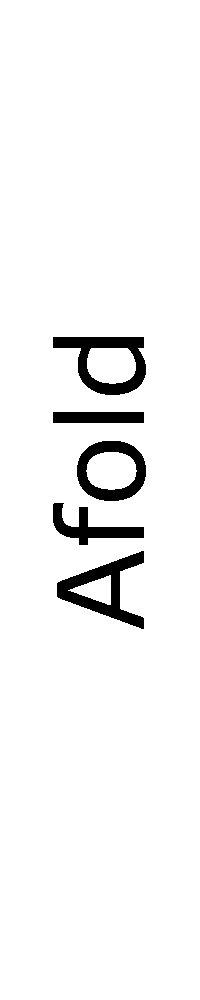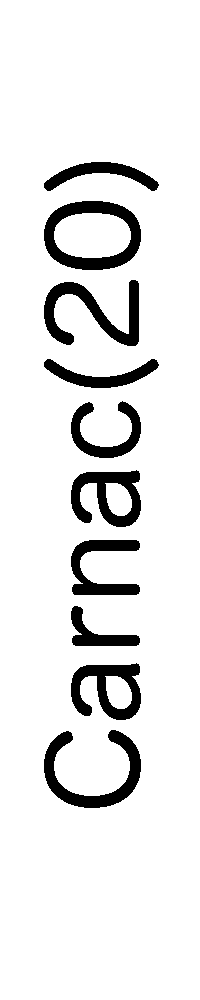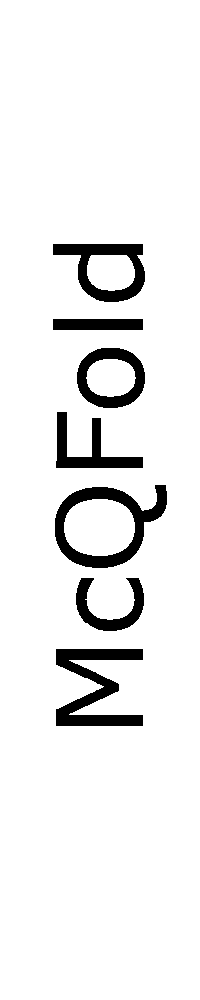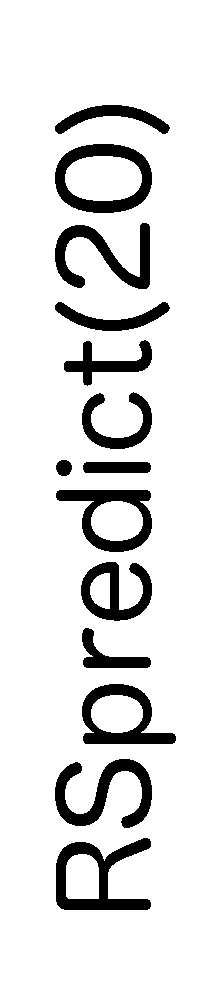| |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
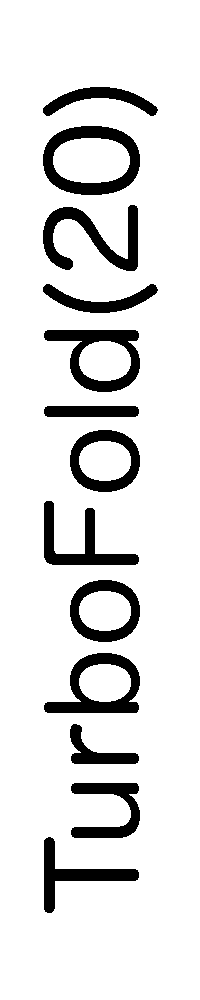 |
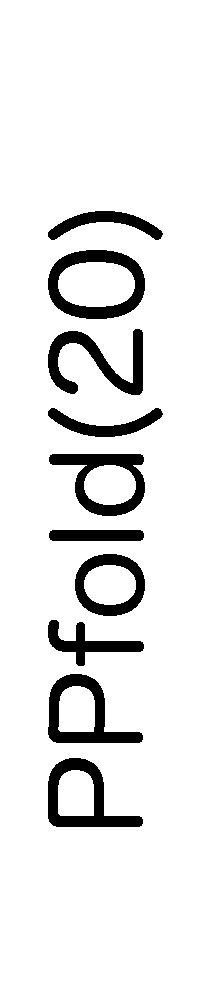 |
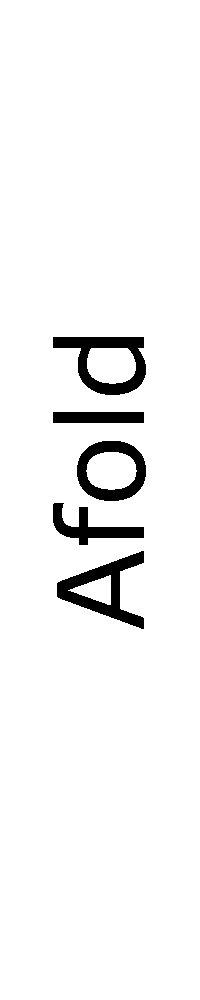 |
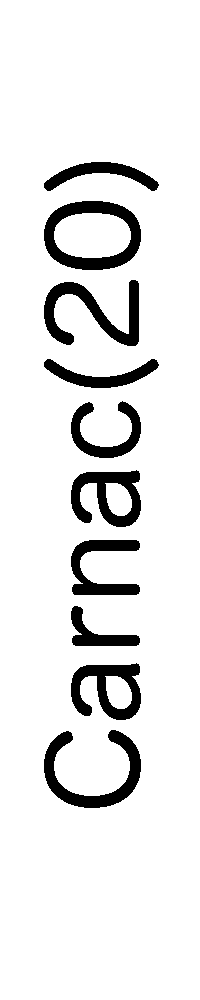 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
| ContextFold |
|
+
ContextFold vs PETfold_pre2.0(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
PETfold_pre2.0(seed):
0.737
Sensitivity
ContextFold:
0.717
PETfold_pre2.0(seed):
0.667
Positive Predictive Value
ContextFold:
0.794
PETfold_pre2.0(seed):
0.814
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidAlifold(seed):
0.611
Sensitivity
ContextFold:
0.717
CentroidAlifold(seed):
0.414
Positive Predictive Value
ContextFold:
0.794
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs IPknot
Matthews Correlation Coefficient
ContextFold:
0.754
IPknot:
0.607
Sensitivity
ContextFold:
0.717
IPknot:
0.551
Positive Predictive Value
ContextFold:
0.794
IPknot:
0.670
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
ContextFold:
0.754
PETfold_pre2.0(20):
0.575
Sensitivity
ContextFold:
0.717
PETfold_pre2.0(20):
0.410
Positive Predictive Value
ContextFold:
0.794
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidFold
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidFold:
0.571
Sensitivity
ContextFold:
0.717
CentroidFold:
0.515
Positive Predictive Value
ContextFold:
0.794
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidAlifold(20):
0.573
Sensitivity
ContextFold:
0.717
CentroidAlifold(20):
0.369
Positive Predictive Value
ContextFold:
0.794
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidHomfold‑LAST:
0.560
Sensitivity
ContextFold:
0.717
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
ContextFold:
0.794
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Contrafold
Matthews Correlation Coefficient
ContextFold:
0.754
Contrafold:
0.556
Sensitivity
ContextFold:
0.717
Contrafold:
0.542
Positive Predictive Value
ContextFold:
0.794
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MXScarna(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
MXScarna(seed):
0.536
Sensitivity
ContextFold:
0.717
MXScarna(seed):
0.382
Positive Predictive Value
ContextFold:
0.794
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAalifold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
RNAalifold(20):
0.532
Sensitivity
ContextFold:
0.717
RNAalifold(20):
0.359
Positive Predictive Value
ContextFold:
0.794
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Sfold
Matthews Correlation Coefficient
ContextFold:
0.754
Sfold:
0.527
Sensitivity
ContextFold:
0.717
Sfold:
0.477
Positive Predictive Value
ContextFold:
0.794
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAalifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
RNAalifold(seed):
0.523
Sensitivity
ContextFold:
0.717
RNAalifold(seed):
0.328
Positive Predictive Value
ContextFold:
0.794
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MaxExpect
Matthews Correlation Coefficient
ContextFold:
0.754
MaxExpect:
0.516
Sensitivity
ContextFold:
0.717
MaxExpect:
0.505
Positive Predictive Value
ContextFold:
0.794
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs ProbKnot
Matthews Correlation Coefficient
ContextFold:
0.754
ProbKnot:
0.512
Sensitivity
ContextFold:
0.717
ProbKnot:
0.508
Positive Predictive Value
ContextFold:
0.794
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MXScarna(20)
Matthews Correlation Coefficient
ContextFold:
0.754
MXScarna(20):
0.500
Sensitivity
ContextFold:
0.717
MXScarna(20):
0.353
Positive Predictive Value
ContextFold:
0.794
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs UNAFold
Matthews Correlation Coefficient
ContextFold:
0.754
UNAFold:
0.494
Sensitivity
ContextFold:
0.717
UNAFold:
0.491
Positive Predictive Value
ContextFold:
0.794
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Fold
Matthews Correlation Coefficient
ContextFold:
0.754
Fold:
0.493
Sensitivity
ContextFold:
0.717
Fold:
0.496
Positive Predictive Value
ContextFold:
0.794
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAfold
Matthews Correlation Coefficient
ContextFold:
0.754
RNAfold:
0.485
Sensitivity
ContextFold:
0.717
RNAfold:
0.486
Positive Predictive Value
ContextFold:
0.794
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PknotsRG
Matthews Correlation Coefficient
ContextFold:
0.754
PknotsRG:
0.474
Sensitivity
ContextFold:
0.717
PknotsRG:
0.440
Positive Predictive Value
ContextFold:
0.794
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CRWrnafold
Matthews Correlation Coefficient
ContextFold:
0.754
CRWrnafold:
0.430
Sensitivity
ContextFold:
0.717
CRWrnafold:
0.379
Positive Predictive Value
ContextFold:
0.794
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAsubopt
Matthews Correlation Coefficient
ContextFold:
0.754
RNAsubopt:
0.422
Sensitivity
ContextFold:
0.717
RNAsubopt:
0.344
Positive Predictive Value
ContextFold:
0.794
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs TurboFold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
TurboFold(20):
0.419
Sensitivity
ContextFold:
0.717
TurboFold(20):
0.210
Positive Predictive Value
ContextFold:
0.794
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PPfold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
PPfold(20):
0.416
Sensitivity
ContextFold:
0.717
PPfold(20):
0.197
Positive Predictive Value
ContextFold:
0.794
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Afold
Matthews Correlation Coefficient
ContextFold:
0.754
Afold:
0.408
Sensitivity
ContextFold:
0.717
Afold:
0.343
Positive Predictive Value
ContextFold:
0.794
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Carnac(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Carnac(20):
0.401
Sensitivity
ContextFold:
0.717
Carnac(20):
0.180
Positive Predictive Value
ContextFold:
0.794
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs McQFold
Matthews Correlation Coefficient
ContextFold:
0.754
McQFold:
0.389
Sensitivity
ContextFold:
0.717
McQFold:
0.308
Positive Predictive Value
ContextFold:
0.794
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RSpredict(20)
Matthews Correlation Coefficient
ContextFold:
0.754
RSpredict(20):
0.377
Sensitivity
ContextFold:
0.717
RSpredict(20):
0.241
Positive Predictive Value
ContextFold:
0.794
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAshapes
Matthews Correlation Coefficient
ContextFold:
0.754
RNAshapes:
0.367
Sensitivity
ContextFold:
0.717
RNAshapes:
0.285
Positive Predictive Value
ContextFold:
0.794
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNASLOpt
Matthews Correlation Coefficient
ContextFold:
0.754
RNASLOpt:
0.364
Sensitivity
ContextFold:
0.717
RNASLOpt:
0.261
Positive Predictive Value
ContextFold:
0.794
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Multilign(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Multilign(20):
0.350
Sensitivity
ContextFold:
0.717
Multilign(20):
0.161
Positive Predictive Value
ContextFold:
0.794
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Murlet(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Murlet(20):
0.321
Sensitivity
ContextFold:
0.717
Murlet(20):
0.131
Positive Predictive Value
ContextFold:
0.794
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNAwolf
Matthews Correlation Coefficient
ContextFold:
0.754
RNAwolf:
0.310
Sensitivity
ContextFold:
0.717
RNAwolf:
0.261
Positive Predictive Value
ContextFold:
0.794
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RSpredict(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
RSpredict(seed):
0.295
Sensitivity
ContextFold:
0.717
RSpredict(seed):
0.146
Positive Predictive Value
ContextFold:
0.794
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CMfinder(20)
Matthews Correlation Coefficient
ContextFold:
0.754
CMfinder(20):
0.292
Sensitivity
ContextFold:
0.717
CMfinder(20):
0.107
Positive Predictive Value
ContextFold:
0.794
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Vsfold4
Matthews Correlation Coefficient
ContextFold:
0.754
Vsfold4:
0.270
Sensitivity
ContextFold:
0.717
Vsfold4:
0.198
Positive Predictive Value
ContextFold:
0.794
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Vsfold5
Matthews Correlation Coefficient
ContextFold:
0.754
Vsfold5:
0.260
Sensitivity
ContextFold:
0.717
Vsfold5:
0.195
Positive Predictive Value
ContextFold:
0.794
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNASampler(20)
Matthews Correlation Coefficient
ContextFold:
0.754
RNASampler(20):
0.246
Sensitivity
ContextFold:
0.717
RNASampler(20):
0.069
Positive Predictive Value
ContextFold:
0.794
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs HotKnots
Matthews Correlation Coefficient
ContextFold:
0.754
HotKnots:
0.212
Sensitivity
ContextFold:
0.717
HotKnots:
0.069
Positive Predictive Value
ContextFold:
0.794
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Mastr(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Mastr(20):
0.187
Sensitivity
ContextFold:
0.717
Mastr(20):
0.044
Positive Predictive Value
ContextFold:
0.794
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Cylofold
Matthews Correlation Coefficient
ContextFold:
0.754
Cylofold:
0.178
Sensitivity
ContextFold:
0.717
Cylofold:
0.053
Positive Predictive Value
ContextFold:
0.794
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Pknots
Matthews Correlation Coefficient
ContextFold:
0.754
Pknots:
0.165
Sensitivity
ContextFold:
0.717
Pknots:
0.041
Positive Predictive Value
ContextFold:
0.794
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Alterna
Matthews Correlation Coefficient
ContextFold:
0.754
Alterna:
0.145
Sensitivity
ContextFold:
0.717
Alterna:
0.036
Positive Predictive Value
ContextFold:
0.794
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs MCFold
Matthews Correlation Coefficient
ContextFold:
0.754
MCFold:
0.138
Sensitivity
ContextFold:
0.717
MCFold:
0.044
Positive Predictive Value
ContextFold:
0.794
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RDfolder
Matthews Correlation Coefficient
ContextFold:
0.754
RDfolder:
0.138
Sensitivity
ContextFold:
0.717
RDfolder:
0.031
Positive Predictive Value
ContextFold:
0.794
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Murlet(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Murlet(seed):
0.110
Sensitivity
ContextFold:
0.717
Murlet(seed):
0.015
Positive Predictive Value
ContextFold:
0.794
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs CMfinder(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
CMfinder(seed):
0.111
Sensitivity
ContextFold:
0.717
CMfinder(seed):
0.016
Positive Predictive Value
ContextFold:
0.794
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs TurboFold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
TurboFold(seed):
0.106
Sensitivity
ContextFold:
0.717
TurboFold(seed):
0.014
Positive Predictive Value
ContextFold:
0.794
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs PPfold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
PPfold(seed):
0.097
Sensitivity
ContextFold:
0.717
PPfold(seed):
0.012
Positive Predictive Value
ContextFold:
0.794
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Carnac(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Carnac(seed):
0.093
Sensitivity
ContextFold:
0.717
Carnac(seed):
0.010
Positive Predictive Value
ContextFold:
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs RNASampler(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
RNASampler(seed):
0.088
Sensitivity
ContextFold:
0.717
RNASampler(seed):
0.009
Positive Predictive Value
ContextFold:
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Mastr(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Mastr(seed):
0.072
Sensitivity
ContextFold:
0.717
Mastr(seed):
0.006
Positive Predictive Value
ContextFold:
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs NanoFolder
Matthews Correlation Coefficient
ContextFold:
0.754
NanoFolder:
0.068
Sensitivity
ContextFold:
0.717
NanoFolder:
0.027
Positive Predictive Value
ContextFold:
0.794
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ContextFold vs Multilign(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Multilign(seed):
0.059
Sensitivity
ContextFold:
0.717
Multilign(seed):
0.005
Positive Predictive Value
ContextFold:
0.794
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PETfold_pre2.0(seed) |
1242
ContextFold vs PETfold_pre2.0(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
PETfold_pre2.0(seed):
0.737
Sensitivity
ContextFold:
0.717
PETfold_pre2.0(seed):
0.667
Positive Predictive Value
ContextFold:
0.794
PETfold_pre2.0(seed):
0.814
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PETfold_pre2.0(seed) vs CentroidAlifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidAlifold(seed):
0.611
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidAlifold(seed):
0.414
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs IPknot
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
IPknot:
0.607
Sensitivity
PETfold_pre2.0(seed):
0.667
IPknot:
0.551
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
IPknot:
0.670
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PETfold_pre2.0(20):
0.575
Sensitivity
PETfold_pre2.0(seed):
0.667
PETfold_pre2.0(20):
0.410
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidFold:
0.571
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidFold:
0.515
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidAlifold(20):
0.573
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidAlifold(20):
0.369
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidHomfold‑LAST:
0.560
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Contrafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Contrafold:
0.556
Sensitivity
PETfold_pre2.0(seed):
0.667
Contrafold:
0.542
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MXScarna(seed):
0.536
Sensitivity
PETfold_pre2.0(seed):
0.667
MXScarna(seed):
0.382
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAalifold(20):
0.532
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAalifold(20):
0.359
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Sfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Sfold:
0.527
Sensitivity
PETfold_pre2.0(seed):
0.667
Sfold:
0.477
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAalifold(seed):
0.523
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAalifold(seed):
0.328
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MaxExpect
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MaxExpect:
0.516
Sensitivity
PETfold_pre2.0(seed):
0.667
MaxExpect:
0.505
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs ProbKnot
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
ProbKnot:
0.512
Sensitivity
PETfold_pre2.0(seed):
0.667
ProbKnot:
0.508
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MXScarna(20):
0.500
Sensitivity
PETfold_pre2.0(seed):
0.667
MXScarna(20):
0.353
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs UNAFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
UNAFold:
0.494
Sensitivity
PETfold_pre2.0(seed):
0.667
UNAFold:
0.491
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Fold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Fold:
0.493
Sensitivity
PETfold_pre2.0(seed):
0.667
Fold:
0.496
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAfold:
0.485
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAfold:
0.486
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PknotsRG:
0.474
Sensitivity
PETfold_pre2.0(seed):
0.667
PknotsRG:
0.440
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CRWrnafold:
0.430
Sensitivity
PETfold_pre2.0(seed):
0.667
CRWrnafold:
0.379
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAsubopt:
0.422
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAsubopt:
0.344
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
TurboFold(20):
0.419
Sensitivity
PETfold_pre2.0(seed):
0.667
TurboFold(20):
0.210
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PPfold(20):
0.416
Sensitivity
PETfold_pre2.0(seed):
0.667
PPfold(20):
0.197
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Afold:
0.408
Sensitivity
PETfold_pre2.0(seed):
0.667
Afold:
0.343
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Carnac(20):
0.401
Sensitivity
PETfold_pre2.0(seed):
0.667
Carnac(20):
0.180
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
McQFold:
0.389
Sensitivity
PETfold_pre2.0(seed):
0.667
McQFold:
0.308
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RSpredict(20):
0.377
Sensitivity
PETfold_pre2.0(seed):
0.667
RSpredict(20):
0.241
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAshapes:
0.367
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAshapes:
0.285
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNASLOpt:
0.364
Sensitivity
PETfold_pre2.0(seed):
0.667
RNASLOpt:
0.261
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Multilign(20):
0.350
Sensitivity
PETfold_pre2.0(seed):
0.667
Multilign(20):
0.161
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Murlet(20):
0.321
Sensitivity
PETfold_pre2.0(seed):
0.667
Murlet(20):
0.131
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAwolf:
0.310
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAwolf:
0.261
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RSpredict(seed):
0.295
Sensitivity
PETfold_pre2.0(seed):
0.667
RSpredict(seed):
0.146
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CMfinder(20):
0.292
Sensitivity
PETfold_pre2.0(seed):
0.667
CMfinder(20):
0.107
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Vsfold4:
0.270
Sensitivity
PETfold_pre2.0(seed):
0.667
Vsfold4:
0.198
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Vsfold5:
0.260
Sensitivity
PETfold_pre2.0(seed):
0.667
Vsfold5:
0.195
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNASampler(20):
0.246
Sensitivity
PETfold_pre2.0(seed):
0.667
RNASampler(20):
0.069
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
HotKnots:
0.212
Sensitivity
PETfold_pre2.0(seed):
0.667
HotKnots:
0.069
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Mastr(20):
0.187
Sensitivity
PETfold_pre2.0(seed):
0.667
Mastr(20):
0.044
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Cylofold:
0.178
Sensitivity
PETfold_pre2.0(seed):
0.667
Cylofold:
0.053
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Pknots:
0.165
Sensitivity
PETfold_pre2.0(seed):
0.667
Pknots:
0.041
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Alterna:
0.145
Sensitivity
PETfold_pre2.0(seed):
0.667
Alterna:
0.036
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MCFold:
0.138
Sensitivity
PETfold_pre2.0(seed):
0.667
MCFold:
0.044
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RDfolder:
0.138
Sensitivity
PETfold_pre2.0(seed):
0.667
RDfolder:
0.031
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Murlet(seed):
0.110
Sensitivity
PETfold_pre2.0(seed):
0.667
Murlet(seed):
0.015
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CMfinder(seed):
0.111
Sensitivity
PETfold_pre2.0(seed):
0.667
CMfinder(seed):
0.016
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
TurboFold(seed):
0.106
Sensitivity
PETfold_pre2.0(seed):
0.667
TurboFold(seed):
0.014
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PPfold(seed):
0.097
Sensitivity
PETfold_pre2.0(seed):
0.667
PPfold(seed):
0.012
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Carnac(seed):
0.093
Sensitivity
PETfold_pre2.0(seed):
0.667
Carnac(seed):
0.010
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNASampler(seed):
0.088
Sensitivity
PETfold_pre2.0(seed):
0.667
RNASampler(seed):
0.009
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Mastr(seed):
0.072
Sensitivity
PETfold_pre2.0(seed):
0.667
Mastr(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
NanoFolder:
0.068
Sensitivity
PETfold_pre2.0(seed):
0.667
NanoFolder:
0.027
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Multilign(seed):
0.059
Sensitivity
PETfold_pre2.0(seed):
0.667
Multilign(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidAlifold(seed) |
1242
ContextFold vs CentroidAlifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidAlifold(seed):
0.611
Sensitivity
ContextFold:
0.717
CentroidAlifold(seed):
0.414
Positive Predictive Value
ContextFold:
0.794
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CentroidAlifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidAlifold(seed):
0.611
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidAlifold(seed):
0.414
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(seed):
0.901
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
CentroidAlifold(seed) vs IPknot
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
IPknot:
0.607
Sensitivity
CentroidAlifold(seed):
0.414
IPknot:
0.551
Positive Predictive Value
CentroidAlifold(seed):
0.901
IPknot:
0.670
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.01962644653e-06
|
+
CentroidAlifold(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PETfold_pre2.0(20):
0.575
Sensitivity
CentroidAlifold(seed):
0.414
PETfold_pre2.0(20):
0.410
Positive Predictive Value
CentroidAlifold(seed):
0.901
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CentroidFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CentroidFold:
0.571
Sensitivity
CentroidAlifold(seed):
0.414
CentroidFold:
0.515
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CentroidAlifold(20):
0.573
Sensitivity
CentroidAlifold(seed):
0.414
CentroidAlifold(20):
0.369
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CentroidHomfold‑LAST:
0.560
Sensitivity
CentroidAlifold(seed):
0.414
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Contrafold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Contrafold:
0.556
Sensitivity
CentroidAlifold(seed):
0.414
Contrafold:
0.542
Positive Predictive Value
CentroidAlifold(seed):
0.901
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MXScarna(seed):
0.536
Sensitivity
CentroidAlifold(seed):
0.414
MXScarna(seed):
0.382
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAalifold(20):
0.532
Sensitivity
CentroidAlifold(seed):
0.414
RNAalifold(20):
0.359
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Sfold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Sfold:
0.527
Sensitivity
CentroidAlifold(seed):
0.414
Sfold:
0.477
Positive Predictive Value
