
RNAalifold(seed):
0.835
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs NanoFolder
Matthews Correlation Coefficient
Afold:
0.358
NanoFolder:
0.052
Sensitivity
Afold:
0.295
NanoFolder:
0.016
Positive Predictive Value
Afold:
0.434
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(20):
0.359
NanoFolder:
0.052
Sensitivity
MXScarna(20):
0.181
NanoFolder:
0.016
Positive Predictive Value
MXScarna(20):
0.710
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs NanoFolder
Matthews Correlation Coefficient
CRWrnafold:
0.346
NanoFolder:
0.052
Sensitivity
CRWrnafold:
0.275
NanoFolder:
0.016
Positive Predictive Value
CRWrnafold:
0.435
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs NanoFolder
Matthews Correlation Coefficient
RNAsubopt:
0.336
NanoFolder:
0.052
Sensitivity
RNAsubopt:
0.217
NanoFolder:
0.016
Positive Predictive Value
RNAsubopt:
0.521
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs NanoFolder
Matthews Correlation Coefficient
McQFold:
0.314
NanoFolder:
0.052
Sensitivity
McQFold:
0.205
NanoFolder:
0.016
Positive Predictive Value
McQFold:
0.480
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(20):
0.301
NanoFolder:
0.052
Sensitivity
TurboFold(20):
0.108
NanoFolder:
0.016
Positive Predictive Value
TurboFold(20):
0.837
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs NanoFolder
Matthews Correlation Coefficient
PPfold(20):
0.298
NanoFolder:
0.052
Sensitivity
PPfold(20):
0.101
NanoFolder:
0.016
Positive Predictive Value
PPfold(20):
0.879
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs NanoFolder
Matthews Correlation Coefficient
RNAshapes:
0.295
NanoFolder:
0.052
Sensitivity
RNAshapes:
0.180
NanoFolder:
0.016
Positive Predictive Value
RNAshapes:
0.484
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs NanoFolder
Matthews Correlation Coefficient
Carnac(20):
0.288
NanoFolder:
0.052
Sensitivity
Carnac(20):
0.093
NanoFolder:
0.016
Positive Predictive Value
Carnac(20):
0.892
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs NanoFolder
Matthews Correlation Coefficient
RNASLOpt:
0.281
NanoFolder:
0.052
Sensitivity
RNASLOpt:
0.158
NanoFolder:
0.016
Positive Predictive Value
RNASLOpt:
0.502
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(20):
0.270
NanoFolder:
0.052
Sensitivity
RSpredict(20):
0.124
NanoFolder:
0.016
Positive Predictive Value
RSpredict(20):
0.590
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs NanoFolder
Matthews Correlation Coefficient
Multilign(20):
0.251
NanoFolder:
0.052
Sensitivity
Multilign(20):
0.083
NanoFolder:
0.016
Positive Predictive Value
Multilign(20):
0.762
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs NanoFolder
Matthews Correlation Coefficient
RNAwolf:
0.251
NanoFolder:
0.052
Sensitivity
RNAwolf:
0.191
NanoFolder:
0.016
Positive Predictive Value
RNAwolf:
0.330
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs NanoFolder
Matthews Correlation Coefficient
Murlet(20):
0.230
NanoFolder:
0.052
Sensitivity
Murlet(20):
0.067
NanoFolder:
0.016
Positive Predictive Value
Murlet(20):
0.787
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs NanoFolder
Matthews Correlation Coefficient
Vsfold4:
0.217
NanoFolder:
0.052
Sensitivity
Vsfold4:
0.125
NanoFolder:
0.016
Positive Predictive Value
Vsfold4:
0.378
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs NanoFolder
Matthews Correlation Coefficient
Vsfold5:
0.211
NanoFolder:
0.052
Sensitivity
Vsfold5:
0.125
NanoFolder:
0.016
Positive Predictive Value
Vsfold5:
0.359
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(seed):
0.212
NanoFolder:
0.052
Sensitivity
RSpredict(seed):
0.075
NanoFolder:
0.016
Positive Predictive Value
RSpredict(seed):
0.597
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(20):
0.210
NanoFolder:
0.052
Sensitivity
CMfinder(20):
0.055
NanoFolder:
0.016
Positive Predictive Value
CMfinder(20):
0.794
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs NanoFolder
Matthews Correlation Coefficient
HotKnots:
0.179
NanoFolder:
0.052
Sensitivity
HotKnots:
0.051
NanoFolder:
0.016
Positive Predictive Value
HotKnots:
0.624
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(20):
0.176
NanoFolder:
0.052
Sensitivity
RNASampler(20):
0.035
NanoFolder:
0.016
Positive Predictive Value
RNASampler(20):
0.877
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs NanoFolder
Matthews Correlation Coefficient