CentroidAlifold(seed):
0.901
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAalifold(seed):
0.523
Sensitivity
CentroidAlifold(seed):
0.414
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MaxExpect
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MaxExpect:
0.516
Sensitivity
CentroidAlifold(seed):
0.414
MaxExpect:
0.505
Positive Predictive Value
CentroidAlifold(seed):
0.901
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs ProbKnot
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
ProbKnot:
0.512
Sensitivity
CentroidAlifold(seed):
0.414
ProbKnot:
0.508
Positive Predictive Value
CentroidAlifold(seed):
0.901
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MXScarna(20):
0.500
Sensitivity
CentroidAlifold(seed):
0.414
MXScarna(20):
0.353
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs UNAFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
UNAFold:
0.494
Sensitivity
CentroidAlifold(seed):
0.414
UNAFold:
0.491
Positive Predictive Value
CentroidAlifold(seed):
0.901
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Fold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Fold:
0.493
Sensitivity
CentroidAlifold(seed):
0.414
Fold:
0.496
Positive Predictive Value
CentroidAlifold(seed):
0.901
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAfold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAfold:
0.485
Sensitivity
CentroidAlifold(seed):
0.414
RNAfold:
0.486
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PknotsRG:
0.474
Sensitivity
CentroidAlifold(seed):
0.414
PknotsRG:
0.440
Positive Predictive Value
CentroidAlifold(seed):
0.901
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CRWrnafold:
0.430
Sensitivity
CentroidAlifold(seed):
0.414
CRWrnafold:
0.379
Positive Predictive Value
CentroidAlifold(seed):
0.901
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAsubopt:
0.422
Sensitivity
CentroidAlifold(seed):
0.414
RNAsubopt:
0.344
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
TurboFold(20):
0.419
Sensitivity
CentroidAlifold(seed):
0.414
TurboFold(20):
0.210
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PPfold(20):
0.416
Sensitivity
CentroidAlifold(seed):
0.414
PPfold(20):
0.197
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Afold:
0.408
Sensitivity
CentroidAlifold(seed):
0.414
Afold:
0.343
Positive Predictive Value
CentroidAlifold(seed):
0.901
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Carnac(20):
0.401
Sensitivity
CentroidAlifold(seed):
0.414
Carnac(20):
0.180
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
McQFold:
0.389
Sensitivity
CentroidAlifold(seed):
0.414
McQFold:
0.308
Positive Predictive Value
CentroidAlifold(seed):
0.901
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RSpredict(20):
0.377
Sensitivity
CentroidAlifold(seed):
0.414
RSpredict(20):
0.241
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAshapes:
0.367
Sensitivity
CentroidAlifold(seed):
0.414
RNAshapes:
0.285
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNASLOpt:
0.364
Sensitivity
CentroidAlifold(seed):
0.414
RNASLOpt:
0.261
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Multilign(20):
0.350
Sensitivity
CentroidAlifold(seed):
0.414
Multilign(20):
0.161
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Murlet(20):
0.321
Sensitivity
CentroidAlifold(seed):
0.414
Murlet(20):
0.131
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAwolf:
0.310
Sensitivity
CentroidAlifold(seed):
0.414
RNAwolf:
0.261
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RSpredict(seed):
0.295
Sensitivity
CentroidAlifold(seed):
0.414
RSpredict(seed):
0.146
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CMfinder(20):
0.292
Sensitivity
CentroidAlifold(seed):
0.414
CMfinder(20):
0.107
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Vsfold4:
0.270
Sensitivity
CentroidAlifold(seed):
0.414
Vsfold4:
0.198
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Vsfold5:
0.260
Sensitivity
CentroidAlifold(seed):
0.414
Vsfold5:
0.195
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNASampler(20):
0.246
Sensitivity
CentroidAlifold(seed):
0.414
RNASampler(20):
0.069
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
HotKnots:
0.212
Sensitivity
CentroidAlifold(seed):
0.414
HotKnots:
0.069
Positive Predictive Value
CentroidAlifold(seed):
0.901
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Mastr(20):
0.187
Sensitivity
CentroidAlifold(seed):
0.414
Mastr(20):
0.044
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Cylofold:
0.178
Sensitivity
CentroidAlifold(seed):
0.414
Cylofold:
0.053
Positive Predictive Value
CentroidAlifold(seed):
0.901
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Pknots:
0.165
Sensitivity
CentroidAlifold(seed):
0.414
Pknots:
0.041
Positive Predictive Value
CentroidAlifold(seed):
0.901
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Alterna:
0.145
Sensitivity
CentroidAlifold(seed):
0.414
Alterna:
0.036
Positive Predictive Value
CentroidAlifold(seed):
0.901
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MCFold:
0.138
Sensitivity
CentroidAlifold(seed):
0.414
MCFold:
0.044
Positive Predictive Value
CentroidAlifold(seed):
0.901
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RDfolder:
0.138
Sensitivity
CentroidAlifold(seed):
0.414
RDfolder:
0.031
Positive Predictive Value
CentroidAlifold(seed):
0.901
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Murlet(seed):
0.110
Sensitivity
CentroidAlifold(seed):
0.414
Murlet(seed):
0.015
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CMfinder(seed):
0.111
Sensitivity
CentroidAlifold(seed):
0.414
CMfinder(seed):
0.016
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
TurboFold(seed):
0.106
Sensitivity
CentroidAlifold(seed):
0.414
TurboFold(seed):
0.014
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PPfold(seed):
0.097
Sensitivity
CentroidAlifold(seed):
0.414
PPfold(seed):
0.012
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Carnac(seed):
0.093
Sensitivity
CentroidAlifold(seed):
0.414
Carnac(seed):
0.010
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNASampler(seed):
0.088
Sensitivity
CentroidAlifold(seed):
0.414
RNASampler(seed):
0.009
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Mastr(seed):
0.072
Sensitivity
CentroidAlifold(seed):
0.414
Mastr(seed):
0.006
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
NanoFolder:
0.068
Sensitivity
CentroidAlifold(seed):
0.414
NanoFolder:
0.027
Positive Predictive Value
CentroidAlifold(seed):
0.901
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Multilign(seed):
0.059
Sensitivity
CentroidAlifold(seed):
0.414
Multilign(seed):
0.005
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| IPknot |
1242
ContextFold vs IPknot
Matthews Correlation Coefficient
ContextFold:
0.754
IPknot:
0.607
Sensitivity
ContextFold:
0.717
IPknot:
0.551
Positive Predictive Value
ContextFold:
0.794
IPknot:
0.670
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs IPknot
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
IPknot:
0.607
Sensitivity
PETfold_pre2.0(seed):
0.667
IPknot:
0.551
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
IPknot:
0.670
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs IPknot
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
IPknot:
0.607
Sensitivity
CentroidAlifold(seed):
0.414
IPknot:
0.551
Positive Predictive Value
CentroidAlifold(seed):
0.901
IPknot:
0.670
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.01962644653e-06
|
|
+
IPknot vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
IPknot:
0.607
PETfold_pre2.0(20):
0.575
Sensitivity
IPknot:
0.551
PETfold_pre2.0(20):
0.410
Positive Predictive Value
IPknot:
0.670
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidFold
Matthews Correlation Coefficient
IPknot:
0.607
CentroidFold:
0.571
Sensitivity
IPknot:
0.551
CentroidFold:
0.515
Positive Predictive Value
IPknot:
0.670
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidAlifold(20)
Matthews Correlation Coefficient
IPknot:
0.607
CentroidAlifold(20):
0.573
Sensitivity
IPknot:
0.551
CentroidAlifold(20):
0.369
Positive Predictive Value
IPknot:
0.670
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
IPknot:
0.607
CentroidHomfold‑LAST:
0.560
Sensitivity
IPknot:
0.551
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
IPknot:
0.670
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Contrafold
Matthews Correlation Coefficient
IPknot:
0.607
Contrafold:
0.556
Sensitivity
IPknot:
0.551
Contrafold:
0.542
Positive Predictive Value
IPknot:
0.670
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MXScarna(seed)
Matthews Correlation Coefficient
IPknot:
0.607
MXScarna(seed):
0.536
Sensitivity
IPknot:
0.551
MXScarna(seed):
0.382
Positive Predictive Value
IPknot:
0.670
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAalifold(20)
Matthews Correlation Coefficient
IPknot:
0.607
RNAalifold(20):
0.532
Sensitivity
IPknot:
0.551
RNAalifold(20):
0.359
Positive Predictive Value
IPknot:
0.670
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Sfold
Matthews Correlation Coefficient
IPknot:
0.607
Sfold:
0.527
Sensitivity
IPknot:
0.551
Sfold:
0.477
Positive Predictive Value
IPknot:
0.670
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAalifold(seed)
Matthews Correlation Coefficient
IPknot:
0.607
RNAalifold(seed):
0.523
Sensitivity
IPknot:
0.551
RNAalifold(seed):
0.328
Positive Predictive Value
IPknot:
0.670
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MaxExpect
Matthews Correlation Coefficient
IPknot:
0.607
MaxExpect:
0.516
Sensitivity
IPknot:
0.551
MaxExpect:
0.505
Positive Predictive Value
IPknot:
0.670
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs ProbKnot
Matthews Correlation Coefficient
IPknot:
0.607
ProbKnot:
0.512
Sensitivity
IPknot:
0.551
ProbKnot:
0.508
Positive Predictive Value
IPknot:
0.670
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MXScarna(20)
Matthews Correlation Coefficient
IPknot:
0.607
MXScarna(20):
0.500
Sensitivity
IPknot:
0.551
MXScarna(20):
0.353
Positive Predictive Value
IPknot:
0.670
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs UNAFold
Matthews Correlation Coefficient
IPknot:
0.607
UNAFold:
0.494
Sensitivity
IPknot:
0.551
UNAFold:
0.491
Positive Predictive Value
IPknot:
0.670
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Fold
Matthews Correlation Coefficient
IPknot:
0.607
Fold:
0.493
Sensitivity
IPknot:
0.551
Fold:
0.496
Positive Predictive Value
IPknot:
0.670
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAfold
Matthews Correlation Coefficient
IPknot:
0.607
RNAfold:
0.485
Sensitivity
IPknot:
0.551
RNAfold:
0.486
Positive Predictive Value
IPknot:
0.670
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PknotsRG
Matthews Correlation Coefficient
IPknot:
0.607
PknotsRG:
0.474
Sensitivity
IPknot:
0.551
PknotsRG:
0.440
Positive Predictive Value
IPknot:
0.670
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CRWrnafold
Matthews Correlation Coefficient
IPknot:
0.607
CRWrnafold:
0.430
Sensitivity
IPknot:
0.551
CRWrnafold:
0.379
Positive Predictive Value
IPknot:
0.670
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAsubopt
Matthews Correlation Coefficient
IPknot:
0.607
RNAsubopt:
0.422
Sensitivity
IPknot:
0.551
RNAsubopt:
0.344
Positive Predictive Value
IPknot:
0.670
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs TurboFold(20)
Matthews Correlation Coefficient
IPknot:
0.607
TurboFold(20):
0.419
Sensitivity
IPknot:
0.551
TurboFold(20):
0.210
Positive Predictive Value
IPknot:
0.670
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PPfold(20)
Matthews Correlation Coefficient
IPknot:
0.607
PPfold(20):
0.416
Sensitivity
IPknot:
0.551
PPfold(20):
0.197
Positive Predictive Value
IPknot:
0.670
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Afold
Matthews Correlation Coefficient
IPknot:
0.607
Afold:
0.408
Sensitivity
IPknot:
0.551
Afold:
0.343
Positive Predictive Value
IPknot:
0.670
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Carnac(20)
Matthews Correlation Coefficient
IPknot:
0.607
Carnac(20):
0.401
Sensitivity
IPknot:
0.551
Carnac(20):
0.180
Positive Predictive Value
IPknot:
0.670
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs McQFold
Matthews Correlation Coefficient
IPknot:
0.607
McQFold:
0.389
Sensitivity
IPknot:
0.551
McQFold:
0.308
Positive Predictive Value
IPknot:
0.670
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RSpredict(20)
Matthews Correlation Coefficient
IPknot:
0.607
RSpredict(20):
0.377
Sensitivity
IPknot:
0.551
RSpredict(20):
0.241
Positive Predictive Value
IPknot:
0.670
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAshapes
Matthews Correlation Coefficient
IPknot:
0.607
RNAshapes:
0.367
Sensitivity
IPknot:
0.551
RNAshapes:
0.285
Positive Predictive Value
IPknot:
0.670
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNASLOpt
Matthews Correlation Coefficient
IPknot:
0.607
RNASLOpt:
0.364
Sensitivity
IPknot:
0.551
RNASLOpt:
0.261
Positive Predictive Value
IPknot:
0.670
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Multilign(20)
Matthews Correlation Coefficient
IPknot:
0.607
Multilign(20):
0.350
Sensitivity
IPknot:
0.551
Multilign(20):
0.161
Positive Predictive Value
IPknot:
0.670
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Murlet(20)
Matthews Correlation Coefficient
IPknot:
0.607
Murlet(20):
0.321
Sensitivity
IPknot:
0.551
Murlet(20):
0.131
Positive Predictive Value
IPknot:
0.670
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNAwolf
Matthews Correlation Coefficient
IPknot:
0.607
RNAwolf:
0.310
Sensitivity
IPknot:
0.551
RNAwolf:
0.261
Positive Predictive Value
IPknot:
0.670
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RSpredict(seed)
Matthews Correlation Coefficient
IPknot:
0.607
RSpredict(seed):
0.295
Sensitivity
IPknot:
0.551
RSpredict(seed):
0.146
Positive Predictive Value
IPknot:
0.670
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CMfinder(20)
Matthews Correlation Coefficient
IPknot:
0.607
CMfinder(20):
0.292
Sensitivity
IPknot:
0.551
CMfinder(20):
0.107
Positive Predictive Value
IPknot:
0.670
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Vsfold4
Matthews Correlation Coefficient
IPknot:
0.607
Vsfold4:
0.270
Sensitivity
IPknot:
0.551
Vsfold4:
0.198
Positive Predictive Value
IPknot:
0.670
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Vsfold5
Matthews Correlation Coefficient
IPknot:
0.607
Vsfold5:
0.260
Sensitivity
IPknot:
0.551
Vsfold5:
0.195
Positive Predictive Value
IPknot:
0.670
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNASampler(20)
Matthews Correlation Coefficient
IPknot:
0.607
RNASampler(20):
0.246
Sensitivity
IPknot:
0.551
RNASampler(20):
0.069
Positive Predictive Value
IPknot:
0.670
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs HotKnots
Matthews Correlation Coefficient
IPknot:
0.607
HotKnots:
0.212
Sensitivity
IPknot:
0.551
HotKnots:
0.069
Positive Predictive Value
IPknot:
0.670
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Mastr(20)
Matthews Correlation Coefficient
IPknot:
0.607
Mastr(20):
0.187
Sensitivity
IPknot:
0.551
Mastr(20):
0.044
Positive Predictive Value
IPknot:
0.670
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Cylofold
Matthews Correlation Coefficient
IPknot:
0.607
Cylofold:
0.178
Sensitivity
IPknot:
0.551
Cylofold:
0.053
Positive Predictive Value
IPknot:
0.670
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Pknots
Matthews Correlation Coefficient
IPknot:
0.607
Pknots:
0.165
Sensitivity
IPknot:
0.551
Pknots:
0.041
Positive Predictive Value
IPknot:
0.670
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Alterna
Matthews Correlation Coefficient
IPknot:
0.607
Alterna:
0.145
Sensitivity
IPknot:
0.551
Alterna:
0.036
Positive Predictive Value
IPknot:
0.670
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs MCFold
Matthews Correlation Coefficient
IPknot:
0.607
MCFold:
0.138
Sensitivity
IPknot:
0.551
MCFold:
0.044
Positive Predictive Value
IPknot:
0.670
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RDfolder
Matthews Correlation Coefficient
IPknot:
0.607
RDfolder:
0.138
Sensitivity
IPknot:
0.551
RDfolder:
0.031
Positive Predictive Value
IPknot:
0.670
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Murlet(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Murlet(seed):
0.110
Sensitivity
IPknot:
0.551
Murlet(seed):
0.015
Positive Predictive Value
IPknot:
0.670
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs CMfinder(seed)
Matthews Correlation Coefficient
IPknot:
0.607
CMfinder(seed):
0.111
Sensitivity
IPknot:
0.551
CMfinder(seed):
0.016
Positive Predictive Value
IPknot:
0.670
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs TurboFold(seed)
Matthews Correlation Coefficient
IPknot:
0.607
TurboFold(seed):
0.106
Sensitivity
IPknot:
0.551
TurboFold(seed):
0.014
Positive Predictive Value
IPknot:
0.670
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs PPfold(seed)
Matthews Correlation Coefficient
IPknot:
0.607
PPfold(seed):
0.097
Sensitivity
IPknot:
0.551
PPfold(seed):
0.012
Positive Predictive Value
IPknot:
0.670
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Carnac(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Carnac(seed):
0.093
Sensitivity
IPknot:
0.551
Carnac(seed):
0.010
Positive Predictive Value
IPknot:
0.670
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs RNASampler(seed)
Matthews Correlation Coefficient
IPknot:
0.607
RNASampler(seed):
0.088
Sensitivity
IPknot:
0.551
RNASampler(seed):
0.009
Positive Predictive Value
IPknot:
0.670
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Mastr(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Mastr(seed):
0.072
Sensitivity
IPknot:
0.551
Mastr(seed):
0.006
Positive Predictive Value
IPknot:
0.670
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs NanoFolder
Matthews Correlation Coefficient
IPknot:
0.607
NanoFolder:
0.068
Sensitivity
IPknot:
0.551
NanoFolder:
0.027
Positive Predictive Value
IPknot:
0.670
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
IPknot vs Multilign(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Multilign(seed):
0.059
Sensitivity
IPknot:
0.551
Multilign(seed):
0.005
Positive Predictive Value
IPknot:
0.670
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PETfold_pre2.0(20) |
1242
ContextFold vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
ContextFold:
0.754
PETfold_pre2.0(20):
0.575
Sensitivity
ContextFold:
0.717
PETfold_pre2.0(20):
0.410
Positive Predictive Value
ContextFold:
0.794
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PETfold_pre2.0(20):
0.575
Sensitivity
PETfold_pre2.0(seed):
0.667
PETfold_pre2.0(20):
0.410
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PETfold_pre2.0(20):
0.575
Sensitivity
CentroidAlifold(seed):
0.414
PETfold_pre2.0(20):
0.410
Positive Predictive Value
CentroidAlifold(seed):
0.901
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs PETfold_pre2.0(20)
Matthews Correlation Coefficient
IPknot:
0.607
PETfold_pre2.0(20):
0.575
Sensitivity
IPknot:
0.551
PETfold_pre2.0(20):
0.410
Positive Predictive Value
IPknot:
0.670
PETfold_pre2.0(20):
0.807
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PETfold_pre2.0(20) vs CentroidFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CentroidFold:
0.571
Sensitivity
PETfold_pre2.0(20):
0.410
CentroidFold:
0.515
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.000706081118372
|
+
PETfold_pre2.0(20) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CentroidAlifold(20):
0.573
Sensitivity
PETfold_pre2.0(20):
0.410
CentroidAlifold(20):
0.369
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CentroidHomfold‑LAST:
0.560
Sensitivity
PETfold_pre2.0(20):
0.410
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Contrafold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Contrafold:
0.556
Sensitivity
PETfold_pre2.0(20):
0.410
Contrafold:
0.542
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MXScarna(seed):
0.536
Sensitivity
PETfold_pre2.0(20):
0.410
MXScarna(seed):
0.382
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAalifold(20):
0.532
Sensitivity
PETfold_pre2.0(20):
0.410
RNAalifold(20):
0.359
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Sfold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Sfold:
0.527
Sensitivity
PETfold_pre2.0(20):
0.410
Sfold:
0.477
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAalifold(seed):
0.523
Sensitivity
PETfold_pre2.0(20):
0.410
RNAalifold(seed):
0.328
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MaxExpect
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MaxExpect:
0.516
Sensitivity
PETfold_pre2.0(20):
0.410
MaxExpect:
0.505
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs ProbKnot
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
ProbKnot:
0.512
Sensitivity
PETfold_pre2.0(20):
0.410
ProbKnot:
0.508
Positive Predictive Value
PETfold_pre2.0(20):
0.807
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MXScarna(20):
0.500
Sensitivity
PETfold_pre2.0(20):
0.410
MXScarna(20):
0.353
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs UNAFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
UNAFold:
0.494
Sensitivity
PETfold_pre2.0(20):
0.410
UNAFold:
0.491
Positive Predictive Value
PETfold_pre2.0(20):
0.807
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Fold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Fold:
0.493
Sensitivity
PETfold_pre2.0(20):
0.410
Fold:
0.496
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAfold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAfold:
0.485
Sensitivity
PETfold_pre2.0(20):
0.410
RNAfold:
0.486
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
PknotsRG:
0.474
Sensitivity
PETfold_pre2.0(20):
0.410
PknotsRG:
0.440
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CRWrnafold:
0.430
Sensitivity
PETfold_pre2.0(20):
0.410
CRWrnafold:
0.379
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAsubopt:
0.422
Sensitivity
PETfold_pre2.0(20):
0.410
RNAsubopt:
0.344
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
TurboFold(20):
0.419
Sensitivity
PETfold_pre2.0(20):
0.410
TurboFold(20):
0.210
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
PPfold(20):
0.416
Sensitivity
PETfold_pre2.0(20):
0.410
PPfold(20):
0.197
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Afold:
0.408
Sensitivity
PETfold_pre2.0(20):
0.410
Afold:
0.343
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Carnac(20):
0.401
Sensitivity
PETfold_pre2.0(20):
0.410
Carnac(20):
0.180
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
McQFold:
0.389
Sensitivity
PETfold_pre2.0(20):
0.410
McQFold:
0.308
Positive Predictive Value
PETfold_pre2.0(20):
0.807
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RSpredict(20):
0.377
Sensitivity
PETfold_pre2.0(20):
0.410
RSpredict(20):
0.241
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAshapes:
0.367
Sensitivity
PETfold_pre2.0(20):
0.410
RNAshapes:
0.285
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNASLOpt:
0.364
Sensitivity
PETfold_pre2.0(20):
0.410
RNASLOpt:
0.261
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Multilign(20):
0.350
Sensitivity
PETfold_pre2.0(20):
0.410
Multilign(20):
0.161
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Murlet(20):
0.321
Sensitivity
PETfold_pre2.0(20):
0.410
Murlet(20):
0.131
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAwolf:
0.310
Sensitivity
PETfold_pre2.0(20):
0.410
RNAwolf:
0.261
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RSpredict(seed):
0.295
Sensitivity
PETfold_pre2.0(20):
0.410
RSpredict(seed):
0.146
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CMfinder(20):
0.292
Sensitivity
PETfold_pre2.0(20):
0.410
CMfinder(20):
0.107
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Vsfold4:
0.270
Sensitivity
PETfold_pre2.0(20):
0.410
Vsfold4:
0.198
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Vsfold5:
0.260
Sensitivity
PETfold_pre2.0(20):
0.410
Vsfold5:
0.195
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNASampler(20):
0.246
Sensitivity
PETfold_pre2.0(20):
0.410
RNASampler(20):
0.069
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
HotKnots:
0.212
Sensitivity
PETfold_pre2.0(20):
0.410
HotKnots:
0.069
Positive Predictive Value
PETfold_pre2.0(20):
0.807
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Mastr(20):
0.187
Sensitivity
PETfold_pre2.0(20):
0.410
Mastr(20):
0.044
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Cylofold:
0.178
Sensitivity
PETfold_pre2.0(20):
0.410
Cylofold:
0.053
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Pknots:
0.165
Sensitivity
PETfold_pre2.0(20):
0.410
Pknots:
0.041
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Alterna:
0.145
Sensitivity
PETfold_pre2.0(20):
0.410
Alterna:
0.036
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MCFold:
0.138
Sensitivity
PETfold_pre2.0(20):
0.410
MCFold:
0.044
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RDfolder:
0.138
Sensitivity
PETfold_pre2.0(20):
0.410
RDfolder:
0.031
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Murlet(seed):
0.110
Sensitivity
PETfold_pre2.0(20):
0.410
Murlet(seed):
0.015
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CMfinder(seed):
0.111
Sensitivity
PETfold_pre2.0(20):
0.410
CMfinder(seed):
0.016
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
TurboFold(seed):
0.106
Sensitivity
PETfold_pre2.0(20):
0.410
TurboFold(seed):
0.014
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
PPfold(seed):
0.097
Sensitivity
PETfold_pre2.0(20):
0.410
PPfold(seed):
0.012
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Carnac(seed):
0.093
Sensitivity
PETfold_pre2.0(20):
0.410
Carnac(seed):
0.010
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNASampler(seed):
0.088
Sensitivity
PETfold_pre2.0(20):
0.410
RNASampler(seed):
0.009
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Mastr(seed):
0.072
Sensitivity
PETfold_pre2.0(20):
0.410
Mastr(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
NanoFolder:
0.068
Sensitivity
PETfold_pre2.0(20):
0.410
NanoFolder:
0.027
Positive Predictive Value
PETfold_pre2.0(20):
0.807
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PETfold_pre2.0(20) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Multilign(seed):
0.059
Sensitivity
PETfold_pre2.0(20):
0.410
Multilign(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidFold |
1242
ContextFold vs CentroidFold
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidFold:
0.571
Sensitivity
ContextFold:
0.717
CentroidFold:
0.515
Positive Predictive Value
ContextFold:
0.794
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CentroidFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidFold:
0.571
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidFold:
0.515
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs CentroidFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CentroidFold:
0.571
Sensitivity
CentroidAlifold(seed):
0.414
CentroidFold:
0.515
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs CentroidFold
Matthews Correlation Coefficient
IPknot:
0.607
CentroidFold:
0.571
Sensitivity
IPknot:
0.551
CentroidFold:
0.515
Positive Predictive Value
IPknot:
0.670
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs CentroidFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CentroidFold:
0.571
Sensitivity
PETfold_pre2.0(20):
0.410
CentroidFold:
0.515
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.000706081118372
|
|
=
CentroidAlifold(20) vs CentroidFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CentroidFold:
0.571
Sensitivity
CentroidAlifold(20):
0.369
CentroidFold:
0.515
Positive Predictive Value
CentroidAlifold(20):
0.891
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.0167317061725
|
+
CentroidFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidFold:
0.571
CentroidHomfold‑LAST:
0.560
Sensitivity
CentroidFold:
0.515
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
CentroidFold:
0.633
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Contrafold
Matthews Correlation Coefficient
CentroidFold:
0.571
Contrafold:
0.556
Sensitivity
CentroidFold:
0.515
Contrafold:
0.542
Positive Predictive Value
CentroidFold:
0.633
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
MXScarna(seed):
0.536
Sensitivity
CentroidFold:
0.515
MXScarna(seed):
0.382
Positive Predictive Value
CentroidFold:
0.633
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAalifold(20):
0.532
Sensitivity
CentroidFold:
0.515
RNAalifold(20):
0.359
Positive Predictive Value
CentroidFold:
0.633
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Sfold
Matthews Correlation Coefficient
CentroidFold:
0.571
Sfold:
0.527
Sensitivity
CentroidFold:
0.515
Sfold:
0.477
Positive Predictive Value
CentroidFold:
0.633
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAalifold(seed):
0.523
Sensitivity
CentroidFold:
0.515
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidFold:
0.633
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MaxExpect
Matthews Correlation Coefficient
CentroidFold:
0.571
MaxExpect:
0.516
Sensitivity
CentroidFold:
0.515
MaxExpect:
0.505
Positive Predictive Value
CentroidFold:
0.633
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs ProbKnot
Matthews Correlation Coefficient
CentroidFold:
0.571
ProbKnot:
0.512
Sensitivity
CentroidFold:
0.515
ProbKnot:
0.508
Positive Predictive Value
CentroidFold:
0.633
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MXScarna(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
MXScarna(20):
0.500
Sensitivity
CentroidFold:
0.515
MXScarna(20):
0.353
Positive Predictive Value
CentroidFold:
0.633
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs UNAFold
Matthews Correlation Coefficient
CentroidFold:
0.571
UNAFold:
0.494
Sensitivity
CentroidFold:
0.515
UNAFold:
0.491
Positive Predictive Value
CentroidFold:
0.633
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Fold
Matthews Correlation Coefficient
CentroidFold:
0.571
Fold:
0.493
Sensitivity
CentroidFold:
0.515
Fold:
0.496
Positive Predictive Value
CentroidFold:
0.633
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAfold
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAfold:
0.485
Sensitivity
CentroidFold:
0.515
RNAfold:
0.486
Positive Predictive Value
CentroidFold:
0.633
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PknotsRG
Matthews Correlation Coefficient
CentroidFold:
0.571
PknotsRG:
0.474
Sensitivity
CentroidFold:
0.515
PknotsRG:
0.440
Positive Predictive Value
CentroidFold:
0.633
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CRWrnafold
Matthews Correlation Coefficient
CentroidFold:
0.571
CRWrnafold:
0.430
Sensitivity
CentroidFold:
0.515
CRWrnafold:
0.379
Positive Predictive Value
CentroidFold:
0.633
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAsubopt
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAsubopt:
0.422
Sensitivity
CentroidFold:
0.515
RNAsubopt:
0.344
Positive Predictive Value
CentroidFold:
0.633
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs TurboFold(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
TurboFold(20):
0.419
Sensitivity
CentroidFold:
0.515
TurboFold(20):
0.210
Positive Predictive Value
CentroidFold:
0.633
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PPfold(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
PPfold(20):
0.416
Sensitivity
CentroidFold:
0.515
PPfold(20):
0.197
Positive Predictive Value
CentroidFold:
0.633
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Afold
Matthews Correlation Coefficient
CentroidFold:
0.571
Afold:
0.408
Sensitivity
CentroidFold:
0.515
Afold:
0.343
Positive Predictive Value
CentroidFold:
0.633
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Carnac(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Carnac(20):
0.401
Sensitivity
CentroidFold:
0.515
Carnac(20):
0.180
Positive Predictive Value
CentroidFold:
0.633
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs McQFold
Matthews Correlation Coefficient
CentroidFold:
0.571
McQFold:
0.389
Sensitivity
CentroidFold:
0.515
McQFold:
0.308
Positive Predictive Value
CentroidFold:
0.633
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RSpredict(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
RSpredict(20):
0.377
Sensitivity
CentroidFold:
0.515
RSpredict(20):
0.241
Positive Predictive Value
CentroidFold:
0.633
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAshapes
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAshapes:
0.367
Sensitivity
CentroidFold:
0.515
RNAshapes:
0.285
Positive Predictive Value
CentroidFold:
0.633
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNASLOpt
Matthews Correlation Coefficient
CentroidFold:
0.571
RNASLOpt:
0.364
Sensitivity
CentroidFold:
0.515
RNASLOpt:
0.261
Positive Predictive Value
CentroidFold:
0.633
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Multilign(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Multilign(20):
0.350
Sensitivity
CentroidFold:
0.515
Multilign(20):
0.161
Positive Predictive Value
CentroidFold:
0.633
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Murlet(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Murlet(20):
0.321
Sensitivity
CentroidFold:
0.515
Murlet(20):
0.131
Positive Predictive Value
CentroidFold:
0.633
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNAwolf
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAwolf:
0.310
Sensitivity
CentroidFold:
0.515
RNAwolf:
0.261
Positive Predictive Value
CentroidFold:
0.633
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
RSpredict(seed):
0.295
Sensitivity
CentroidFold:
0.515
RSpredict(seed):
0.146
Positive Predictive Value
CentroidFold:
0.633
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CMfinder(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
CMfinder(20):
0.292
Sensitivity
CentroidFold:
0.515
CMfinder(20):
0.107
Positive Predictive Value
CentroidFold:
0.633
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Vsfold4
Matthews Correlation Coefficient
CentroidFold:
0.571
Vsfold4:
0.270
Sensitivity
CentroidFold:
0.515
Vsfold4:
0.198
Positive Predictive Value
CentroidFold:
0.633
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Vsfold5
Matthews Correlation Coefficient
CentroidFold:
0.571
Vsfold5:
0.260
Sensitivity
CentroidFold:
0.515
Vsfold5:
0.195
Positive Predictive Value
CentroidFold:
0.633
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNASampler(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNASampler(20):
0.246
Sensitivity
CentroidFold:
0.515
RNASampler(20):
0.069
Positive Predictive Value
CentroidFold:
0.633
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs HotKnots
Matthews Correlation Coefficient
CentroidFold:
0.571
HotKnots:
0.212
Sensitivity
CentroidFold:
0.515
HotKnots:
0.069
Positive Predictive Value
CentroidFold:
0.633
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Mastr(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Mastr(20):
0.187
Sensitivity
CentroidFold:
0.515
Mastr(20):
0.044
Positive Predictive Value
CentroidFold:
0.633
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Cylofold
Matthews Correlation Coefficient
CentroidFold:
0.571
Cylofold:
0.178
Sensitivity
CentroidFold:
0.515
Cylofold:
0.053
Positive Predictive Value
CentroidFold:
0.633
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Pknots
Matthews Correlation Coefficient
CentroidFold:
0.571
Pknots:
0.165
Sensitivity
CentroidFold:
0.515
Pknots:
0.041
Positive Predictive Value
CentroidFold:
0.633
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Alterna
Matthews Correlation Coefficient
CentroidFold:
0.571
Alterna:
0.145
Sensitivity
CentroidFold:
0.515
Alterna:
0.036
Positive Predictive Value
CentroidFold:
0.633
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs MCFold
Matthews Correlation Coefficient
CentroidFold:
0.571
MCFold:
0.138
Sensitivity
CentroidFold:
0.515
MCFold:
0.044
Positive Predictive Value
CentroidFold:
0.633
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RDfolder
Matthews Correlation Coefficient
CentroidFold:
0.571
RDfolder:
0.138
Sensitivity
CentroidFold:
0.515
RDfolder:
0.031
Positive Predictive Value
CentroidFold:
0.633
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Murlet(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Murlet(seed):
0.110
Sensitivity
CentroidFold:
0.515
Murlet(seed):
0.015
Positive Predictive Value
CentroidFold:
0.633
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
CMfinder(seed):
0.111
Sensitivity
CentroidFold:
0.515
CMfinder(seed):
0.016
Positive Predictive Value
CentroidFold:
0.633
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
TurboFold(seed):
0.106
Sensitivity
CentroidFold:
0.515
TurboFold(seed):
0.014
Positive Predictive Value
CentroidFold:
0.633
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs PPfold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
PPfold(seed):
0.097
Sensitivity
CentroidFold:
0.515
PPfold(seed):
0.012
Positive Predictive Value
CentroidFold:
0.633
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Carnac(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Carnac(seed):
0.093
Sensitivity
CentroidFold:
0.515
Carnac(seed):
0.010
Positive Predictive Value
CentroidFold:
0.633
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNASampler(seed):
0.088
Sensitivity
CentroidFold:
0.515
RNASampler(seed):
0.009
Positive Predictive Value
CentroidFold:
0.633
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Mastr(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Mastr(seed):
0.072
Sensitivity
CentroidFold:
0.515
Mastr(seed):
0.006
Positive Predictive Value
CentroidFold:
0.633
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs NanoFolder
Matthews Correlation Coefficient
CentroidFold:
0.571
NanoFolder:
0.068
Sensitivity
CentroidFold:
0.515
NanoFolder:
0.027
Positive Predictive Value
CentroidFold:
0.633
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidFold vs Multilign(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Multilign(seed):
0.059
Sensitivity
CentroidFold:
0.515
Multilign(seed):
0.005
Positive Predictive Value
CentroidFold:
0.633
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidAlifold(20) |
1242
ContextFold vs CentroidAlifold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidAlifold(20):
0.573
Sensitivity
ContextFold:
0.717
CentroidAlifold(20):
0.369
Positive Predictive Value
ContextFold:
0.794
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidAlifold(20):
0.573
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidAlifold(20):
0.369
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs CentroidAlifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CentroidAlifold(20):
0.573
Sensitivity
CentroidAlifold(seed):
0.414
CentroidAlifold(20):
0.369
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs CentroidAlifold(20)
Matthews Correlation Coefficient
IPknot:
0.607
CentroidAlifold(20):
0.573
Sensitivity
IPknot:
0.551
CentroidAlifold(20):
0.369
Positive Predictive Value
IPknot:
0.670
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs CentroidAlifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CentroidAlifold(20):
0.573
Sensitivity
PETfold_pre2.0(20):
0.410
CentroidAlifold(20):
0.369
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidAlifold(20):
0.891
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs CentroidFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CentroidFold:
0.571
Sensitivity
CentroidAlifold(20):
0.369
CentroidFold:
0.515
Positive Predictive Value
CentroidAlifold(20):
0.891
CentroidFold:
0.633
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.0167317061725
|
|
+
CentroidAlifold(20) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CentroidHomfold‑LAST:
0.560
Sensitivity
CentroidAlifold(20):
0.369
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
CentroidAlifold(20):
0.891
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.83792096436e-08
|
+
CentroidAlifold(20) vs Contrafold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Contrafold:
0.556
Sensitivity
CentroidAlifold(20):
0.369
Contrafold:
0.542
Positive Predictive Value
CentroidAlifold(20):
0.891
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MXScarna(seed):
0.536
Sensitivity
CentroidAlifold(20):
0.369
MXScarna(seed):
0.382
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAalifold(20):
0.532
Sensitivity
CentroidAlifold(20):
0.369
RNAalifold(20):
0.359
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Sfold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Sfold:
0.527
Sensitivity
CentroidAlifold(20):
0.369
Sfold:
0.477
Positive Predictive Value
CentroidAlifold(20):
0.891
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAalifold(seed):
0.523
Sensitivity
CentroidAlifold(20):
0.369
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MaxExpect
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MaxExpect:
0.516
Sensitivity
CentroidAlifold(20):
0.369
MaxExpect:
0.505
Positive Predictive Value
CentroidAlifold(20):
0.891
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs ProbKnot
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
ProbKnot:
0.512
Sensitivity
CentroidAlifold(20):
0.369
ProbKnot:
0.508
Positive Predictive Value
CentroidAlifold(20):
0.891
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MXScarna(20):
0.500
Sensitivity
CentroidAlifold(20):
0.369
MXScarna(20):
0.353
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs UNAFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
UNAFold:
0.494
Sensitivity
CentroidAlifold(20):
0.369
UNAFold:
0.491
Positive Predictive Value
CentroidAlifold(20):
0.891
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Fold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Fold:
0.493
Sensitivity
CentroidAlifold(20):
0.369
Fold:
0.496
Positive Predictive Value
CentroidAlifold(20):
0.891
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAfold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAfold:
0.485
Sensitivity
CentroidAlifold(20):
0.369
RNAfold:
0.486
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
PknotsRG:
0.474
Sensitivity
CentroidAlifold(20):
0.369
PknotsRG:
0.440
Positive Predictive Value
CentroidAlifold(20):
0.891
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CRWrnafold:
0.430
Sensitivity
CentroidAlifold(20):
0.369
CRWrnafold:
0.379
Positive Predictive Value
CentroidAlifold(20):
0.891
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAsubopt:
0.422
Sensitivity
CentroidAlifold(20):
0.369
RNAsubopt:
0.344
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
TurboFold(20):
0.419
Sensitivity
CentroidAlifold(20):
0.369
TurboFold(20):
0.210
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
PPfold(20):
0.416
Sensitivity
CentroidAlifold(20):
0.369
PPfold(20):
0.197
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Afold:
0.408
Sensitivity
CentroidAlifold(20):
0.369
Afold:
0.343
Positive Predictive Value
CentroidAlifold(20):
0.891
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Carnac(20):
0.401
Sensitivity
CentroidAlifold(20):
0.369
Carnac(20):
0.180
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
McQFold:
0.389
Sensitivity
CentroidAlifold(20):
0.369
McQFold:
0.308
Positive Predictive Value
CentroidAlifold(20):
0.891
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RSpredict(20):
0.377
Sensitivity
CentroidAlifold(20):
0.369
RSpredict(20):
0.241
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAshapes:
0.367
Sensitivity
CentroidAlifold(20):
0.369
RNAshapes:
0.285
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNASLOpt:
0.364
Sensitivity
CentroidAlifold(20):
0.369
RNASLOpt:
0.261
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Multilign(20):
0.350
Sensitivity
CentroidAlifold(20):
0.369
Multilign(20):
0.161
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Murlet(20):
0.321
Sensitivity
CentroidAlifold(20):
0.369
Murlet(20):
0.131
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAwolf:
0.310
Sensitivity
CentroidAlifold(20):
0.369
RNAwolf:
0.261
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RSpredict(seed):
0.295
Sensitivity
CentroidAlifold(20):
0.369
RSpredict(seed):
0.146
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CMfinder(20):
0.292
Sensitivity
CentroidAlifold(20):
0.369
CMfinder(20):
0.107
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Vsfold4:
0.270
Sensitivity
CentroidAlifold(20):
0.369
Vsfold4:
0.198
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Vsfold5:
0.260
Sensitivity
CentroidAlifold(20):
0.369
Vsfold5:
0.195
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNASampler(20):
0.246
Sensitivity
CentroidAlifold(20):
0.369
RNASampler(20):
0.069
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
HotKnots:
0.212
Sensitivity
CentroidAlifold(20):
0.369
HotKnots:
0.069
Positive Predictive Value
CentroidAlifold(20):
0.891