Cylofold:
0.155
NanoFolder:
0.052
Sensitivity
Cylofold:
0.040
NanoFolder:
0.016
Positive Predictive Value
Cylofold:
0.606
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs NanoFolder
Matthews Correlation Coefficient
Pknots:
0.143
NanoFolder:
0.052
Sensitivity
Pknots:
0.033
NanoFolder:
0.016
Positive Predictive Value
Pknots:
0.624
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs NanoFolder
Matthews Correlation Coefficient
Mastr(20):
0.134
NanoFolder:
0.052
Sensitivity
Mastr(20):
0.023
NanoFolder:
0.016
Positive Predictive Value
Mastr(20):
0.790
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs NanoFolder
Matthews Correlation Coefficient
Alterna:
0.129
NanoFolder:
0.052
Sensitivity
Alterna:
0.030
NanoFolder:
0.016
Positive Predictive Value
Alterna:
0.566
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs NanoFolder
Matthews Correlation Coefficient
MCFold:
0.125
NanoFolder:
0.052
Sensitivity
MCFold:
0.036
NanoFolder:
0.016
Positive Predictive Value
MCFold:
0.435
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs NanoFolder
Matthews Correlation Coefficient
RDfolder:
0.122
NanoFolder:
0.052
Sensitivity
RDfolder:
0.025
NanoFolder:
0.016
Positive Predictive Value
RDfolder:
0.592
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs NanoFolder
Matthews Correlation Coefficient
Murlet(seed):
0.079
NanoFolder:
0.052
Sensitivity
Murlet(seed):
0.008
NanoFolder:
0.016
Positive Predictive Value
Murlet(seed):
0.811
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(seed):
0.080
NanoFolder:
0.052
Sensitivity
CMfinder(seed):
0.008
NanoFolder:
0.016
Positive Predictive Value
CMfinder(seed):
0.757
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(seed) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(seed):
0.076
NanoFolder:
0.052
Sensitivity
TurboFold(seed):
0.007
NanoFolder:
0.016
Positive Predictive Value
TurboFold(seed):
0.822
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(seed) vs NanoFolder
Matthews Correlation Coefficient
PPfold(seed):
0.069
NanoFolder:
0.052
Sensitivity
PPfold(seed):
0.006
NanoFolder:
0.016
Positive Predictive Value
PPfold(seed):
0.794
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(seed) vs NanoFolder
Matthews Correlation Coefficient
Carnac(seed):
0.066
NanoFolder:
0.052
Sensitivity
Carnac(seed):
0.005
NanoFolder:
0.016
Positive Predictive Value
Carnac(seed):
0.867
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(seed) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(seed):
0.063
NanoFolder:
0.052
Sensitivity
RNASampler(seed):
0.005
NanoFolder:
0.016
Positive Predictive Value
RNASampler(seed):
0.847
NanoFolder:
0.168
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
NanoFolder vs Mastr(seed)
Matthews Correlation Coefficient
NanoFolder:
0.052
Mastr(seed):
0.051
Sensitivity
NanoFolder:
0.016
Mastr(seed):
0.003
Positive Predictive Value
NanoFolder:
0.168
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.0529232299595
|
+
NanoFolder vs Multilign(seed)
Matthews Correlation Coefficient
NanoFolder:
0.052
Multilign(seed):
0.043
Sensitivity
NanoFolder:
0.016
Multilign(seed):
0.003
Positive Predictive Value
NanoFolder:
0.168
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Mastr(seed) |
1987
ContextFold vs Mastr(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Mastr(seed):
0.051
Sensitivity
ContextFold:
0.644
Mastr(seed):
0.003
Positive Predictive Value
ContextFold:
0.718
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Mastr(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Mastr(seed):
0.051
Sensitivity
IPknot:
0.475
Mastr(seed):
0.003
Positive Predictive Value
IPknot:
0.632
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Mastr(seed):
0.051
Sensitivity
PETfold_pre2.0(seed):
0.343
Mastr(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Mastr(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Mastr(seed):
0.051
Sensitivity
Contrafold:
0.481
Mastr(seed):
0.003
Positive Predictive Value
Contrafold:
0.537
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Mastr(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Mastr(seed):
0.051
Sensitivity
CentroidFold:
0.369
Mastr(seed):
0.003
Positive Predictive Value
CentroidFold:
0.600
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Mastr(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Mastr(seed):
0.051