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Mastr(20):
0.187
Sensitivity
CentroidAlifold(20):
0.369
Mastr(20):
0.044
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Cylofold:
0.178
Sensitivity
CentroidAlifold(20):
0.369
Cylofold:
0.053
Positive Predictive Value
CentroidAlifold(20):
0.891
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Pknots:
0.165
Sensitivity
CentroidAlifold(20):
0.369
Pknots:
0.041
Positive Predictive Value
CentroidAlifold(20):
0.891
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Alterna:
0.145
Sensitivity
CentroidAlifold(20):
0.369
Alterna:
0.036
Positive Predictive Value
CentroidAlifold(20):
0.891
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MCFold:
0.138
Sensitivity
CentroidAlifold(20):
0.369
MCFold:
0.044
Positive Predictive Value
CentroidAlifold(20):
0.891
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RDfolder:
0.138
Sensitivity
CentroidAlifold(20):
0.369
RDfolder:
0.031
Positive Predictive Value
CentroidAlifold(20):
0.891
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Murlet(seed):
0.110
Sensitivity
CentroidAlifold(20):
0.369
Murlet(seed):
0.015
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CMfinder(seed):
0.111
Sensitivity
CentroidAlifold(20):
0.369
CMfinder(seed):
0.016
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
TurboFold(seed):
0.106
Sensitivity
CentroidAlifold(20):
0.369
TurboFold(seed):
0.014
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
PPfold(seed):
0.097
Sensitivity
CentroidAlifold(20):
0.369
PPfold(seed):
0.012
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Carnac(seed):
0.093
Sensitivity
CentroidAlifold(20):
0.369
Carnac(seed):
0.010
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNASampler(seed):
0.088
Sensitivity
CentroidAlifold(20):
0.369
RNASampler(seed):
0.009
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Mastr(seed):
0.072
Sensitivity
CentroidAlifold(20):
0.369
Mastr(seed):
0.006
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
NanoFolder:
0.068
Sensitivity
CentroidAlifold(20):
0.369
NanoFolder:
0.027
Positive Predictive Value
CentroidAlifold(20):
0.891
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidAlifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Multilign(seed):
0.059
Sensitivity
CentroidAlifold(20):
0.369
Multilign(seed):
0.005
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CentroidHomfold‑LAST |
1242
ContextFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
ContextFold:
0.754
CentroidHomfold‑LAST:
0.560
Sensitivity
ContextFold:
0.717
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
ContextFold:
0.794
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CentroidHomfold‑LAST:
0.560
Sensitivity
PETfold_pre2.0(seed):
0.667
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CentroidHomfold‑LAST:
0.560
Sensitivity
CentroidAlifold(seed):
0.414
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
CentroidAlifold(seed):
0.901
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
IPknot:
0.607
CentroidHomfold‑LAST:
0.560
Sensitivity
IPknot:
0.551
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
IPknot:
0.670
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CentroidHomfold‑LAST:
0.560
Sensitivity
PETfold_pre2.0(20):
0.410
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidFold:
0.571
CentroidHomfold‑LAST:
0.560
Sensitivity
CentroidFold:
0.515
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
CentroidFold:
0.633
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs CentroidHomfold‑LAST
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CentroidHomfold‑LAST:
0.560
Sensitivity
CentroidAlifold(20):
0.369
CentroidHomfold‑LAST:
0.359
Positive Predictive Value
CentroidAlifold(20):
0.891
CentroidHomfold‑LAST:
0.874
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.83792096436e-08
|
|
+
CentroidHomfold‑LAST vs Contrafold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Contrafold:
0.556
Sensitivity
CentroidHomfold‑LAST:
0.359
Contrafold:
0.542
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 9.94962292175e-07
|
+
CentroidHomfold‑LAST vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MXScarna(seed):
0.536
Sensitivity
CentroidHomfold‑LAST:
0.359
MXScarna(seed):
0.382
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAalifold(20):
0.532
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAalifold(20):
0.359
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Sfold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Sfold:
0.527
Sensitivity
CentroidHomfold‑LAST:
0.359
Sfold:
0.477
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAalifold(seed):
0.523
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs MaxExpect
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MaxExpect:
0.516
Sensitivity
CentroidHomfold‑LAST:
0.359
MaxExpect:
0.505
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs ProbKnot
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
ProbKnot:
0.512
Sensitivity
CentroidHomfold‑LAST:
0.359
ProbKnot:
0.508
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs MXScarna(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MXScarna(20):
0.500
Sensitivity
CentroidHomfold‑LAST:
0.359
MXScarna(20):
0.353
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs UNAFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
UNAFold:
0.494
Sensitivity
CentroidHomfold‑LAST:
0.359
UNAFold:
0.491
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Fold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Fold:
0.493
Sensitivity
CentroidHomfold‑LAST:
0.359
Fold:
0.496
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAfold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAfold:
0.485
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAfold:
0.486
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs PknotsRG
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
PknotsRG:
0.474
Sensitivity
CentroidHomfold‑LAST:
0.359
PknotsRG:
0.440
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CRWrnafold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
CRWrnafold:
0.430
Sensitivity
CentroidHomfold‑LAST:
0.359
CRWrnafold:
0.379
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAsubopt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAsubopt:
0.422
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAsubopt:
0.344
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs TurboFold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
TurboFold(20):
0.419
Sensitivity
CentroidHomfold‑LAST:
0.359
TurboFold(20):
0.210
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs PPfold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
PPfold(20):
0.416
Sensitivity
CentroidHomfold‑LAST:
0.359
PPfold(20):
0.197
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Afold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Afold:
0.408
Sensitivity
CentroidHomfold‑LAST:
0.359
Afold:
0.343
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Carnac(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Carnac(20):
0.401
Sensitivity
CentroidHomfold‑LAST:
0.359
Carnac(20):
0.180
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs McQFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
McQFold:
0.389
Sensitivity
CentroidHomfold‑LAST:
0.359
McQFold:
0.308
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RSpredict(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RSpredict(20):
0.377
Sensitivity
CentroidHomfold‑LAST:
0.359
RSpredict(20):
0.241
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAshapes
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAshapes:
0.367
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAshapes:
0.285
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNASLOpt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNASLOpt:
0.364
Sensitivity
CentroidHomfold‑LAST:
0.359
RNASLOpt:
0.261
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Multilign(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Multilign(20):
0.350
Sensitivity
CentroidHomfold‑LAST:
0.359
Multilign(20):
0.161
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Murlet(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Murlet(20):
0.321
Sensitivity
CentroidHomfold‑LAST:
0.359
Murlet(20):
0.131
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNAwolf
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAwolf:
0.310
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAwolf:
0.261
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RSpredict(seed):
0.295
Sensitivity
CentroidHomfold‑LAST:
0.359
RSpredict(seed):
0.146
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CMfinder(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
CMfinder(20):
0.292
Sensitivity
CentroidHomfold‑LAST:
0.359
CMfinder(20):
0.107
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Vsfold4
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Vsfold4:
0.270
Sensitivity
CentroidHomfold‑LAST:
0.359
Vsfold4:
0.198
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Vsfold5
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Vsfold5:
0.260
Sensitivity
CentroidHomfold‑LAST:
0.359
Vsfold5:
0.195
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNASampler(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNASampler(20):
0.246
Sensitivity
CentroidHomfold‑LAST:
0.359
RNASampler(20):
0.069
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs HotKnots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
HotKnots:
0.212
Sensitivity
CentroidHomfold‑LAST:
0.359
HotKnots:
0.069
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Mastr(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Mastr(20):
0.187
Sensitivity
CentroidHomfold‑LAST:
0.359
Mastr(20):
0.044
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Cylofold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Cylofold:
0.178
Sensitivity
CentroidHomfold‑LAST:
0.359
Cylofold:
0.053
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Pknots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Pknots:
0.165
Sensitivity
CentroidHomfold‑LAST:
0.359
Pknots:
0.041
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Alterna
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Alterna:
0.145
Sensitivity
CentroidHomfold‑LAST:
0.359
Alterna:
0.036
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs MCFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MCFold:
0.138
Sensitivity
CentroidHomfold‑LAST:
0.359
MCFold:
0.044
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RDfolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RDfolder:
0.138
Sensitivity
CentroidHomfold‑LAST:
0.359
RDfolder:
0.031
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Murlet(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Murlet(seed):
0.110
Sensitivity
CentroidHomfold‑LAST:
0.359
Murlet(seed):
0.015
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
CMfinder(seed):
0.111
Sensitivity
CentroidHomfold‑LAST:
0.359
CMfinder(seed):
0.016
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
TurboFold(seed):
0.106
Sensitivity
CentroidHomfold‑LAST:
0.359
TurboFold(seed):
0.014
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs PPfold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
PPfold(seed):
0.097
Sensitivity
CentroidHomfold‑LAST:
0.359
PPfold(seed):
0.012
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Carnac(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Carnac(seed):
0.093
Sensitivity
CentroidHomfold‑LAST:
0.359
Carnac(seed):
0.010
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNASampler(seed):
0.088
Sensitivity
CentroidHomfold‑LAST:
0.359
RNASampler(seed):
0.009
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Mastr(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Mastr(seed):
0.072
Sensitivity
CentroidHomfold‑LAST:
0.359
Mastr(seed):
0.006
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs NanoFolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
NanoFolder:
0.068
Sensitivity
CentroidHomfold‑LAST:
0.359
NanoFolder:
0.027
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CentroidHomfold‑LAST vs Multilign(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Multilign(seed):
0.059
Sensitivity
CentroidHomfold‑LAST:
0.359
Multilign(seed):
0.005
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Contrafold |
1242
ContextFold vs Contrafold
Matthews Correlation Coefficient
ContextFold:
0.754
Contrafold:
0.556
Sensitivity
ContextFold:
0.717
Contrafold:
0.542
Positive Predictive Value
ContextFold:
0.794
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Contrafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Contrafold:
0.556
Sensitivity
PETfold_pre2.0(seed):
0.667
Contrafold:
0.542
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Contrafold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Contrafold:
0.556
Sensitivity
CentroidAlifold(seed):
0.414
Contrafold:
0.542
Positive Predictive Value
CentroidAlifold(seed):
0.901
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Contrafold
Matthews Correlation Coefficient
IPknot:
0.607
Contrafold:
0.556
Sensitivity
IPknot:
0.551
Contrafold:
0.542
Positive Predictive Value
IPknot:
0.670
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Contrafold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Contrafold:
0.556
Sensitivity
PETfold_pre2.0(20):
0.410
Contrafold:
0.542
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Contrafold
Matthews Correlation Coefficient
CentroidFold:
0.571
Contrafold:
0.556
Sensitivity
CentroidFold:
0.515
Contrafold:
0.542
Positive Predictive Value
CentroidFold:
0.633
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Contrafold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Contrafold:
0.556
Sensitivity
CentroidAlifold(20):
0.369
Contrafold:
0.542
Positive Predictive Value
CentroidAlifold(20):
0.891
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Contrafold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Contrafold:
0.556
Sensitivity
CentroidHomfold‑LAST:
0.359
Contrafold:
0.542
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Contrafold:
0.571
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 9.94962292175e-07
|
|
+
Contrafold vs MXScarna(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
MXScarna(seed):
0.536
Sensitivity
Contrafold:
0.542
MXScarna(seed):
0.382
Positive Predictive Value
Contrafold:
0.571
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAalifold(20)
Matthews Correlation Coefficient
Contrafold:
0.556
RNAalifold(20):
0.532
Sensitivity
Contrafold:
0.542
RNAalifold(20):
0.359
Positive Predictive Value
Contrafold:
0.571
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Sfold
Matthews Correlation Coefficient
Contrafold:
0.556
Sfold:
0.527
Sensitivity
Contrafold:
0.542
Sfold:
0.477
Positive Predictive Value
Contrafold:
0.571
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAalifold(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
RNAalifold(seed):
0.523
Sensitivity
Contrafold:
0.542
RNAalifold(seed):
0.328
Positive Predictive Value
Contrafold:
0.571
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MaxExpect
Matthews Correlation Coefficient
Contrafold:
0.556
MaxExpect:
0.516
Sensitivity
Contrafold:
0.542
MaxExpect:
0.505
Positive Predictive Value
Contrafold:
0.571
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs ProbKnot
Matthews Correlation Coefficient
Contrafold:
0.556
ProbKnot:
0.512
Sensitivity
Contrafold:
0.542
ProbKnot:
0.508
Positive Predictive Value
Contrafold:
0.571
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MXScarna(20)
Matthews Correlation Coefficient
Contrafold:
0.556
MXScarna(20):
0.500
Sensitivity
Contrafold:
0.542
MXScarna(20):
0.353
Positive Predictive Value
Contrafold:
0.571
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs UNAFold
Matthews Correlation Coefficient
Contrafold:
0.556
UNAFold:
0.494
Sensitivity
Contrafold:
0.542
UNAFold:
0.491
Positive Predictive Value
Contrafold:
0.571
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Fold
Matthews Correlation Coefficient
Contrafold:
0.556
Fold:
0.493
Sensitivity
Contrafold:
0.542
Fold:
0.496
Positive Predictive Value
Contrafold:
0.571
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAfold
Matthews Correlation Coefficient
Contrafold:
0.556
RNAfold:
0.485
Sensitivity
Contrafold:
0.542
RNAfold:
0.486
Positive Predictive Value
Contrafold:
0.571
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PknotsRG
Matthews Correlation Coefficient
Contrafold:
0.556
PknotsRG:
0.474
Sensitivity
Contrafold:
0.542
PknotsRG:
0.440
Positive Predictive Value
Contrafold:
0.571
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CRWrnafold
Matthews Correlation Coefficient
Contrafold:
0.556
CRWrnafold:
0.430
Sensitivity
Contrafold:
0.542
CRWrnafold:
0.379
Positive Predictive Value
Contrafold:
0.571
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAsubopt
Matthews Correlation Coefficient
Contrafold:
0.556
RNAsubopt:
0.422
Sensitivity
Contrafold:
0.542
RNAsubopt:
0.344
Positive Predictive Value
Contrafold:
0.571
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs TurboFold(20)
Matthews Correlation Coefficient
Contrafold:
0.556
TurboFold(20):
0.419
Sensitivity
Contrafold:
0.542
TurboFold(20):
0.210
Positive Predictive Value
Contrafold:
0.571
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PPfold(20)
Matthews Correlation Coefficient
Contrafold:
0.556
PPfold(20):
0.416
Sensitivity
Contrafold:
0.542
PPfold(20):
0.197
Positive Predictive Value
Contrafold:
0.571
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Afold
Matthews Correlation Coefficient
Contrafold:
0.556
Afold:
0.408
Sensitivity
Contrafold:
0.542
Afold:
0.343
Positive Predictive Value
Contrafold:
0.571
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Carnac(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Carnac(20):
0.401
Sensitivity
Contrafold:
0.542
Carnac(20):
0.180
Positive Predictive Value
Contrafold:
0.571
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs McQFold
Matthews Correlation Coefficient
Contrafold:
0.556
McQFold:
0.389
Sensitivity
Contrafold:
0.542
McQFold:
0.308
Positive Predictive Value
Contrafold:
0.571
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RSpredict(20)
Matthews Correlation Coefficient
Contrafold:
0.556
RSpredict(20):
0.377
Sensitivity
Contrafold:
0.542
RSpredict(20):
0.241
Positive Predictive Value
Contrafold:
0.571
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAshapes
Matthews Correlation Coefficient
Contrafold:
0.556
RNAshapes:
0.367
Sensitivity
Contrafold:
0.542
RNAshapes:
0.285
Positive Predictive Value
Contrafold:
0.571
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNASLOpt
Matthews Correlation Coefficient
Contrafold:
0.556
RNASLOpt:
0.364
Sensitivity
Contrafold:
0.542
RNASLOpt:
0.261
Positive Predictive Value
Contrafold:
0.571
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Multilign(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Multilign(20):
0.350
Sensitivity
Contrafold:
0.542
Multilign(20):
0.161
Positive Predictive Value
Contrafold:
0.571
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Murlet(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Murlet(20):
0.321
Sensitivity
Contrafold:
0.542
Murlet(20):
0.131
Positive Predictive Value
Contrafold:
0.571
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNAwolf
Matthews Correlation Coefficient
Contrafold:
0.556
RNAwolf:
0.310
Sensitivity
Contrafold:
0.542
RNAwolf:
0.261
Positive Predictive Value
Contrafold:
0.571
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RSpredict(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
RSpredict(seed):
0.295
Sensitivity
Contrafold:
0.542
RSpredict(seed):
0.146
Positive Predictive Value
Contrafold:
0.571
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CMfinder(20)
Matthews Correlation Coefficient
Contrafold:
0.556
CMfinder(20):
0.292
Sensitivity
Contrafold:
0.542
CMfinder(20):
0.107
Positive Predictive Value
Contrafold:
0.571
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Vsfold4
Matthews Correlation Coefficient
Contrafold:
0.556
Vsfold4:
0.270
Sensitivity
Contrafold:
0.542
Vsfold4:
0.198
Positive Predictive Value
Contrafold:
0.571
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Vsfold5
Matthews Correlation Coefficient
Contrafold:
0.556
Vsfold5:
0.260
Sensitivity
Contrafold:
0.542
Vsfold5:
0.195
Positive Predictive Value
Contrafold:
0.571
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNASampler(20)
Matthews Correlation Coefficient
Contrafold:
0.556
RNASampler(20):
0.246
Sensitivity
Contrafold:
0.542
RNASampler(20):
0.069
Positive Predictive Value
Contrafold:
0.571
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs HotKnots
Matthews Correlation Coefficient
Contrafold:
0.556
HotKnots:
0.212
Sensitivity
Contrafold:
0.542
HotKnots:
0.069
Positive Predictive Value
Contrafold:
0.571
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Mastr(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Mastr(20):
0.187
Sensitivity
Contrafold:
0.542
Mastr(20):
0.044
Positive Predictive Value
Contrafold:
0.571
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Cylofold
Matthews Correlation Coefficient
Contrafold:
0.556
Cylofold:
0.178
Sensitivity
Contrafold:
0.542
Cylofold:
0.053
Positive Predictive Value
Contrafold:
0.571
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Pknots
Matthews Correlation Coefficient
Contrafold:
0.556
Pknots:
0.165
Sensitivity
Contrafold:
0.542
Pknots:
0.041
Positive Predictive Value
Contrafold:
0.571
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Alterna
Matthews Correlation Coefficient
Contrafold:
0.556
Alterna:
0.145
Sensitivity
Contrafold:
0.542
Alterna:
0.036
Positive Predictive Value
Contrafold:
0.571
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs MCFold
Matthews Correlation Coefficient
Contrafold:
0.556
MCFold:
0.138
Sensitivity
Contrafold:
0.542
MCFold:
0.044
Positive Predictive Value
Contrafold:
0.571
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RDfolder
Matthews Correlation Coefficient
Contrafold:
0.556
RDfolder:
0.138
Sensitivity
Contrafold:
0.542
RDfolder:
0.031
Positive Predictive Value
Contrafold:
0.571
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Murlet(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Murlet(seed):
0.110
Sensitivity
Contrafold:
0.542
Murlet(seed):
0.015
Positive Predictive Value
Contrafold:
0.571
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs CMfinder(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
CMfinder(seed):
0.111
Sensitivity
Contrafold:
0.542
CMfinder(seed):
0.016
Positive Predictive Value
Contrafold:
0.571
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs TurboFold(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
TurboFold(seed):
0.106
Sensitivity
Contrafold:
0.542
TurboFold(seed):
0.014
Positive Predictive Value
Contrafold:
0.571
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs PPfold(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
PPfold(seed):
0.097
Sensitivity
Contrafold:
0.542
PPfold(seed):
0.012
Positive Predictive Value
Contrafold:
0.571
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Carnac(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Carnac(seed):
0.093
Sensitivity
Contrafold:
0.542
Carnac(seed):
0.010
Positive Predictive Value
Contrafold:
0.571
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs RNASampler(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
RNASampler(seed):
0.088
Sensitivity
Contrafold:
0.542
RNASampler(seed):
0.009
Positive Predictive Value
Contrafold:
0.571
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Mastr(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Mastr(seed):
0.072
Sensitivity
Contrafold:
0.542
Mastr(seed):
0.006
Positive Predictive Value
Contrafold:
0.571
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs NanoFolder
Matthews Correlation Coefficient
Contrafold:
0.556
NanoFolder:
0.068
Sensitivity
Contrafold:
0.542
NanoFolder:
0.027
Positive Predictive Value
Contrafold:
0.571
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Contrafold vs Multilign(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Multilign(seed):
0.059
Sensitivity
Contrafold:
0.542
Multilign(seed):
0.005
Positive Predictive Value
Contrafold:
0.571
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MXScarna(seed) |
1242
ContextFold vs MXScarna(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
MXScarna(seed):
0.536
Sensitivity
ContextFold:
0.717
MXScarna(seed):
0.382
Positive Predictive Value
ContextFold:
0.794
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MXScarna(seed):
0.536
Sensitivity
PETfold_pre2.0(seed):
0.667
MXScarna(seed):
0.382
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MXScarna(seed):
0.536
Sensitivity
CentroidAlifold(seed):
0.414
MXScarna(seed):
0.382
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs MXScarna(seed)
Matthews Correlation Coefficient
IPknot:
0.607
MXScarna(seed):
0.536
Sensitivity
IPknot:
0.551
MXScarna(seed):
0.382
Positive Predictive Value
IPknot:
0.670
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs MXScarna(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MXScarna(seed):
0.536
Sensitivity
PETfold_pre2.0(20):
0.410
MXScarna(seed):
0.382
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
MXScarna(seed):
0.536
Sensitivity
CentroidFold:
0.515
MXScarna(seed):
0.382
Positive Predictive Value
CentroidFold:
0.633
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MXScarna(seed):
0.536
Sensitivity
CentroidAlifold(20):
0.369
MXScarna(seed):
0.382
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs MXScarna(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MXScarna(seed):
0.536
Sensitivity
CentroidHomfold‑LAST:
0.359
MXScarna(seed):
0.382
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs MXScarna(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
MXScarna(seed):
0.536
Sensitivity
Contrafold:
0.542
MXScarna(seed):
0.382
Positive Predictive Value
Contrafold:
0.571
MXScarna(seed):
0.753
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
MXScarna(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAalifold(20):
0.532
Sensitivity
MXScarna(seed):
0.382
RNAalifold(20):
0.359
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.1569883686e-08
|
+
MXScarna(seed) vs Sfold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Sfold:
0.527
Sensitivity
MXScarna(seed):
0.382
Sfold:
0.477
Positive Predictive Value
MXScarna(seed):
0.753
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAalifold(seed):
0.523
Sensitivity
MXScarna(seed):
0.382
RNAalifold(seed):
0.328
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs MaxExpect
Matthews Correlation Coefficient
MXScarna(seed):
0.536
MaxExpect:
0.516
Sensitivity
MXScarna(seed):
0.382
MaxExpect:
0.505
Positive Predictive Value
MXScarna(seed):
0.753
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs ProbKnot
Matthews Correlation Coefficient
MXScarna(seed):
0.536
ProbKnot:
0.512
Sensitivity
MXScarna(seed):
0.382
ProbKnot:
0.508
Positive Predictive Value
MXScarna(seed):
0.753
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs MXScarna(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
MXScarna(20):
0.500
Sensitivity
MXScarna(seed):
0.382
MXScarna(20):
0.353
Positive Predictive Value
MXScarna(seed):
0.753
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs UNAFold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
UNAFold:
0.494
Sensitivity
MXScarna(seed):
0.382
UNAFold:
0.491
Positive Predictive Value
MXScarna(seed):
0.753
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Fold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Fold:
0.493
Sensitivity
MXScarna(seed):
0.382
Fold:
0.496
Positive Predictive Value
MXScarna(seed):
0.753
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAfold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAfold:
0.485
Sensitivity
MXScarna(seed):
0.382
RNAfold:
0.486
Positive Predictive Value
MXScarna(seed):
0.753
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs PknotsRG
Matthews Correlation Coefficient
MXScarna(seed):
0.536
PknotsRG:
0.474
Sensitivity
MXScarna(seed):
0.382
PknotsRG:
0.440
Positive Predictive Value
MXScarna(seed):
0.753
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
CRWrnafold:
0.430
Sensitivity
MXScarna(seed):
0.382
CRWrnafold:
0.379
Positive Predictive Value
MXScarna(seed):
0.753
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAsubopt:
0.422
Sensitivity
MXScarna(seed):
0.382
RNAsubopt:
0.344
Positive Predictive Value
MXScarna(seed):
0.753
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
TurboFold(20):
0.419
Sensitivity
MXScarna(seed):
0.382
TurboFold(20):
0.210
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
PPfold(20):
0.416
Sensitivity
MXScarna(seed):
0.382
PPfold(20):
0.197
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Afold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Afold:
0.408
Sensitivity
MXScarna(seed):
0.382
Afold:
0.343
Positive Predictive Value
MXScarna(seed):
0.753
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Carnac(20):
0.401
Sensitivity
MXScarna(seed):
0.382
Carnac(20):
0.180
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs McQFold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
McQFold:
0.389
Sensitivity
MXScarna(seed):
0.382
McQFold:
0.308
Positive Predictive Value
MXScarna(seed):
0.753
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RSpredict(20):
0.377
Sensitivity
MXScarna(seed):
0.382
RSpredict(20):
0.241
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAshapes:
0.367
Sensitivity
MXScarna(seed):
0.382
RNAshapes:
0.285
Positive Predictive Value
MXScarna(seed):
0.753
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNASLOpt:
0.364
Sensitivity
MXScarna(seed):
0.382
RNASLOpt:
0.261
Positive Predictive Value
MXScarna(seed):
0.753
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Multilign(20):
0.350
Sensitivity
MXScarna(seed):
0.382
Multilign(20):
0.161
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Murlet(20):
0.321
Sensitivity
MXScarna(seed):
0.382
Murlet(20):
0.131
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAwolf:
0.310
Sensitivity
MXScarna(seed):
0.382
RNAwolf:
0.261
Positive Predictive Value
MXScarna(seed):
0.753
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RSpredict(seed):
0.295
Sensitivity
MXScarna(seed):
0.382
RSpredict(seed):
0.146
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
CMfinder(20):
0.292
Sensitivity
MXScarna(seed):
0.382
CMfinder(20):
0.107
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Vsfold4:
0.270
Sensitivity
MXScarna(seed):
0.382
Vsfold4:
0.198
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Vsfold5:
0.260
Sensitivity
MXScarna(seed):
0.382
Vsfold5:
0.195
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNASampler(20):
0.246
Sensitivity
MXScarna(seed):
0.382
RNASampler(20):
0.069
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs HotKnots
Matthews Correlation Coefficient
MXScarna(seed):
0.536
HotKnots:
0.212
Sensitivity
MXScarna(seed):
0.382
HotKnots:
0.069
Positive Predictive Value
MXScarna(seed):
0.753
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Mastr(20):
0.187
Sensitivity
MXScarna(seed):
0.382
Mastr(20):
0.044
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Cylofold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Cylofold:
0.178
Sensitivity
MXScarna(seed):
0.382
Cylofold:
0.053
Positive Predictive Value
MXScarna(seed):
0.753
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Pknots
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Pknots:
0.165
Sensitivity
MXScarna(seed):
0.382
Pknots:
0.041
Positive Predictive Value
MXScarna(seed):
0.753
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Alterna
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Alterna:
0.145
Sensitivity
MXScarna(seed):
0.382
Alterna:
0.036
Positive Predictive Value
MXScarna(seed):
0.753
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs MCFold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
MCFold:
0.138
Sensitivity
MXScarna(seed):
0.382
MCFold:
0.044
Positive Predictive Value
MXScarna(seed):
0.753
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RDfolder
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RDfolder:
0.138
Sensitivity
MXScarna(seed):
0.382
RDfolder:
0.031
Positive Predictive Value
MXScarna(seed):
0.753
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Murlet(seed):
0.110
Sensitivity
MXScarna(seed):
0.382
Murlet(seed):
0.015
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
CMfinder(seed):
0.111
Sensitivity
MXScarna(seed):
0.382
CMfinder(seed):
0.016
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
TurboFold(seed):
0.106
Sensitivity
MXScarna(seed):
0.382
TurboFold(seed):
0.014
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
PPfold(seed):
0.097
Sensitivity
MXScarna(seed):
0.382
PPfold(seed):
0.012
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Carnac(seed):
0.093
Sensitivity
MXScarna(seed):
0.382
Carnac(seed):
0.010
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNASampler(seed):
0.088
Sensitivity
MXScarna(seed):
0.382
RNASampler(seed):
0.009
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Mastr(seed):
0.072
Sensitivity
MXScarna(seed):
0.382
Mastr(seed):
0.006
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(seed):
0.536
NanoFolder:
0.068
Sensitivity
MXScarna(seed):
0.382
NanoFolder:
0.027
Positive Predictive Value
MXScarna(seed):
0.753
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(seed) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Multilign(seed):
0.059
Sensitivity
MXScarna(seed):
0.382
Multilign(seed):
0.005
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAalifold(20) |
1242
ContextFold vs RNAalifold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
RNAalifold(20):
0.532
Sensitivity
ContextFold:
0.717
RNAalifold(20):
0.359
Positive Predictive Value
ContextFold:
0.794
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAalifold(20):
0.532
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAalifold(20):
0.359
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAalifold(20):
0.532
Sensitivity
CentroidAlifold(seed):
0.414
RNAalifold(20):
0.359
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNAalifold(20)
Matthews Correlation Coefficient
IPknot:
0.607
RNAalifold(20):
0.532
Sensitivity
IPknot:
0.551
RNAalifold(20):
0.359
Positive Predictive Value
IPknot:
0.670
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNAalifold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAalifold(20):
0.532
Sensitivity
PETfold_pre2.0(20):
0.410
RNAalifold(20):
0.359
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAalifold(20):
0.532
Sensitivity
CentroidFold:
0.515
RNAalifold(20):
0.359
Positive Predictive Value
CentroidFold:
0.633
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAalifold(20):
0.532
Sensitivity
CentroidAlifold(20):
0.369
RNAalifold(20):
0.359
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNAalifold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAalifold(20):
0.532
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAalifold(20):
0.359
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNAalifold(20)
Matthews Correlation Coefficient
Contrafold:
0.556
RNAalifold(20):
0.532
Sensitivity
Contrafold:
0.542
RNAalifold(20):
0.359
Positive Predictive Value
Contrafold:
0.571
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNAalifold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAalifold(20):
0.532
Sensitivity
MXScarna(seed):
0.382
RNAalifold(20):
0.359
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.1569883686e-08
|
|
+
RNAalifold(20) vs Sfold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Sfold:
0.527
Sensitivity
RNAalifold(20):
0.359
Sfold:
0.477
Positive Predictive Value
RNAalifold(20):
0.787
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.14023792096e-06
|
+
RNAalifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAalifold(seed):
0.523
Sensitivity
RNAalifold(20):
0.359
RNAalifold(seed):
0.328
Positive Predictive Value
RNAalifold(20):
0.787
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs MaxExpect
Matthews Correlation Coefficient
RNAalifold(20):
0.532
MaxExpect:
0.516
Sensitivity
RNAalifold(20):
0.359
MaxExpect:
0.505
Positive Predictive Value
RNAalifold(20):
0.787
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs ProbKnot
Matthews Correlation Coefficient
RNAalifold(20):
0.532
ProbKnot:
0.512
Sensitivity
RNAalifold(20):
0.359
ProbKnot:
0.508
Positive Predictive Value
RNAalifold(20):
0.787
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
MXScarna(20):
0.500
Sensitivity
RNAalifold(20):
0.359
MXScarna(20):
0.353
Positive Predictive Value
RNAalifold(20):
0.787
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs UNAFold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
UNAFold:
0.494
Sensitivity
RNAalifold(20):
0.359
UNAFold:
0.491
Positive Predictive Value
RNAalifold(20):
0.787
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Fold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Fold:
0.493
Sensitivity
RNAalifold(20):
0.359
Fold:
0.496
Positive Predictive Value
RNAalifold(20):
0.787
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAfold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAfold:
0.485
Sensitivity
RNAalifold(20):
0.359
RNAfold:
0.486
Positive Predictive Value
RNAalifold(20):
0.787
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs PknotsRG
Matthews Correlation Coefficient
RNAalifold(20):
0.532
PknotsRG:
0.474
Sensitivity
RNAalifold(20):
0.359
PknotsRG:
0.440
Positive Predictive Value
RNAalifold(20):
0.787
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
CRWrnafold:
0.430
Sensitivity
RNAalifold(20):
0.359
CRWrnafold:
0.379
Positive Predictive Value
RNAalifold(20):
0.787
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAsubopt:
0.422
Sensitivity
RNAalifold(20):
0.359
RNAsubopt:
0.344
Positive Predictive Value
RNAalifold(20):
0.787
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
TurboFold(20):
0.419
Sensitivity
RNAalifold(20):
0.359
TurboFold(20):
0.210
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
PPfold(20):
0.416
Sensitivity
RNAalifold(20):
0.359
PPfold(20):
0.197
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Afold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Afold:
0.408
Sensitivity
RNAalifold(20):
0.359
Afold:
0.343
Positive Predictive Value
RNAalifold(20):
0.787
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Carnac(20):
0.401
Sensitivity
RNAalifold(20):
0.359
Carnac(20):
0.180
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs McQFold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
McQFold:
0.389
Sensitivity
RNAalifold(20):
0.359
McQFold:
0.308
Positive Predictive Value
RNAalifold(20):
0.787
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RSpredict(20):
0.377
Sensitivity
RNAalifold(20):
0.359
RSpredict(20):
0.241
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAshapes:
0.367
Sensitivity
RNAalifold(20):
0.359
RNAshapes:
0.285
Positive Predictive Value
RNAalifold(20):
0.787
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNASLOpt:
0.364
Sensitivity
RNAalifold(20):
0.359
RNASLOpt:
0.261
Positive Predictive Value
RNAalifold(20):
0.787
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Multilign(20):
0.350
Sensitivity
RNAalifold(20):
0.359
Multilign(20):
0.161
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Murlet(20):
0.321
Sensitivity
RNAalifold(20):
0.359
Murlet(20):
0.131
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAwolf:
0.310
Sensitivity
RNAalifold(20):
0.359
RNAwolf:
0.261
Positive Predictive Value
RNAalifold(20):
0.787
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RSpredict(seed):
0.295
Sensitivity
RNAalifold(20):
0.359
RSpredict(seed):
0.146
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
CMfinder(20):
0.292
Sensitivity
RNAalifold(20):
0.359
CMfinder(20):
0.107
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Vsfold4:
0.270
Sensitivity
RNAalifold(20):
0.359
Vsfold4:
0.198
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Vsfold5:
0.260
Sensitivity
RNAalifold(20):
0.359
Vsfold5:
0.195
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNASampler(20):
0.246
Sensitivity
RNAalifold(20):
0.359
RNASampler(20):
0.069
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(20):
0.532
HotKnots:
0.212
Sensitivity
RNAalifold(20):
0.359
HotKnots:
0.069
Positive Predictive Value
RNAalifold(20):
0.787
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Mastr(20):
0.187
Sensitivity
RNAalifold(20):
0.359
Mastr(20):
0.044
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Cylofold:
0.178
Sensitivity
RNAalifold(20):
0.359
Cylofold:
0.053
Positive Predictive Value
RNAalifold(20):
0.787
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Pknots
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Pknots:
0.165
Sensitivity
RNAalifold(20):
0.359
Pknots:
0.041
Positive Predictive Value
RNAalifold(20):
0.787
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Alterna
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Alterna:
0.145
Sensitivity
RNAalifold(20):
0.359
Alterna:
0.036
Positive Predictive Value
RNAalifold(20):
0.787
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs MCFold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
MCFold:
0.138
Sensitivity
RNAalifold(20):
0.359
MCFold:
0.044
Positive Predictive Value
RNAalifold(20):
0.787
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RDfolder:
0.138
Sensitivity
RNAalifold(20):
0.359
RDfolder:
0.031
Positive Predictive Value
RNAalifold(20):
0.787
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Murlet(seed):
0.110
Sensitivity
RNAalifold(20):
0.359
Murlet(seed):
0.015
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
CMfinder(seed):
0.111
Sensitivity
RNAalifold(20):
0.359
CMfinder(seed):
0.016
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
TurboFold(seed):
0.106
Sensitivity
RNAalifold(20):
0.359
TurboFold(seed):
0.014
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
PPfold(seed):
0.097
Sensitivity
RNAalifold(20):
0.359
PPfold(seed):
0.012
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Carnac(seed):
0.093
Sensitivity
RNAalifold(20):
0.359
Carnac(seed):
0.010
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNASampler(seed):
0.088
Sensitivity
RNAalifold(20):
0.359
RNASampler(seed):
0.009
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Mastr(seed):
0.072
Sensitivity
RNAalifold(20):
0.359
Mastr(seed):
0.006
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(20):
0.532
NanoFolder:
0.068
Sensitivity
RNAalifold(20):
0.359
NanoFolder:
0.027
Positive Predictive Value
RNAalifold(20):
0.787
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Multilign(seed):
0.059
Sensitivity
RNAalifold(20):
0.359
Multilign(seed):
0.005
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Sfold |
1242
ContextFold vs Sfold
Matthews Correlation Coefficient
ContextFold:
0.754
Sfold:
0.527
Sensitivity
ContextFold:
0.717
Sfold:
0.477
Positive Predictive Value
ContextFold:
0.794
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Sfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Sfold:
0.527
Sensitivity
PETfold_pre2.0(seed):
0.667
Sfold:
0.477
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Sfold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Sfold:
0.527
Sensitivity
CentroidAlifold(seed):
0.414
Sfold:
0.477
Positive Predictive Value
CentroidAlifold(seed):
0.901
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Sfold
Matthews Correlation Coefficient
IPknot:
0.607
Sfold:
0.527
Sensitivity
IPknot:
0.551
Sfold:
0.477
Positive Predictive Value
IPknot:
0.670
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Sfold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Sfold:
0.527
Sensitivity
PETfold_pre2.0(20):
0.410
Sfold:
0.477
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Sfold
Matthews Correlation Coefficient
CentroidFold:
0.571
Sfold:
0.527
Sensitivity
CentroidFold:
0.515
Sfold:
0.477
Positive Predictive Value
CentroidFold:
0.633
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Sfold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Sfold:
0.527
Sensitivity
CentroidAlifold(20):
0.369
Sfold:
0.477
Positive Predictive Value
CentroidAlifold(20):
0.891
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Sfold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Sfold:
0.527
Sensitivity
CentroidHomfold‑LAST:
0.359
Sfold:
0.477
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Sfold
Matthews Correlation Coefficient
Contrafold:
0.556
Sfold:
0.527
Sensitivity
Contrafold:
0.542
Sfold:
0.477
Positive Predictive Value