Sensitivity
Sfold:
0.412
Mastr(seed):
0.003
Positive Predictive Value
Sfold:
0.535
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Mastr(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Mastr(seed):
0.051
Sensitivity
MaxExpect:
0.443
Mastr(seed):
0.003
Positive Predictive Value
MaxExpect:
0.488
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Mastr(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Mastr(seed):
0.051
Sensitivity
ProbKnot:
0.446
Mastr(seed):
0.003
Positive Predictive Value
ProbKnot:
0.475
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Mastr(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Mastr(seed):
0.051
Sensitivity
UNAFold:
0.445
Mastr(seed):
0.003
Positive Predictive Value
UNAFold:
0.446
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Mastr(seed):
0.051
Sensitivity
CentroidAlifold(seed):
0.213
Mastr(seed):
0.003
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Mastr(seed)
Matthews Correlation Coefficient
Fold:
0.437
Mastr(seed):
0.051
Sensitivity
Fold:
0.432
Mastr(seed):
0.003
Positive Predictive Value
Fold:
0.444
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Mastr(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Mastr(seed):
0.051
Sensitivity
RNAfold:
0.422
Mastr(seed):
0.003
Positive Predictive Value
RNAfold:
0.437
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Mastr(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Mastr(seed):
0.051
Sensitivity
CentroidHomfold‑LAST:
0.216
Mastr(seed):
0.003
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Mastr(seed):
0.051
Sensitivity
PETfold_pre2.0(20):
0.211
Mastr(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Mastr(seed):
0.051
Sensitivity
CentroidAlifold(20):
0.190
Mastr(seed):
0.003
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Mastr(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Mastr(seed):
0.051
Sensitivity
PknotsRG:
0.320
Mastr(seed):
0.003
Positive Predictive Value
PknotsRG:
0.465
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Mastr(seed):
0.051
Sensitivity
MXScarna(seed):
0.196
Mastr(seed):
0.003
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Mastr(seed):
0.051
Sensitivity
RNAalifold(20):
0.185
Mastr(seed):
0.003
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Mastr(seed):
0.051
Sensitivity
RNAalifold(seed):
0.169
Mastr(seed):
0.003
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Mastr(seed)
Matthews Correlation Coefficient
Afold:
0.358
Mastr(seed):
0.051
Sensitivity
Afold:
0.295
Mastr(seed):
0.003
Positive Predictive Value
Afold:
0.434
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Mastr(seed):
0.051
Sensitivity
MXScarna(20):
0.181
Mastr(seed):
0.003
Positive Predictive Value
MXScarna(20):
0.710
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Mastr(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Mastr(seed):
0.051
Sensitivity
CRWrnafold:
0.275
Mastr(seed):
0.003
Positive Predictive Value
CRWrnafold:
0.435
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Mastr(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Mastr(seed):
0.051
Sensitivity
RNAsubopt:
0.217
Mastr(seed):
0.003
Positive Predictive Value
RNAsubopt:
0.521
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Mastr(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Mastr(seed):
0.051
Sensitivity
McQFold:
0.205
Mastr(seed):
0.003
Positive Predictive Value
McQFold:
0.480
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Mastr(seed):
0.051
Sensitivity
TurboFold(20):
0.108
Mastr(seed):
0.003
Positive Predictive Value
TurboFold(20):
0.837
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Mastr(seed):
0.051
Sensitivity
PPfold(20):
0.101
Mastr(seed):
0.003
Positive Predictive Value
PPfold(20):
0.879
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Mastr(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Mastr(seed):
0.051
Sensitivity
RNAshapes:
0.180
Mastr(seed):
0.003
Positive Predictive Value
RNAshapes:
0.484
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Mastr(seed):
0.051
Sensitivity
Carnac(20):
0.093
Mastr(seed):
0.003
Positive Predictive Value
Carnac(20):
0.892
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Mastr(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Mastr(seed):
0.051
Sensitivity
RNASLOpt:
0.158
Mastr(seed):
0.003
Positive Predictive Value
RNASLOpt:
0.502
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Mastr(seed):
0.051
Sensitivity
RSpredict(20):
0.124
Mastr(seed):
0.003
Positive Predictive Value
RSpredict(20):
0.590
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Mastr(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Mastr(seed):
0.051
Sensitivity
Multilign(20):
0.083
Mastr(seed):
0.003
Positive Predictive Value
Multilign(20):
0.762
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Mastr(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Mastr(seed):
0.051
Sensitivity
RNAwolf:
0.191
Mastr(seed):
0.003
Positive Predictive Value
RNAwolf:
0.330
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Mastr(seed):
0.051
Sensitivity
Murlet(20):
0.067
Mastr(seed):
0.003
Positive Predictive Value
Murlet(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Mastr(seed):
0.051
Sensitivity
Vsfold4:
0.125
Mastr(seed):
0.003
Positive Predictive Value
Vsfold4:
0.378
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Mastr(seed):
0.051
Sensitivity
Vsfold5:
0.125
Mastr(seed):
0.003
Positive Predictive Value
Vsfold5:
0.359
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Mastr(seed):
0.051
Sensitivity
RSpredict(seed):
0.075
Mastr(seed):
0.003
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Mastr(seed):
0.051
Sensitivity
CMfinder(20):
0.055
Mastr(seed):
0.003
Positive Predictive Value
CMfinder(20):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Mastr(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Mastr(seed):
0.051
Sensitivity
HotKnots:
0.051
Mastr(seed):
0.003
Positive Predictive Value
HotKnots:
0.624
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Mastr(seed):
0.051
Sensitivity
RNASampler(20):
0.035
Mastr(seed):
0.003
Positive Predictive Value
RNASampler(20):
0.877
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Mastr(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Mastr(seed):
0.051
Sensitivity
Cylofold:
0.040
Mastr(seed):
0.003
Positive Predictive Value
Cylofold:
0.606
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs Mastr(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Mastr(seed):
0.051
Sensitivity
Pknots:
0.033
Mastr(seed):
0.003
Positive Predictive Value
Pknots:
0.624
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs Mastr(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Mastr(seed):
0.051
Sensitivity
Mastr(20):
0.023
Mastr(seed):
0.003
Positive Predictive Value
Mastr(20):
0.790
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs Mastr(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Mastr(seed):
0.051
Sensitivity
Alterna:
0.030
Mastr(seed):
0.003
Positive Predictive Value
Alterna:
0.566
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs Mastr(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Mastr(seed):
0.051
Sensitivity
MCFold:
0.036
Mastr(seed):
0.003
Positive Predictive Value
MCFold:
0.435
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs Mastr(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Mastr(seed):
0.051
Sensitivity
RDfolder:
0.025
Mastr(seed):
0.003
Positive Predictive Value
RDfolder:
0.592
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
Mastr(seed):
0.051
Sensitivity
Murlet(seed):
0.008
Mastr(seed):
0.003
Positive Predictive Value
Murlet(seed):
0.811
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Mastr(seed):
0.051
Sensitivity
CMfinder(seed):
0.008
Mastr(seed):
0.003
Positive Predictive Value
CMfinder(seed):
0.757
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
Mastr(seed):
0.051
Sensitivity
TurboFold(seed):
0.007
Mastr(seed):
0.003
Positive Predictive Value
TurboFold(seed):
0.822
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
Mastr(seed):
0.051
Sensitivity
PPfold(seed):
0.006
Mastr(seed):
0.003
Positive Predictive Value
PPfold(seed):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.066
Mastr(seed):
0.051
Sensitivity
Carnac(seed):
0.005
Mastr(seed):
0.003
Positive Predictive Value
Carnac(seed):
0.867
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.063
Mastr(seed):
0.051
Sensitivity
RNASampler(seed):
0.005
Mastr(seed):
0.003
Positive Predictive Value
RNASampler(seed):
0.847
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
NanoFolder vs Mastr(seed)
Matthews Correlation Coefficient
NanoFolder:
0.052
Mastr(seed):
0.051
Sensitivity
NanoFolder:
0.016
Mastr(seed):