Contrafold:
0.571
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Sfold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Sfold:
0.527
Sensitivity
MXScarna(seed):
0.382
Sfold:
0.477
Positive Predictive Value
MXScarna(seed):
0.753
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Sfold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Sfold:
0.527
Sensitivity
RNAalifold(20):
0.359
Sfold:
0.477
Positive Predictive Value
RNAalifold(20):
0.787
Sfold:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.14023792096e-06
|
|
+
Sfold vs RNAalifold(seed)
Matthews Correlation Coefficient
Sfold:
0.527
RNAalifold(seed):
0.523
Sensitivity
Sfold:
0.477
RNAalifold(seed):
0.328
Positive Predictive Value
Sfold:
0.582
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.000150377120024
|
+
Sfold vs MaxExpect
Matthews Correlation Coefficient
Sfold:
0.527
MaxExpect:
0.516
Sensitivity
Sfold:
0.477
MaxExpect:
0.505
Positive Predictive Value
Sfold:
0.582
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs ProbKnot
Matthews Correlation Coefficient
Sfold:
0.527
ProbKnot:
0.512
Sensitivity
Sfold:
0.477
ProbKnot:
0.508
Positive Predictive Value
Sfold:
0.582
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs MXScarna(20)
Matthews Correlation Coefficient
Sfold:
0.527
MXScarna(20):
0.500
Sensitivity
Sfold:
0.477
MXScarna(20):
0.353
Positive Predictive Value
Sfold:
0.582
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs UNAFold
Matthews Correlation Coefficient
Sfold:
0.527
UNAFold:
0.494
Sensitivity
Sfold:
0.477
UNAFold:
0.491
Positive Predictive Value
Sfold:
0.582
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Fold
Matthews Correlation Coefficient
Sfold:
0.527
Fold:
0.493
Sensitivity
Sfold:
0.477
Fold:
0.496
Positive Predictive Value
Sfold:
0.582
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAfold
Matthews Correlation Coefficient
Sfold:
0.527
RNAfold:
0.485
Sensitivity
Sfold:
0.477
RNAfold:
0.486
Positive Predictive Value
Sfold:
0.582
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PknotsRG
Matthews Correlation Coefficient
Sfold:
0.527
PknotsRG:
0.474
Sensitivity
Sfold:
0.477
PknotsRG:
0.440
Positive Predictive Value
Sfold:
0.582
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CRWrnafold
Matthews Correlation Coefficient
Sfold:
0.527
CRWrnafold:
0.430
Sensitivity
Sfold:
0.477
CRWrnafold:
0.379
Positive Predictive Value
Sfold:
0.582
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAsubopt
Matthews Correlation Coefficient
Sfold:
0.527
RNAsubopt:
0.422
Sensitivity
Sfold:
0.477
RNAsubopt:
0.344
Positive Predictive Value
Sfold:
0.582
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs TurboFold(20)
Matthews Correlation Coefficient
Sfold:
0.527
TurboFold(20):
0.419
Sensitivity
Sfold:
0.477
TurboFold(20):
0.210
Positive Predictive Value
Sfold:
0.582
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PPfold(20)
Matthews Correlation Coefficient
Sfold:
0.527
PPfold(20):
0.416
Sensitivity
Sfold:
0.477
PPfold(20):
0.197
Positive Predictive Value
Sfold:
0.582
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Afold
Matthews Correlation Coefficient
Sfold:
0.527
Afold:
0.408
Sensitivity
Sfold:
0.477
Afold:
0.343
Positive Predictive Value
Sfold:
0.582
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Carnac(20)
Matthews Correlation Coefficient
Sfold:
0.527
Carnac(20):
0.401
Sensitivity
Sfold:
0.477
Carnac(20):
0.180
Positive Predictive Value
Sfold:
0.582
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs McQFold
Matthews Correlation Coefficient
Sfold:
0.527
McQFold:
0.389
Sensitivity
Sfold:
0.477
McQFold:
0.308
Positive Predictive Value
Sfold:
0.582
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RSpredict(20)
Matthews Correlation Coefficient
Sfold:
0.527
RSpredict(20):
0.377
Sensitivity
Sfold:
0.477
RSpredict(20):
0.241
Positive Predictive Value
Sfold:
0.582
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAshapes
Matthews Correlation Coefficient
Sfold:
0.527
RNAshapes:
0.367
Sensitivity
Sfold:
0.477
RNAshapes:
0.285
Positive Predictive Value
Sfold:
0.582
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNASLOpt
Matthews Correlation Coefficient
Sfold:
0.527
RNASLOpt:
0.364
Sensitivity
Sfold:
0.477
RNASLOpt:
0.261
Positive Predictive Value
Sfold:
0.582
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Multilign(20)
Matthews Correlation Coefficient
Sfold:
0.527
Multilign(20):
0.350
Sensitivity
Sfold:
0.477
Multilign(20):
0.161
Positive Predictive Value
Sfold:
0.582
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Murlet(20)
Matthews Correlation Coefficient
Sfold:
0.527
Murlet(20):
0.321
Sensitivity
Sfold:
0.477
Murlet(20):
0.131
Positive Predictive Value
Sfold:
0.582
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNAwolf
Matthews Correlation Coefficient
Sfold:
0.527
RNAwolf:
0.310
Sensitivity
Sfold:
0.477
RNAwolf:
0.261
Positive Predictive Value
Sfold:
0.582
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RSpredict(seed)
Matthews Correlation Coefficient
Sfold:
0.527
RSpredict(seed):
0.295
Sensitivity
Sfold:
0.477
RSpredict(seed):
0.146
Positive Predictive Value
Sfold:
0.582
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CMfinder(20)
Matthews Correlation Coefficient
Sfold:
0.527
CMfinder(20):
0.292
Sensitivity
Sfold:
0.477
CMfinder(20):
0.107
Positive Predictive Value
Sfold:
0.582
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Vsfold4
Matthews Correlation Coefficient
Sfold:
0.527
Vsfold4:
0.270
Sensitivity
Sfold:
0.477
Vsfold4:
0.198
Positive Predictive Value
Sfold:
0.582
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Vsfold5
Matthews Correlation Coefficient
Sfold:
0.527
Vsfold5:
0.260
Sensitivity
Sfold:
0.477
Vsfold5:
0.195
Positive Predictive Value
Sfold:
0.582
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNASampler(20)
Matthews Correlation Coefficient
Sfold:
0.527
RNASampler(20):
0.246
Sensitivity
Sfold:
0.477
RNASampler(20):
0.069
Positive Predictive Value
Sfold:
0.582
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs HotKnots
Matthews Correlation Coefficient
Sfold:
0.527
HotKnots:
0.212
Sensitivity
Sfold:
0.477
HotKnots:
0.069
Positive Predictive Value
Sfold:
0.582
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Mastr(20)
Matthews Correlation Coefficient
Sfold:
0.527
Mastr(20):
0.187
Sensitivity
Sfold:
0.477
Mastr(20):
0.044
Positive Predictive Value
Sfold:
0.582
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Cylofold
Matthews Correlation Coefficient
Sfold:
0.527
Cylofold:
0.178
Sensitivity
Sfold:
0.477
Cylofold:
0.053
Positive Predictive Value
Sfold:
0.582
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Pknots
Matthews Correlation Coefficient
Sfold:
0.527
Pknots:
0.165
Sensitivity
Sfold:
0.477
Pknots:
0.041
Positive Predictive Value
Sfold:
0.582
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Alterna
Matthews Correlation Coefficient
Sfold:
0.527
Alterna:
0.145
Sensitivity
Sfold:
0.477
Alterna:
0.036
Positive Predictive Value
Sfold:
0.582
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs MCFold
Matthews Correlation Coefficient
Sfold:
0.527
MCFold:
0.138
Sensitivity
Sfold:
0.477
MCFold:
0.044
Positive Predictive Value
Sfold:
0.582
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RDfolder
Matthews Correlation Coefficient
Sfold:
0.527
RDfolder:
0.138
Sensitivity
Sfold:
0.477
RDfolder:
0.031
Positive Predictive Value
Sfold:
0.582
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Murlet(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Murlet(seed):
0.110
Sensitivity
Sfold:
0.477
Murlet(seed):
0.015
Positive Predictive Value
Sfold:
0.582
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs CMfinder(seed)
Matthews Correlation Coefficient
Sfold:
0.527
CMfinder(seed):
0.111
Sensitivity
Sfold:
0.477
CMfinder(seed):
0.016
Positive Predictive Value
Sfold:
0.582
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs TurboFold(seed)
Matthews Correlation Coefficient
Sfold:
0.527
TurboFold(seed):
0.106
Sensitivity
Sfold:
0.477
TurboFold(seed):
0.014
Positive Predictive Value
Sfold:
0.582
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs PPfold(seed)
Matthews Correlation Coefficient
Sfold:
0.527
PPfold(seed):
0.097
Sensitivity
Sfold:
0.477
PPfold(seed):
0.012
Positive Predictive Value
Sfold:
0.582
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Carnac(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Carnac(seed):
0.093
Sensitivity
Sfold:
0.477
Carnac(seed):
0.010
Positive Predictive Value
Sfold:
0.582
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs RNASampler(seed)
Matthews Correlation Coefficient
Sfold:
0.527
RNASampler(seed):
0.088
Sensitivity
Sfold:
0.477
RNASampler(seed):
0.009
Positive Predictive Value
Sfold:
0.582
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Mastr(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Mastr(seed):
0.072
Sensitivity
Sfold:
0.477
Mastr(seed):
0.006
Positive Predictive Value
Sfold:
0.582
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs NanoFolder
Matthews Correlation Coefficient
Sfold:
0.527
NanoFolder:
0.068
Sensitivity
Sfold:
0.477
NanoFolder:
0.027
Positive Predictive Value
Sfold:
0.582
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Sfold vs Multilign(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Multilign(seed):
0.059
Sensitivity
Sfold:
0.477
Multilign(seed):
0.005
Positive Predictive Value
Sfold:
0.582
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAalifold(seed) |
1242
ContextFold vs RNAalifold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
RNAalifold(seed):
0.523
Sensitivity
ContextFold:
0.717
RNAalifold(seed):
0.328
Positive Predictive Value
ContextFold:
0.794
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAalifold(seed):
0.523
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAalifold(seed):
0.328
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAalifold(seed):
0.523
Sensitivity
CentroidAlifold(seed):
0.414
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNAalifold(seed)
Matthews Correlation Coefficient
IPknot:
0.607
RNAalifold(seed):
0.523
Sensitivity
IPknot:
0.551
RNAalifold(seed):
0.328
Positive Predictive Value
IPknot:
0.670
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAalifold(seed):
0.523
Sensitivity
PETfold_pre2.0(20):
0.410
RNAalifold(seed):
0.328
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAalifold(seed):
0.523
Sensitivity
CentroidFold:
0.515
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidFold:
0.633
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAalifold(seed):
0.523
Sensitivity
CentroidAlifold(20):
0.369
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNAalifold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAalifold(seed):
0.523
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAalifold(seed):
0.328
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNAalifold(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
RNAalifold(seed):
0.523
Sensitivity
Contrafold:
0.542
RNAalifold(seed):
0.328
Positive Predictive Value
Contrafold:
0.571
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNAalifold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAalifold(seed):
0.523
Sensitivity
MXScarna(seed):
0.382
RNAalifold(seed):
0.328
Positive Predictive Value
MXScarna(seed):
0.753
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNAalifold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAalifold(seed):
0.523
Sensitivity
RNAalifold(20):
0.359
RNAalifold(seed):
0.328
Positive Predictive Value
RNAalifold(20):
0.787
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNAalifold(seed)
Matthews Correlation Coefficient
Sfold:
0.527
RNAalifold(seed):
0.523
Sensitivity
Sfold:
0.477
RNAalifold(seed):
0.328
Positive Predictive Value
Sfold:
0.582
RNAalifold(seed):
0.835
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.000150377120024
|
|
+
RNAalifold(seed) vs MaxExpect
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
MaxExpect:
0.516
Sensitivity
RNAalifold(seed):
0.328
MaxExpect:
0.505
Positive Predictive Value
RNAalifold(seed):
0.835
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
RNAalifold(seed) vs ProbKnot
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
ProbKnot:
0.512
Sensitivity
RNAalifold(seed):
0.328
ProbKnot:
0.508
Positive Predictive Value
RNAalifold(seed):
0.835
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
MXScarna(20):
0.500
Sensitivity
RNAalifold(seed):
0.328
MXScarna(20):
0.353
Positive Predictive Value
RNAalifold(seed):
0.835
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs UNAFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
UNAFold:
0.494
Sensitivity
RNAalifold(seed):
0.328
UNAFold:
0.491
Positive Predictive Value
RNAalifold(seed):
0.835
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Fold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Fold:
0.493
Sensitivity
RNAalifold(seed):
0.328
Fold:
0.496
Positive Predictive Value
RNAalifold(seed):
0.835
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAfold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAfold:
0.485
Sensitivity
RNAalifold(seed):
0.328
RNAfold:
0.486
Positive Predictive Value
RNAalifold(seed):
0.835
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs PknotsRG
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
PknotsRG:
0.474
Sensitivity
RNAalifold(seed):
0.328
PknotsRG:
0.440
Positive Predictive Value
RNAalifold(seed):
0.835
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
CRWrnafold:
0.430
Sensitivity
RNAalifold(seed):
0.328
CRWrnafold:
0.379
Positive Predictive Value
RNAalifold(seed):
0.835
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAsubopt:
0.422
Sensitivity
RNAalifold(seed):
0.328
RNAsubopt:
0.344
Positive Predictive Value
RNAalifold(seed):
0.835
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
TurboFold(20):
0.419
Sensitivity
RNAalifold(seed):
0.328
TurboFold(20):
0.210
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
PPfold(20):
0.416
Sensitivity
RNAalifold(seed):
0.328
PPfold(20):
0.197
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Afold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Afold:
0.408
Sensitivity
RNAalifold(seed):
0.328
Afold:
0.343
Positive Predictive Value
RNAalifold(seed):
0.835
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Carnac(20):
0.401
Sensitivity
RNAalifold(seed):
0.328
Carnac(20):
0.180
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs McQFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
McQFold:
0.389
Sensitivity
RNAalifold(seed):
0.328
McQFold:
0.308
Positive Predictive Value
RNAalifold(seed):
0.835
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RSpredict(20):
0.377
Sensitivity
RNAalifold(seed):
0.328
RSpredict(20):
0.241
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAshapes:
0.367
Sensitivity
RNAalifold(seed):
0.328
RNAshapes:
0.285
Positive Predictive Value
RNAalifold(seed):
0.835
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNASLOpt:
0.364
Sensitivity
RNAalifold(seed):
0.328
RNASLOpt:
0.261
Positive Predictive Value
RNAalifold(seed):
0.835
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Multilign(20):
0.350
Sensitivity
RNAalifold(seed):
0.328
Multilign(20):
0.161
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Murlet(20):
0.321
Sensitivity
RNAalifold(seed):
0.328
Murlet(20):
0.131
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAwolf:
0.310
Sensitivity
RNAalifold(seed):
0.328
RNAwolf:
0.261
Positive Predictive Value
RNAalifold(seed):
0.835
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RSpredict(seed):
0.295
Sensitivity
RNAalifold(seed):
0.328
RSpredict(seed):
0.146
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
CMfinder(20):
0.292
Sensitivity
RNAalifold(seed):
0.328
CMfinder(20):
0.107
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Vsfold4:
0.270
Sensitivity
RNAalifold(seed):
0.328
Vsfold4:
0.198
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Vsfold5:
0.260
Sensitivity
RNAalifold(seed):
0.328
Vsfold5:
0.195
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNASampler(20):
0.246
Sensitivity
RNAalifold(seed):
0.328
RNASampler(20):
0.069
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
HotKnots:
0.212
Sensitivity
RNAalifold(seed):
0.328
HotKnots:
0.069
Positive Predictive Value
RNAalifold(seed):
0.835
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Mastr(20):
0.187
Sensitivity
RNAalifold(seed):
0.328
Mastr(20):
0.044
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Cylofold:
0.178
Sensitivity
RNAalifold(seed):
0.328
Cylofold:
0.053
Positive Predictive Value
RNAalifold(seed):
0.835
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Pknots
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Pknots:
0.165
Sensitivity
RNAalifold(seed):
0.328
Pknots:
0.041
Positive Predictive Value
RNAalifold(seed):
0.835
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Alterna
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Alterna:
0.145
Sensitivity
RNAalifold(seed):
0.328
Alterna:
0.036
Positive Predictive Value
RNAalifold(seed):
0.835
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs MCFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
MCFold:
0.138
Sensitivity
RNAalifold(seed):
0.328
MCFold:
0.044
Positive Predictive Value
RNAalifold(seed):
0.835
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RDfolder:
0.138
Sensitivity
RNAalifold(seed):
0.328
RDfolder:
0.031
Positive Predictive Value
RNAalifold(seed):
0.835
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Murlet(seed):
0.110
Sensitivity
RNAalifold(seed):
0.328
Murlet(seed):
0.015
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
CMfinder(seed):
0.111
Sensitivity
RNAalifold(seed):
0.328
CMfinder(seed):
0.016
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
TurboFold(seed):
0.106
Sensitivity
RNAalifold(seed):
0.328
TurboFold(seed):
0.014
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
PPfold(seed):
0.097
Sensitivity
RNAalifold(seed):
0.328
PPfold(seed):
0.012
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Carnac(seed):
0.093
Sensitivity
RNAalifold(seed):
0.328
Carnac(seed):
0.010
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNASampler(seed):
0.088
Sensitivity
RNAalifold(seed):
0.328
RNASampler(seed):
0.009
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Mastr(seed):
0.072
Sensitivity
RNAalifold(seed):
0.328
Mastr(seed):
0.006
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
NanoFolder:
0.068
Sensitivity
RNAalifold(seed):
0.328
NanoFolder:
0.027
Positive Predictive Value
RNAalifold(seed):
0.835
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAalifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Multilign(seed):
0.059
Sensitivity
RNAalifold(seed):
0.328
Multilign(seed):
0.005
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MaxExpect |
1242
ContextFold vs MaxExpect
Matthews Correlation Coefficient
ContextFold:
0.754
MaxExpect:
0.516
Sensitivity
ContextFold:
0.717
MaxExpect:
0.505
Positive Predictive Value
ContextFold:
0.794
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs MaxExpect
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MaxExpect:
0.516
Sensitivity
PETfold_pre2.0(seed):
0.667
MaxExpect:
0.505
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs MaxExpect
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MaxExpect:
0.516
Sensitivity
CentroidAlifold(seed):
0.414
MaxExpect:
0.505
Positive Predictive Value
CentroidAlifold(seed):
0.901
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs MaxExpect
Matthews Correlation Coefficient
IPknot:
0.607
MaxExpect:
0.516
Sensitivity
IPknot:
0.551
MaxExpect:
0.505
Positive Predictive Value
IPknot:
0.670
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs MaxExpect
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MaxExpect:
0.516
Sensitivity
PETfold_pre2.0(20):
0.410
MaxExpect:
0.505
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs MaxExpect
Matthews Correlation Coefficient
CentroidFold:
0.571
MaxExpect:
0.516
Sensitivity
CentroidFold:
0.515
MaxExpect:
0.505
Positive Predictive Value
CentroidFold:
0.633
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs MaxExpect
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MaxExpect:
0.516
Sensitivity
CentroidAlifold(20):
0.369
MaxExpect:
0.505
Positive Predictive Value
CentroidAlifold(20):
0.891
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs MaxExpect
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MaxExpect:
0.516
Sensitivity
CentroidHomfold‑LAST:
0.359
MaxExpect:
0.505
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs MaxExpect
Matthews Correlation Coefficient
Contrafold:
0.556
MaxExpect:
0.516
Sensitivity
Contrafold:
0.542
MaxExpect:
0.505
Positive Predictive Value
Contrafold:
0.571
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs MaxExpect
Matthews Correlation Coefficient
MXScarna(seed):
0.536
MaxExpect:
0.516
Sensitivity
MXScarna(seed):
0.382
MaxExpect:
0.505
Positive Predictive Value
MXScarna(seed):
0.753
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs MaxExpect
Matthews Correlation Coefficient
RNAalifold(20):
0.532
MaxExpect:
0.516
Sensitivity
RNAalifold(20):
0.359
MaxExpect:
0.505
Positive Predictive Value
RNAalifold(20):
0.787
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs MaxExpect
Matthews Correlation Coefficient
Sfold:
0.527
MaxExpect:
0.516
Sensitivity
Sfold:
0.477
MaxExpect:
0.505
Positive Predictive Value
Sfold:
0.582
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs MaxExpect
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
MaxExpect:
0.516
Sensitivity
RNAalifold(seed):
0.328
MaxExpect:
0.505
Positive Predictive Value
RNAalifold(seed):
0.835
MaxExpect:
0.528
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
|
+
MaxExpect vs ProbKnot
Matthews Correlation Coefficient
MaxExpect:
0.516
ProbKnot:
0.512
Sensitivity
MaxExpect:
0.505
ProbKnot:
0.508
Positive Predictive Value
MaxExpect:
0.528
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs MXScarna(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
MXScarna(20):
0.500
Sensitivity
MaxExpect:
0.505
MXScarna(20):
0.353
Positive Predictive Value
MaxExpect:
0.528
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs UNAFold
Matthews Correlation Coefficient
MaxExpect:
0.516
UNAFold:
0.494
Sensitivity
MaxExpect:
0.505
UNAFold:
0.491
Positive Predictive Value