0.003
Positive Predictive Value
NanoFolder:
0.168
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 0.0529232299595
|
|
+
Mastr(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(seed):
0.051
Multilign(seed):
0.043
Sensitivity
Mastr(seed):
0.003
Multilign(seed):
0.003
Positive Predictive Value
Mastr(seed):
0.820
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Multilign(seed) |
1987
ContextFold vs Multilign(seed)
Matthews Correlation Coefficient
ContextFold:
0.680
Multilign(seed):
0.043
Sensitivity
ContextFold:
0.644
Multilign(seed):
0.003
Positive Predictive Value
ContextFold:
0.718
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
IPknot vs Multilign(seed)
Matthews Correlation Coefficient
IPknot:
0.548
Multilign(seed):
0.043
Sensitivity
IPknot:
0.475
Multilign(seed):
0.003
Positive Predictive Value
IPknot:
0.632
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.529
Multilign(seed):
0.043
Sensitivity
PETfold_pre2.0(seed):
0.343
Multilign(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Contrafold vs Multilign(seed)
Matthews Correlation Coefficient
Contrafold:
0.508
Multilign(seed):
0.043
Sensitivity
Contrafold:
0.481
Multilign(seed):
0.003
Positive Predictive Value
Contrafold:
0.537
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidFold vs Multilign(seed)
Matthews Correlation Coefficient
CentroidFold:
0.470
Multilign(seed):
0.043
Sensitivity
CentroidFold:
0.369
Multilign(seed):
0.003
Positive Predictive Value
CentroidFold:
0.600
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Sfold vs Multilign(seed)
Matthews Correlation Coefficient
Sfold:
0.469
Multilign(seed):
0.043
Sensitivity
Sfold:
0.412
Multilign(seed):
0.003
Positive Predictive Value
Sfold:
0.535
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MaxExpect vs Multilign(seed)
Matthews Correlation Coefficient
MaxExpect:
0.465
Multilign(seed):
0.043
Sensitivity
MaxExpect:
0.443
Multilign(seed):
0.003
Positive Predictive Value
MaxExpect:
0.488
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
ProbKnot vs Multilign(seed)
Matthews Correlation Coefficient
ProbKnot:
0.460
Multilign(seed):
0.043
Sensitivity
ProbKnot:
0.446
Multilign(seed):
0.003
Positive Predictive Value
ProbKnot:
0.475
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
UNAFold vs Multilign(seed)
Matthews Correlation Coefficient
UNAFold:
0.445
Multilign(seed):
0.043
Sensitivity
UNAFold:
0.445
Multilign(seed):
0.003
Positive Predictive Value
UNAFold:
0.446
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.438
Multilign(seed):
0.043
Sensitivity
CentroidAlifold(seed):
0.213
Multilign(seed):
0.003
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Fold vs Multilign(seed)
Matthews Correlation Coefficient
Fold:
0.437
Multilign(seed):
0.043
Sensitivity
Fold:
0.432
Multilign(seed):
0.003
Positive Predictive Value
Fold:
0.444
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAfold vs Multilign(seed)
Matthews Correlation Coefficient
RNAfold:
0.429
Multilign(seed):
0.043
Sensitivity
RNAfold:
0.422
Multilign(seed):
0.003
Positive Predictive Value
RNAfold:
0.437
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidHomfold‑LAST vs Multilign(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.428
Multilign(seed):
0.043
Sensitivity
CentroidHomfold‑LAST:
0.216
Multilign(seed):
0.003
Positive Predictive Value
CentroidHomfold‑LAST:
0.850
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PETfold_pre2.0(20) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.413
Multilign(seed):
0.043
Sensitivity
PETfold_pre2.0(20):
0.211
Multilign(seed):
0.003
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CentroidAlifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.411
Multilign(seed):
0.043
Sensitivity
CentroidAlifold(20):
0.190
Multilign(seed):
0.003
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PknotsRG vs Multilign(seed)
Matthews Correlation Coefficient
PknotsRG:
0.386
Multilign(seed):
0.043
Sensitivity
PknotsRG:
0.320
Multilign(seed):
0.003
Positive Predictive Value
PknotsRG:
0.465
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(seed) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.385
Multilign(seed):
0.043
Sensitivity
MXScarna(seed):
0.196
Multilign(seed):
0.003
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.381
Multilign(seed):
0.043
Sensitivity
RNAalifold(20):