MaxExpect:
0.528
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Fold
Matthews Correlation Coefficient
MaxExpect:
0.516
Fold:
0.493
Sensitivity
MaxExpect:
0.505
Fold:
0.496
Positive Predictive Value
MaxExpect:
0.528
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAfold
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAfold:
0.485
Sensitivity
MaxExpect:
0.505
RNAfold:
0.486
Positive Predictive Value
MaxExpect:
0.528
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PknotsRG
Matthews Correlation Coefficient
MaxExpect:
0.516
PknotsRG:
0.474
Sensitivity
MaxExpect:
0.505
PknotsRG:
0.440
Positive Predictive Value
MaxExpect:
0.528
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CRWrnafold
Matthews Correlation Coefficient
MaxExpect:
0.516
CRWrnafold:
0.430
Sensitivity
MaxExpect:
0.505
CRWrnafold:
0.379
Positive Predictive Value
MaxExpect:
0.528
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAsubopt
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAsubopt:
0.422
Sensitivity
MaxExpect:
0.505
RNAsubopt:
0.344
Positive Predictive Value
MaxExpect:
0.528
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs TurboFold(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
TurboFold(20):
0.419
Sensitivity
MaxExpect:
0.505
TurboFold(20):
0.210
Positive Predictive Value
MaxExpect:
0.528
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PPfold(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
PPfold(20):
0.416
Sensitivity
MaxExpect:
0.505
PPfold(20):
0.197
Positive Predictive Value
MaxExpect:
0.528
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Afold
Matthews Correlation Coefficient
MaxExpect:
0.516
Afold:
0.408
Sensitivity
MaxExpect:
0.505
Afold:
0.343
Positive Predictive Value
MaxExpect:
0.528
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Carnac(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Carnac(20):
0.401
Sensitivity
MaxExpect:
0.505
Carnac(20):
0.180
Positive Predictive Value
MaxExpect:
0.528
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs McQFold
Matthews Correlation Coefficient
MaxExpect:
0.516
McQFold:
0.389
Sensitivity
MaxExpect:
0.505
McQFold:
0.308
Positive Predictive Value
MaxExpect:
0.528
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RSpredict(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
RSpredict(20):
0.377
Sensitivity
MaxExpect:
0.505
RSpredict(20):
0.241
Positive Predictive Value
MaxExpect:
0.528
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAshapes
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAshapes:
0.367
Sensitivity
MaxExpect:
0.505
RNAshapes:
0.285
Positive Predictive Value
MaxExpect:
0.528
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNASLOpt
Matthews Correlation Coefficient
MaxExpect:
0.516
RNASLOpt:
0.364
Sensitivity
MaxExpect:
0.505
RNASLOpt:
0.261
Positive Predictive Value
MaxExpect:
0.528
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Multilign(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Multilign(20):
0.350
Sensitivity
MaxExpect:
0.505
Multilign(20):
0.161
Positive Predictive Value
MaxExpect:
0.528
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Murlet(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Murlet(20):
0.321
Sensitivity
MaxExpect:
0.505
Murlet(20):
0.131
Positive Predictive Value
MaxExpect:
0.528
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNAwolf
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAwolf:
0.310
Sensitivity
MaxExpect:
0.505
RNAwolf:
0.261
Positive Predictive Value
MaxExpect:
0.528
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RSpredict(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
RSpredict(seed):
0.295
Sensitivity
MaxExpect:
0.505
RSpredict(seed):
0.146
Positive Predictive Value
MaxExpect:
0.528
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CMfinder(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
CMfinder(20):
0.292
Sensitivity
MaxExpect:
0.505
CMfinder(20):
0.107
Positive Predictive Value
MaxExpect:
0.528
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Vsfold4
Matthews Correlation Coefficient
MaxExpect:
0.516
Vsfold4:
0.270
Sensitivity
MaxExpect:
0.505
Vsfold4:
0.198
Positive Predictive Value
MaxExpect:
0.528
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Vsfold5
Matthews Correlation Coefficient
MaxExpect:
0.516
Vsfold5:
0.260
Sensitivity
MaxExpect:
0.505
Vsfold5:
0.195
Positive Predictive Value
MaxExpect:
0.528
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNASampler(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
RNASampler(20):
0.246
Sensitivity
MaxExpect:
0.505
RNASampler(20):
0.069
Positive Predictive Value
MaxExpect:
0.528
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs HotKnots
Matthews Correlation Coefficient
MaxExpect:
0.516
HotKnots:
0.212
Sensitivity
MaxExpect:
0.505
HotKnots:
0.069
Positive Predictive Value
MaxExpect:
0.528
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Mastr(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Mastr(20):
0.187
Sensitivity
MaxExpect:
0.505
Mastr(20):
0.044
Positive Predictive Value
MaxExpect:
0.528
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Cylofold
Matthews Correlation Coefficient
MaxExpect:
0.516
Cylofold:
0.178
Sensitivity
MaxExpect:
0.505
Cylofold:
0.053
Positive Predictive Value
MaxExpect:
0.528
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Pknots
Matthews Correlation Coefficient
MaxExpect:
0.516
Pknots:
0.165
Sensitivity
MaxExpect:
0.505
Pknots:
0.041
Positive Predictive Value
MaxExpect:
0.528
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Alterna
Matthews Correlation Coefficient
MaxExpect:
0.516
Alterna:
0.145
Sensitivity
MaxExpect:
0.505
Alterna:
0.036
Positive Predictive Value
MaxExpect:
0.528
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs MCFold
Matthews Correlation Coefficient
MaxExpect:
0.516
MCFold:
0.138
Sensitivity
MaxExpect:
0.505
MCFold:
0.044
Positive Predictive Value
MaxExpect:
0.528
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RDfolder
Matthews Correlation Coefficient
MaxExpect:
0.516
RDfolder:
0.138
Sensitivity
MaxExpect:
0.505
RDfolder:
0.031
Positive Predictive Value
MaxExpect:
0.528
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Murlet(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Murlet(seed):
0.110
Sensitivity
MaxExpect:
0.505
Murlet(seed):
0.015
Positive Predictive Value
MaxExpect:
0.528
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs CMfinder(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
CMfinder(seed):
0.111
Sensitivity
MaxExpect:
0.505
CMfinder(seed):
0.016
Positive Predictive Value
MaxExpect:
0.528
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs TurboFold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
TurboFold(seed):
0.106
Sensitivity
MaxExpect:
0.505
TurboFold(seed):
0.014
Positive Predictive Value
MaxExpect:
0.528
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs PPfold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
PPfold(seed):
0.097
Sensitivity
MaxExpect:
0.505
PPfold(seed):
0.012
Positive Predictive Value
MaxExpect:
0.528
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Carnac(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Carnac(seed):
0.093
Sensitivity
MaxExpect:
0.505
Carnac(seed):
0.010
Positive Predictive Value
MaxExpect:
0.528
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs RNASampler(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
RNASampler(seed):
0.088
Sensitivity
MaxExpect:
0.505
RNASampler(seed):
0.009
Positive Predictive Value
MaxExpect:
0.528
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Mastr(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Mastr(seed):
0.072
Sensitivity
MaxExpect:
0.505
Mastr(seed):
0.006
Positive Predictive Value
MaxExpect:
0.528
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs NanoFolder
Matthews Correlation Coefficient
MaxExpect:
0.516
NanoFolder:
0.068
Sensitivity
MaxExpect:
0.505
NanoFolder:
0.027
Positive Predictive Value
MaxExpect:
0.528
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MaxExpect vs Multilign(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Multilign(seed):
0.059
Sensitivity
MaxExpect:
0.505
Multilign(seed):
0.005
Positive Predictive Value
MaxExpect:
0.528
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| ProbKnot |
1242
ContextFold vs ProbKnot
Matthews Correlation Coefficient
ContextFold:
0.754
ProbKnot:
0.512
Sensitivity
ContextFold:
0.717
ProbKnot:
0.508
Positive Predictive Value
ContextFold:
0.794
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs ProbKnot
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
ProbKnot:
0.512
Sensitivity
PETfold_pre2.0(seed):
0.667
ProbKnot:
0.508
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs ProbKnot
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
ProbKnot:
0.512
Sensitivity
CentroidAlifold(seed):
0.414
ProbKnot:
0.508
Positive Predictive Value
CentroidAlifold(seed):
0.901
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs ProbKnot
Matthews Correlation Coefficient
IPknot:
0.607
ProbKnot:
0.512
Sensitivity
IPknot:
0.551
ProbKnot:
0.508
Positive Predictive Value
IPknot:
0.670
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs ProbKnot
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
ProbKnot:
0.512
Sensitivity
PETfold_pre2.0(20):
0.410
ProbKnot:
0.508
Positive Predictive Value
PETfold_pre2.0(20):
0.807
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs ProbKnot
Matthews Correlation Coefficient
CentroidFold:
0.571
ProbKnot:
0.512
Sensitivity
CentroidFold:
0.515
ProbKnot:
0.508
Positive Predictive Value
CentroidFold:
0.633
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs ProbKnot
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
ProbKnot:
0.512
Sensitivity
CentroidAlifold(20):
0.369
ProbKnot:
0.508
Positive Predictive Value
CentroidAlifold(20):
0.891
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs ProbKnot
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
ProbKnot:
0.512
Sensitivity
CentroidHomfold‑LAST:
0.359
ProbKnot:
0.508
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs ProbKnot
Matthews Correlation Coefficient
Contrafold:
0.556
ProbKnot:
0.512
Sensitivity
Contrafold:
0.542
ProbKnot:
0.508
Positive Predictive Value
Contrafold:
0.571
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs ProbKnot
Matthews Correlation Coefficient
MXScarna(seed):
0.536
ProbKnot:
0.512
Sensitivity
MXScarna(seed):
0.382
ProbKnot:
0.508
Positive Predictive Value
MXScarna(seed):
0.753
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs ProbKnot
Matthews Correlation Coefficient
RNAalifold(20):
0.532
ProbKnot:
0.512
Sensitivity
RNAalifold(20):
0.359
ProbKnot:
0.508
Positive Predictive Value
RNAalifold(20):
0.787
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs ProbKnot
Matthews Correlation Coefficient
Sfold:
0.527
ProbKnot:
0.512
Sensitivity
Sfold:
0.477
ProbKnot:
0.508
Positive Predictive Value
Sfold:
0.582
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs ProbKnot
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
ProbKnot:
0.512
Sensitivity
RNAalifold(seed):
0.328
ProbKnot:
0.508
Positive Predictive Value
RNAalifold(seed):
0.835
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs ProbKnot
Matthews Correlation Coefficient
MaxExpect:
0.516
ProbKnot:
0.512
Sensitivity
MaxExpect:
0.505
ProbKnot:
0.508
Positive Predictive Value
MaxExpect:
0.528
ProbKnot:
0.517
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
ProbKnot vs MXScarna(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
MXScarna(20):
0.500
Sensitivity
ProbKnot:
0.508
MXScarna(20):
0.353
Positive Predictive Value
ProbKnot:
0.517
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs UNAFold
Matthews Correlation Coefficient
ProbKnot:
0.512
UNAFold:
0.494
Sensitivity
ProbKnot:
0.508
UNAFold:
0.491
Positive Predictive Value
ProbKnot:
0.517
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Fold
Matthews Correlation Coefficient
ProbKnot:
0.512
Fold:
0.493
Sensitivity
ProbKnot:
0.508
Fold:
0.496
Positive Predictive Value
ProbKnot:
0.517
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAfold
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAfold:
0.485
Sensitivity
ProbKnot:
0.508
RNAfold:
0.486
Positive Predictive Value
ProbKnot:
0.517
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PknotsRG
Matthews Correlation Coefficient
ProbKnot:
0.512
PknotsRG:
0.474
Sensitivity
ProbKnot:
0.508
PknotsRG:
0.440
Positive Predictive Value
ProbKnot:
0.517
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CRWrnafold
Matthews Correlation Coefficient
ProbKnot:
0.512
CRWrnafold:
0.430
Sensitivity
ProbKnot:
0.508
CRWrnafold:
0.379
Positive Predictive Value
ProbKnot:
0.517
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAsubopt
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAsubopt:
0.422
Sensitivity
ProbKnot:
0.508
RNAsubopt:
0.344
Positive Predictive Value
ProbKnot:
0.517
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs TurboFold(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
TurboFold(20):
0.419
Sensitivity
ProbKnot:
0.508
TurboFold(20):
0.210
Positive Predictive Value
ProbKnot:
0.517
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PPfold(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
PPfold(20):
0.416
Sensitivity
ProbKnot:
0.508
PPfold(20):
0.197
Positive Predictive Value
ProbKnot:
0.517
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Afold
Matthews Correlation Coefficient
ProbKnot:
0.512
Afold:
0.408
Sensitivity
ProbKnot:
0.508
Afold:
0.343
Positive Predictive Value
ProbKnot:
0.517
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Carnac(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Carnac(20):
0.401
Sensitivity
ProbKnot:
0.508
Carnac(20):
0.180
Positive Predictive Value
ProbKnot:
0.517
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs McQFold
Matthews Correlation Coefficient
ProbKnot:
0.512
McQFold:
0.389
Sensitivity
ProbKnot:
0.508
McQFold:
0.308
Positive Predictive Value
ProbKnot:
0.517
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RSpredict(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
RSpredict(20):
0.377
Sensitivity
ProbKnot:
0.508
RSpredict(20):
0.241
Positive Predictive Value
ProbKnot:
0.517
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAshapes
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAshapes:
0.367
Sensitivity
ProbKnot:
0.508
RNAshapes:
0.285
Positive Predictive Value
ProbKnot:
0.517
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNASLOpt
Matthews Correlation Coefficient
ProbKnot:
0.512
RNASLOpt:
0.364
Sensitivity
ProbKnot:
0.508
RNASLOpt:
0.261
Positive Predictive Value
ProbKnot:
0.517
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Multilign(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Multilign(20):
0.350
Sensitivity
ProbKnot:
0.508
Multilign(20):
0.161
Positive Predictive Value
ProbKnot:
0.517
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Murlet(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Murlet(20):
0.321
Sensitivity
ProbKnot:
0.508
Murlet(20):
0.131
Positive Predictive Value
ProbKnot:
0.517
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNAwolf
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAwolf:
0.310
Sensitivity
ProbKnot:
0.508
RNAwolf:
0.261
Positive Predictive Value
ProbKnot:
0.517
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RSpredict(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
RSpredict(seed):
0.295
Sensitivity
ProbKnot:
0.508
RSpredict(seed):
0.146
Positive Predictive Value
ProbKnot:
0.517
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CMfinder(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
CMfinder(20):
0.292
Sensitivity
ProbKnot:
0.508
CMfinder(20):
0.107
Positive Predictive Value
ProbKnot:
0.517
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Vsfold4
Matthews Correlation Coefficient
ProbKnot:
0.512
Vsfold4:
0.270
Sensitivity
ProbKnot:
0.508
Vsfold4:
0.198
Positive Predictive Value
ProbKnot:
0.517
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Vsfold5
Matthews Correlation Coefficient
ProbKnot:
0.512
Vsfold5:
0.260
Sensitivity
ProbKnot:
0.508
Vsfold5:
0.195
Positive Predictive Value
ProbKnot:
0.517
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNASampler(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
RNASampler(20):
0.246
Sensitivity
ProbKnot:
0.508
RNASampler(20):
0.069
Positive Predictive Value
ProbKnot:
0.517
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs HotKnots
Matthews Correlation Coefficient
ProbKnot:
0.512
HotKnots:
0.212
Sensitivity
ProbKnot:
0.508
HotKnots:
0.069
Positive Predictive Value
ProbKnot:
0.517
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Mastr(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Mastr(20):
0.187
Sensitivity
ProbKnot:
0.508
Mastr(20):
0.044
Positive Predictive Value
ProbKnot:
0.517
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Cylofold
Matthews Correlation Coefficient
ProbKnot:
0.512
Cylofold:
0.178
Sensitivity
ProbKnot:
0.508
Cylofold:
0.053
Positive Predictive Value
ProbKnot:
0.517
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Pknots
Matthews Correlation Coefficient
ProbKnot:
0.512
Pknots:
0.165
Sensitivity
ProbKnot:
0.508
Pknots:
0.041
Positive Predictive Value
ProbKnot:
0.517
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Alterna
Matthews Correlation Coefficient
ProbKnot:
0.512
Alterna:
0.145
Sensitivity
ProbKnot:
0.508
Alterna:
0.036
Positive Predictive Value
ProbKnot:
0.517
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs MCFold
Matthews Correlation Coefficient
ProbKnot:
0.512
MCFold:
0.138
Sensitivity
ProbKnot:
0.508
MCFold:
0.044
Positive Predictive Value
ProbKnot:
0.517
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RDfolder
Matthews Correlation Coefficient
ProbKnot:
0.512
RDfolder:
0.138
Sensitivity
ProbKnot:
0.508
RDfolder:
0.031
Positive Predictive Value
ProbKnot:
0.517
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Murlet(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Murlet(seed):
0.110
Sensitivity
ProbKnot:
0.508
Murlet(seed):
0.015
Positive Predictive Value
ProbKnot:
0.517
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs CMfinder(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
CMfinder(seed):
0.111
Sensitivity
ProbKnot:
0.508
CMfinder(seed):
0.016
Positive Predictive Value
ProbKnot:
0.517
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs TurboFold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
TurboFold(seed):
0.106
Sensitivity
ProbKnot:
0.508
TurboFold(seed):
0.014
Positive Predictive Value
ProbKnot:
0.517
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs PPfold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
PPfold(seed):
0.097
Sensitivity
ProbKnot:
0.508
PPfold(seed):
0.012
Positive Predictive Value
ProbKnot:
0.517
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Carnac(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Carnac(seed):
0.093
Sensitivity
ProbKnot:
0.508
Carnac(seed):
0.010
Positive Predictive Value
ProbKnot:
0.517
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs RNASampler(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
RNASampler(seed):
0.088
Sensitivity
ProbKnot:
0.508
RNASampler(seed):
0.009
Positive Predictive Value
ProbKnot:
0.517
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Mastr(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Mastr(seed):
0.072
Sensitivity
ProbKnot:
0.508
Mastr(seed):
0.006
Positive Predictive Value
ProbKnot:
0.517
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs NanoFolder
Matthews Correlation Coefficient
ProbKnot:
0.512
NanoFolder:
0.068
Sensitivity
ProbKnot:
0.508
NanoFolder:
0.027
Positive Predictive Value
ProbKnot:
0.517
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
ProbKnot vs Multilign(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Multilign(seed):
0.059
Sensitivity
ProbKnot:
0.508
Multilign(seed):
0.005
Positive Predictive Value
ProbKnot:
0.517
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MXScarna(20) |
1242
ContextFold vs MXScarna(20)
Matthews Correlation Coefficient
ContextFold:
0.754
MXScarna(20):
0.500
Sensitivity
ContextFold:
0.717
MXScarna(20):
0.353
Positive Predictive Value
ContextFold:
0.794
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MXScarna(20):
0.500
Sensitivity
PETfold_pre2.0(seed):
0.667
MXScarna(20):
0.353
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MXScarna(20):
0.500
Sensitivity
CentroidAlifold(seed):
0.414
MXScarna(20):
0.353
Positive Predictive Value
CentroidAlifold(seed):
0.901
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs MXScarna(20)
Matthews Correlation Coefficient
IPknot:
0.607
MXScarna(20):
0.500
Sensitivity
IPknot:
0.551
MXScarna(20):
0.353
Positive Predictive Value
IPknot:
0.670
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs MXScarna(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MXScarna(20):
0.500
Sensitivity
PETfold_pre2.0(20):
0.410
MXScarna(20):
0.353
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs MXScarna(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
MXScarna(20):
0.500
Sensitivity
CentroidFold:
0.515
MXScarna(20):
0.353
Positive Predictive Value
CentroidFold:
0.633
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MXScarna(20):
0.500
Sensitivity
CentroidAlifold(20):
0.369
MXScarna(20):
0.353
Positive Predictive Value
CentroidAlifold(20):
0.891
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs MXScarna(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MXScarna(20):
0.500
Sensitivity
CentroidHomfold‑LAST:
0.359
MXScarna(20):
0.353
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs MXScarna(20)
Matthews Correlation Coefficient
Contrafold:
0.556
MXScarna(20):
0.500
Sensitivity
Contrafold:
0.542
MXScarna(20):
0.353
Positive Predictive Value
Contrafold:
0.571
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs MXScarna(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
MXScarna(20):
0.500
Sensitivity
MXScarna(seed):
0.382
MXScarna(20):
0.353
Positive Predictive Value
MXScarna(seed):
0.753
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
MXScarna(20):
0.500
Sensitivity
RNAalifold(20):
0.359
MXScarna(20):
0.353
Positive Predictive Value
RNAalifold(20):
0.787
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs MXScarna(20)