0.185
Multilign(seed):
0.003
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAalifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.375
Multilign(seed):
0.043
Sensitivity
RNAalifold(seed):
0.169
Multilign(seed):
0.003
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Afold vs Multilign(seed)
Matthews Correlation Coefficient
Afold:
0.358
Multilign(seed):
0.043
Sensitivity
Afold:
0.295
Multilign(seed):
0.003
Positive Predictive Value
Afold:
0.434
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MXScarna(20) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.359
Multilign(seed):
0.043
Sensitivity
MXScarna(20):
0.181
Multilign(seed):
0.003
Positive Predictive Value
MXScarna(20):
0.710
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CRWrnafold vs Multilign(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.346
Multilign(seed):
0.043
Sensitivity
CRWrnafold:
0.275
Multilign(seed):
0.003
Positive Predictive Value
CRWrnafold:
0.435
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAsubopt vs Multilign(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.336
Multilign(seed):
0.043
Sensitivity
RNAsubopt:
0.217
Multilign(seed):
0.003
Positive Predictive Value
RNAsubopt:
0.521
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
McQFold vs Multilign(seed)
Matthews Correlation Coefficient
McQFold:
0.314
Multilign(seed):
0.043
Sensitivity
McQFold:
0.205
Multilign(seed):
0.003
Positive Predictive Value
McQFold:
0.480
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(20) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.301
Multilign(seed):
0.043
Sensitivity
TurboFold(20):
0.108
Multilign(seed):
0.003
Positive Predictive Value
TurboFold(20):
0.837
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(20) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(20):
0.298
Multilign(seed):
0.043
Sensitivity
PPfold(20):
0.101
Multilign(seed):
0.003
Positive Predictive Value
PPfold(20):
0.879
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAshapes vs Multilign(seed)
Matthews Correlation Coefficient
RNAshapes:
0.295
Multilign(seed):
0.043
Sensitivity
RNAshapes:
0.180
Multilign(seed):
0.003
Positive Predictive Value
RNAshapes:
0.484
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(20) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(20):
0.288
Multilign(seed):
0.043
Sensitivity
Carnac(20):
0.093
Multilign(seed):
0.003
Positive Predictive Value
Carnac(20):
0.892
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASLOpt vs Multilign(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.281
Multilign(seed):
0.043
Sensitivity
RNASLOpt:
0.158
Multilign(seed):
0.003
Positive Predictive Value
RNASLOpt:
0.502
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(20) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.270
Multilign(seed):
0.043
Sensitivity
RSpredict(20):
0.124
Multilign(seed):
0.003
Positive Predictive Value
RSpredict(20):
0.590
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Multilign(20) vs Multilign(seed)
Matthews Correlation Coefficient
Multilign(20):
0.251
Multilign(seed):
0.043
Sensitivity
Multilign(20):
0.083
Multilign(seed):
0.003
Positive Predictive Value
Multilign(20):
0.762
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNAwolf vs Multilign(seed)
Matthews Correlation Coefficient
RNAwolf:
0.251
Multilign(seed):
0.043
Sensitivity
RNAwolf:
0.191
Multilign(seed):
0.003
Positive Predictive Value
RNAwolf:
0.330
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(20) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(20):
0.230
Multilign(seed):
0.043
Sensitivity
Murlet(20):
0.067
Multilign(seed):
0.003
Positive Predictive Value
Murlet(20):
0.787
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold4 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold4:
0.217
Multilign(seed):
0.043
Sensitivity
Vsfold4:
0.125
Multilign(seed):
0.003
Positive Predictive Value
Vsfold4:
0.378
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Vsfold5 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold5:
0.211
Multilign(seed):
0.043
Sensitivity
Vsfold5:
0.125
Multilign(seed):
0.003
Positive Predictive Value
Vsfold5:
0.359
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RSpredict(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.212
Multilign(seed):
0.043
Sensitivity
RSpredict(seed):
0.075
Multilign(seed):
0.003
Positive Predictive Value
RSpredict(seed):
0.597
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(20) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.210
Multilign(seed):
0.043
Sensitivity
CMfinder(20):
0.055
Multilign(seed):
0.003
Positive Predictive Value
CMfinder(20):
0.794
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
HotKnots vs Multilign(seed)
Matthews Correlation Coefficient
HotKnots:
0.179
Multilign(seed):
0.043
Sensitivity
HotKnots:
0.051
Multilign(seed):
0.003
Positive Predictive Value
HotKnots:
0.624
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.176
Multilign(seed):
0.043
Sensitivity
RNASampler(20):
0.035
Multilign(seed):
0.003
Positive Predictive Value
RNASampler(20):
0.877
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Cylofold vs Multilign(seed)
Matthews Correlation Coefficient
Cylofold:
0.155
Multilign(seed):
0.043
Sensitivity
Cylofold:
0.040
Multilign(seed):
0.003
Positive Predictive Value
Cylofold:
0.606
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Pknots vs Multilign(seed)
Matthews Correlation Coefficient
Pknots:
0.143
Multilign(seed):
0.043
Sensitivity
Pknots:
0.033
Multilign(seed):
0.003
Positive Predictive Value
Pknots:
0.624
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(20) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(20):
0.134
Multilign(seed):
0.043
Sensitivity
Mastr(20):
0.023
Multilign(seed):
0.003
Positive Predictive Value
Mastr(20):
0.790
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Alterna vs Multilign(seed)
Matthews Correlation Coefficient
Alterna:
0.129
Multilign(seed):
0.043
Sensitivity
Alterna:
0.030
Multilign(seed):
0.003
Positive Predictive Value
Alterna:
0.566
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
MCFold vs Multilign(seed)
Matthews Correlation Coefficient
MCFold:
0.125
Multilign(seed):
0.043
Sensitivity
MCFold:
0.036
Multilign(seed):
0.003
Positive Predictive Value
MCFold:
0.435
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RDfolder vs Multilign(seed)
Matthews Correlation Coefficient
RDfolder:
0.122
Multilign(seed):
0.043
Sensitivity
RDfolder:
0.025
Multilign(seed):
0.003
Positive Predictive Value
RDfolder:
0.592
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Murlet(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.079
Multilign(seed):
0.043
Sensitivity
Murlet(seed):
0.008
Multilign(seed):
0.003
Positive Predictive Value
Murlet(seed):
0.811
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
CMfinder(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.080
Multilign(seed):
0.043
Sensitivity
CMfinder(seed):
0.008
Multilign(seed):
0.003
Positive Predictive Value
CMfinder(seed):
0.757
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
TurboFold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.076
Multilign(seed):
0.043
Sensitivity
TurboFold(seed):
0.007
Multilign(seed):
0.003
Positive Predictive Value
TurboFold(seed):
0.822
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
PPfold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.069
Multilign(seed):
0.043
Sensitivity
PPfold(seed):
0.006
Multilign(seed):
0.003
Positive Predictive Value
PPfold(seed):
0.794
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Carnac(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.066
Multilign(seed):
0.043
Sensitivity
Carnac(seed):
0.005
Multilign(seed):
0.003
Positive Predictive Value
Carnac(seed):
0.867
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
RNASampler(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.063
Multilign(seed):
0.043
Sensitivity
RNASampler(seed):
0.005
Multilign(seed):
0.003
Positive Predictive Value
RNASampler(seed):
0.847
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
NanoFolder vs Multilign(seed)
Matthews Correlation Coefficient
NanoFolder:
0.052
Multilign(seed):
0.043
Sensitivity
NanoFolder:
0.016
Multilign(seed):
0.003
Positive Predictive Value
NanoFolder:
0.168
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1987
Mastr(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(seed):
0.051
Multilign(seed):
0.043
Sensitivity
Mastr(seed):
0.003
Multilign(seed):
0.003
Positive Predictive Value
Mastr(seed):
0.820
Multilign(seed):
0.717
Number of pairs reference - predicted secondary structure: 1987
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|