Matthews Correlation Coefficient
Sfold:
0.527
MXScarna(20):
0.500
Sensitivity
Sfold:
0.477
MXScarna(20):
0.353
Positive Predictive Value
Sfold:
0.582
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs MXScarna(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
MXScarna(20):
0.500
Sensitivity
RNAalifold(seed):
0.328
MXScarna(20):
0.353
Positive Predictive Value
RNAalifold(seed):
0.835
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs MXScarna(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
MXScarna(20):
0.500
Sensitivity
MaxExpect:
0.505
MXScarna(20):
0.353
Positive Predictive Value
MaxExpect:
0.528
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs MXScarna(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
MXScarna(20):
0.500
Sensitivity
ProbKnot:
0.508
MXScarna(20):
0.353
Positive Predictive Value
ProbKnot:
0.517
MXScarna(20):
0.710
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
MXScarna(20) vs UNAFold
Matthews Correlation Coefficient
MXScarna(20):
0.500
UNAFold:
0.494
Sensitivity
MXScarna(20):
0.353
UNAFold:
0.491
Positive Predictive Value
MXScarna(20):
0.710
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Fold
Matthews Correlation Coefficient
MXScarna(20):
0.500
Fold:
0.493
Sensitivity
MXScarna(20):
0.353
Fold:
0.496
Positive Predictive Value
MXScarna(20):
0.710
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
MXScarna(20) vs RNAfold
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAfold:
0.485
Sensitivity
MXScarna(20):
0.353
RNAfold:
0.486
Positive Predictive Value
MXScarna(20):
0.710
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs PknotsRG
Matthews Correlation Coefficient
MXScarna(20):
0.500
PknotsRG:
0.474
Sensitivity
MXScarna(20):
0.353
PknotsRG:
0.440
Positive Predictive Value
MXScarna(20):
0.710
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(20):
0.500
CRWrnafold:
0.430
Sensitivity
MXScarna(20):
0.353
CRWrnafold:
0.379
Positive Predictive Value
MXScarna(20):
0.710
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAsubopt:
0.422
Sensitivity
MXScarna(20):
0.353
RNAsubopt:
0.344
Positive Predictive Value
MXScarna(20):
0.710
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
TurboFold(20):
0.419
Sensitivity
MXScarna(20):
0.353
TurboFold(20):
0.210
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
PPfold(20):
0.416
Sensitivity
MXScarna(20):
0.353
PPfold(20):
0.197
Positive Predictive Value
MXScarna(20):
0.710
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Afold
Matthews Correlation Coefficient
MXScarna(20):
0.500
Afold:
0.408
Sensitivity
MXScarna(20):
0.353
Afold:
0.343
Positive Predictive Value
MXScarna(20):
0.710
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Carnac(20):
0.401
Sensitivity
MXScarna(20):
0.353
Carnac(20):
0.180
Positive Predictive Value
MXScarna(20):
0.710
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs McQFold
Matthews Correlation Coefficient
MXScarna(20):
0.500
McQFold:
0.389
Sensitivity
MXScarna(20):
0.353
McQFold:
0.308
Positive Predictive Value
MXScarna(20):
0.710
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RSpredict(20):
0.377
Sensitivity
MXScarna(20):
0.353
RSpredict(20):
0.241
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAshapes:
0.367
Sensitivity
MXScarna(20):
0.353
RNAshapes:
0.285
Positive Predictive Value
MXScarna(20):
0.710
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNASLOpt:
0.364
Sensitivity
MXScarna(20):
0.353
RNASLOpt:
0.261
Positive Predictive Value
MXScarna(20):
0.710
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Multilign(20):
0.350
Sensitivity
MXScarna(20):
0.353
Multilign(20):
0.161
Positive Predictive Value
MXScarna(20):
0.710
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Murlet(20):
0.321
Sensitivity
MXScarna(20):
0.353
Murlet(20):
0.131
Positive Predictive Value
MXScarna(20):
0.710
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAwolf:
0.310
Sensitivity
MXScarna(20):
0.353
RNAwolf:
0.261
Positive Predictive Value
MXScarna(20):
0.710
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RSpredict(seed):
0.295
Sensitivity
MXScarna(20):
0.353
RSpredict(seed):
0.146
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
CMfinder(20):
0.292
Sensitivity
MXScarna(20):
0.353
CMfinder(20):
0.107
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(20):
0.500
Vsfold4:
0.270
Sensitivity
MXScarna(20):
0.353
Vsfold4:
0.198
Positive Predictive Value
MXScarna(20):
0.710
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(20):
0.500
Vsfold5:
0.260
Sensitivity
MXScarna(20):
0.353
Vsfold5:
0.195
Positive Predictive Value
MXScarna(20):
0.710
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNASampler(20):
0.246
Sensitivity
MXScarna(20):
0.353
RNASampler(20):
0.069
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs HotKnots
Matthews Correlation Coefficient
MXScarna(20):
0.500
HotKnots:
0.212
Sensitivity
MXScarna(20):
0.353
HotKnots:
0.069
Positive Predictive Value
MXScarna(20):
0.710
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Mastr(20):
0.187
Sensitivity
MXScarna(20):
0.353
Mastr(20):
0.044
Positive Predictive Value
MXScarna(20):
0.710
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Cylofold
Matthews Correlation Coefficient
MXScarna(20):
0.500
Cylofold:
0.178
Sensitivity
MXScarna(20):
0.353
Cylofold:
0.053
Positive Predictive Value
MXScarna(20):
0.710
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Pknots
Matthews Correlation Coefficient
MXScarna(20):
0.500
Pknots:
0.165
Sensitivity
MXScarna(20):
0.353
Pknots:
0.041
Positive Predictive Value
MXScarna(20):
0.710
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Alterna
Matthews Correlation Coefficient
MXScarna(20):
0.500
Alterna:
0.145
Sensitivity
MXScarna(20):
0.353
Alterna:
0.036
Positive Predictive Value
MXScarna(20):
0.710
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs MCFold
Matthews Correlation Coefficient
MXScarna(20):
0.500
MCFold:
0.138
Sensitivity
MXScarna(20):
0.353
MCFold:
0.044
Positive Predictive Value
MXScarna(20):
0.710
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RDfolder
Matthews Correlation Coefficient
MXScarna(20):
0.500
RDfolder:
0.138
Sensitivity
MXScarna(20):
0.353
RDfolder:
0.031
Positive Predictive Value
MXScarna(20):
0.710
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Murlet(seed):
0.110
Sensitivity
MXScarna(20):
0.353
Murlet(seed):
0.015
Positive Predictive Value
MXScarna(20):
0.710
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
CMfinder(seed):
0.111
Sensitivity
MXScarna(20):
0.353
CMfinder(seed):
0.016
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
TurboFold(seed):
0.106
Sensitivity
MXScarna(20):
0.353
TurboFold(seed):
0.014
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
PPfold(seed):
0.097
Sensitivity
MXScarna(20):
0.353
PPfold(seed):
0.012
Positive Predictive Value
MXScarna(20):
0.710
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Carnac(seed):
0.093
Sensitivity
MXScarna(20):
0.353
Carnac(seed):
0.010
Positive Predictive Value
MXScarna(20):
0.710
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNASampler(seed):
0.088
Sensitivity
MXScarna(20):
0.353
RNASampler(seed):
0.009
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Mastr(seed):
0.072
Sensitivity
MXScarna(20):
0.353
Mastr(seed):
0.006
Positive Predictive Value
MXScarna(20):
0.710
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(20):
0.500
NanoFolder:
0.068
Sensitivity
MXScarna(20):
0.353
NanoFolder:
0.027
Positive Predictive Value
MXScarna(20):
0.710
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MXScarna(20) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Multilign(seed):
0.059
Sensitivity
MXScarna(20):
0.353
Multilign(seed):
0.005
Positive Predictive Value
MXScarna(20):
0.710
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| UNAFold |
1242
ContextFold vs UNAFold
Matthews Correlation Coefficient
ContextFold:
0.754
UNAFold:
0.494
Sensitivity
ContextFold:
0.717
UNAFold:
0.491
Positive Predictive Value
ContextFold:
0.794
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs UNAFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
UNAFold:
0.494
Sensitivity
PETfold_pre2.0(seed):
0.667
UNAFold:
0.491
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs UNAFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
UNAFold:
0.494
Sensitivity
CentroidAlifold(seed):
0.414
UNAFold:
0.491
Positive Predictive Value
CentroidAlifold(seed):
0.901
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs UNAFold
Matthews Correlation Coefficient
IPknot:
0.607
UNAFold:
0.494
Sensitivity
IPknot:
0.551
UNAFold:
0.491
Positive Predictive Value
IPknot:
0.670
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs UNAFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
UNAFold:
0.494
Sensitivity
PETfold_pre2.0(20):
0.410
UNAFold:
0.491
Positive Predictive Value
PETfold_pre2.0(20):
0.807
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs UNAFold
Matthews Correlation Coefficient
CentroidFold:
0.571
UNAFold:
0.494
Sensitivity
CentroidFold:
0.515
UNAFold:
0.491
Positive Predictive Value
CentroidFold:
0.633
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs UNAFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
UNAFold:
0.494
Sensitivity
CentroidAlifold(20):
0.369
UNAFold:
0.491
Positive Predictive Value
CentroidAlifold(20):
0.891
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs UNAFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
UNAFold:
0.494
Sensitivity
CentroidHomfold‑LAST:
0.359
UNAFold:
0.491
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs UNAFold
Matthews Correlation Coefficient
Contrafold:
0.556
UNAFold:
0.494
Sensitivity
Contrafold:
0.542
UNAFold:
0.491
Positive Predictive Value
Contrafold:
0.571
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs UNAFold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
UNAFold:
0.494
Sensitivity
MXScarna(seed):
0.382
UNAFold:
0.491
Positive Predictive Value
MXScarna(seed):
0.753
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs UNAFold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
UNAFold:
0.494
Sensitivity
RNAalifold(20):
0.359
UNAFold:
0.491
Positive Predictive Value
RNAalifold(20):
0.787
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs UNAFold
Matthews Correlation Coefficient
Sfold:
0.527
UNAFold:
0.494
Sensitivity
Sfold:
0.477
UNAFold:
0.491
Positive Predictive Value
Sfold:
0.582
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs UNAFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
UNAFold:
0.494
Sensitivity
RNAalifold(seed):
0.328
UNAFold:
0.491
Positive Predictive Value
RNAalifold(seed):
0.835
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs UNAFold
Matthews Correlation Coefficient
MaxExpect:
0.516
UNAFold:
0.494
Sensitivity
MaxExpect:
0.505
UNAFold:
0.491
Positive Predictive Value
MaxExpect:
0.528
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs UNAFold
Matthews Correlation Coefficient
ProbKnot:
0.512
UNAFold:
0.494
Sensitivity
ProbKnot:
0.508
UNAFold:
0.491
Positive Predictive Value
ProbKnot:
0.517
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs UNAFold
Matthews Correlation Coefficient
MXScarna(20):
0.500
UNAFold:
0.494
Sensitivity
MXScarna(20):
0.353
UNAFold:
0.491
Positive Predictive Value
MXScarna(20):
0.710
UNAFold:
0.498
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
UNAFold vs Fold
Matthews Correlation Coefficient
UNAFold:
0.494
Fold:
0.493
Sensitivity
UNAFold:
0.491
Fold:
0.496
Positive Predictive Value
UNAFold:
0.498
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.00238364898467
|
+
UNAFold vs RNAfold
Matthews Correlation Coefficient
UNAFold:
0.494
RNAfold:
0.485
Sensitivity
UNAFold:
0.491
RNAfold:
0.486
Positive Predictive Value
UNAFold:
0.498
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PknotsRG
Matthews Correlation Coefficient
UNAFold:
0.494
PknotsRG:
0.474
Sensitivity
UNAFold:
0.491
PknotsRG:
0.440
Positive Predictive Value
UNAFold:
0.498
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CRWrnafold
Matthews Correlation Coefficient
UNAFold:
0.494
CRWrnafold:
0.430
Sensitivity
UNAFold:
0.491
CRWrnafold:
0.379
Positive Predictive Value
UNAFold:
0.498
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAsubopt
Matthews Correlation Coefficient
UNAFold:
0.494
RNAsubopt:
0.422
Sensitivity
UNAFold:
0.491
RNAsubopt:
0.344
Positive Predictive Value
UNAFold:
0.498
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs TurboFold(20)
Matthews Correlation Coefficient
UNAFold:
0.494
TurboFold(20):
0.419
Sensitivity
UNAFold:
0.491
TurboFold(20):
0.210
Positive Predictive Value
UNAFold:
0.498
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PPfold(20)
Matthews Correlation Coefficient
UNAFold:
0.494
PPfold(20):
0.416
Sensitivity
UNAFold:
0.491
PPfold(20):
0.197
Positive Predictive Value
UNAFold:
0.498
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Afold
Matthews Correlation Coefficient
UNAFold:
0.494
Afold:
0.408
Sensitivity
UNAFold:
0.491
Afold:
0.343
Positive Predictive Value
UNAFold:
0.498
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Carnac(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Carnac(20):
0.401
Sensitivity
UNAFold:
0.491
Carnac(20):
0.180
Positive Predictive Value
UNAFold:
0.498
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs McQFold
Matthews Correlation Coefficient
UNAFold:
0.494
McQFold:
0.389
Sensitivity
UNAFold:
0.491
McQFold:
0.308
Positive Predictive Value
UNAFold:
0.498
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RSpredict(20)
Matthews Correlation Coefficient
UNAFold:
0.494
RSpredict(20):
0.377
Sensitivity
UNAFold:
0.491
RSpredict(20):
0.241
Positive Predictive Value
UNAFold:
0.498
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAshapes
Matthews Correlation Coefficient
UNAFold:
0.494
RNAshapes:
0.367
Sensitivity
UNAFold:
0.491
RNAshapes:
0.285
Positive Predictive Value
UNAFold:
0.498
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNASLOpt
Matthews Correlation Coefficient
UNAFold:
0.494
RNASLOpt:
0.364
Sensitivity
UNAFold:
0.491
RNASLOpt:
0.261
Positive Predictive Value
UNAFold:
0.498
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Multilign(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Multilign(20):
0.350
Sensitivity
UNAFold:
0.491
Multilign(20):
0.161
Positive Predictive Value
UNAFold:
0.498
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Murlet(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Murlet(20):
0.321
Sensitivity
UNAFold:
0.491
Murlet(20):
0.131
Positive Predictive Value
UNAFold:
0.498
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNAwolf
Matthews Correlation Coefficient
UNAFold:
0.494
RNAwolf:
0.310
Sensitivity
UNAFold:
0.491
RNAwolf:
0.261
Positive Predictive Value
UNAFold:
0.498
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RSpredict(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
RSpredict(seed):
0.295
Sensitivity
UNAFold:
0.491
RSpredict(seed):
0.146
Positive Predictive Value
UNAFold:
0.498
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CMfinder(20)
Matthews Correlation Coefficient
UNAFold:
0.494
CMfinder(20):
0.292
Sensitivity
UNAFold:
0.491
CMfinder(20):
0.107
Positive Predictive Value
UNAFold:
0.498
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Vsfold4
Matthews Correlation Coefficient
UNAFold:
0.494
Vsfold4:
0.270
Sensitivity
UNAFold:
0.491
Vsfold4:
0.198
Positive Predictive Value
UNAFold:
0.498
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Vsfold5
Matthews Correlation Coefficient
UNAFold:
0.494
Vsfold5:
0.260
Sensitivity
UNAFold:
0.491
Vsfold5:
0.195
Positive Predictive Value
UNAFold:
0.498
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNASampler(20)
Matthews Correlation Coefficient
UNAFold:
0.494
RNASampler(20):
0.246
Sensitivity
UNAFold:
0.491
RNASampler(20):
0.069
Positive Predictive Value
UNAFold:
0.498
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs HotKnots
Matthews Correlation Coefficient
UNAFold:
0.494
HotKnots:
0.212
Sensitivity
UNAFold:
0.491
HotKnots:
0.069
Positive Predictive Value
UNAFold:
0.498
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Mastr(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Mastr(20):
0.187
Sensitivity
UNAFold:
0.491
Mastr(20):
0.044
Positive Predictive Value
UNAFold:
0.498
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Cylofold
Matthews Correlation Coefficient
UNAFold:
0.494
Cylofold:
0.178
Sensitivity
UNAFold:
0.491
Cylofold:
0.053
Positive Predictive Value
UNAFold:
0.498
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Pknots
Matthews Correlation Coefficient
UNAFold:
0.494
Pknots:
0.165
Sensitivity
UNAFold:
0.491
Pknots:
0.041
Positive Predictive Value
UNAFold:
0.498
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Alterna
Matthews Correlation Coefficient
UNAFold:
0.494
Alterna:
0.145
Sensitivity
UNAFold:
0.491
Alterna:
0.036
Positive Predictive Value
UNAFold:
0.498
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs MCFold
Matthews Correlation Coefficient
UNAFold:
0.494
MCFold:
0.138
Sensitivity
UNAFold:
0.491
MCFold:
0.044
Positive Predictive Value
UNAFold:
0.498
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RDfolder
Matthews Correlation Coefficient
UNAFold:
0.494
RDfolder:
0.138
Sensitivity
UNAFold:
0.491
RDfolder:
0.031
Positive Predictive Value
UNAFold:
0.498
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Murlet(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Murlet(seed):
0.110
Sensitivity
UNAFold:
0.491
Murlet(seed):
0.015
Positive Predictive Value
UNAFold:
0.498
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs CMfinder(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
CMfinder(seed):
0.111
Sensitivity
UNAFold:
0.491
CMfinder(seed):
0.016
Positive Predictive Value
UNAFold:
0.498
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs TurboFold(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
TurboFold(seed):
0.106
Sensitivity
UNAFold:
0.491
TurboFold(seed):
0.014
Positive Predictive Value
UNAFold:
0.498
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs PPfold(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
PPfold(seed):
0.097
Sensitivity
UNAFold:
0.491
PPfold(seed):
0.012
Positive Predictive Value
UNAFold:
0.498
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Carnac(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Carnac(seed):
0.093
Sensitivity
UNAFold:
0.491
Carnac(seed):
0.010
Positive Predictive Value
UNAFold:
0.498
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs RNASampler(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
RNASampler(seed):
0.088
Sensitivity
UNAFold:
0.491
RNASampler(seed):
0.009
Positive Predictive Value
UNAFold:
0.498
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Mastr(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Mastr(seed):
0.072
Sensitivity
UNAFold:
0.491
Mastr(seed):
0.006
Positive Predictive Value
UNAFold:
0.498
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs NanoFolder
Matthews Correlation Coefficient
UNAFold:
0.494
NanoFolder:
0.068
Sensitivity
UNAFold:
0.491
NanoFolder:
0.027
Positive Predictive Value
UNAFold:
0.498
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
UNAFold vs Multilign(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Multilign(seed):
0.059
Sensitivity
UNAFold:
0.491
Multilign(seed):
0.005
Positive Predictive Value
UNAFold:
0.498
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Fold |
1242
ContextFold vs Fold
Matthews Correlation Coefficient
ContextFold:
0.754
Fold:
0.493
Sensitivity
ContextFold:
0.717
Fold:
0.496
Positive Predictive Value
ContextFold:
0.794
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Fold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Fold:
0.493
Sensitivity
PETfold_pre2.0(seed):
0.667
Fold:
0.496
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Fold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Fold:
0.493
Sensitivity
CentroidAlifold(seed):
0.414
Fold:
0.496
Positive Predictive Value
CentroidAlifold(seed):
0.901
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Fold
Matthews Correlation Coefficient
IPknot:
0.607
Fold:
0.493
Sensitivity
IPknot:
0.551
Fold:
0.496
Positive Predictive Value
IPknot:
0.670
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Fold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Fold:
0.493
Sensitivity
PETfold_pre2.0(20):
0.410
Fold:
0.496
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Fold
Matthews Correlation Coefficient
CentroidFold:
0.571
Fold:
0.493
Sensitivity
CentroidFold:
0.515
Fold:
0.496
Positive Predictive Value
CentroidFold:
0.633
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Fold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Fold:
0.493
Sensitivity
CentroidAlifold(20):
0.369
Fold:
0.496
Positive Predictive Value
CentroidAlifold(20):
0.891
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Fold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Fold:
0.493
Sensitivity
CentroidHomfold‑LAST:
0.359
Fold:
0.496
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Fold
Matthews Correlation Coefficient
Contrafold:
0.556
Fold:
0.493
Sensitivity
Contrafold:
0.542
Fold:
0.496
Positive Predictive Value
Contrafold:
0.571
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Fold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Fold:
0.493
Sensitivity
MXScarna(seed):
0.382
Fold:
0.496
Positive Predictive Value
MXScarna(seed):
0.753
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Fold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Fold:
0.493
Sensitivity
RNAalifold(20):
0.359
Fold:
0.496
Positive Predictive Value
RNAalifold(20):
0.787
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Fold
Matthews Correlation Coefficient
Sfold:
0.527
Fold:
0.493
Sensitivity
Sfold:
0.477
Fold:
0.496
Positive Predictive Value
Sfold:
0.582
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Fold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Fold:
0.493
Sensitivity
RNAalifold(seed):
0.328
Fold:
0.496
Positive Predictive Value
RNAalifold(seed):
0.835
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Fold
Matthews Correlation Coefficient
MaxExpect:
0.516
Fold:
0.493
Sensitivity
MaxExpect:
0.505
Fold:
0.496
Positive Predictive Value
MaxExpect:
0.528
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Fold
Matthews Correlation Coefficient
ProbKnot:
0.512
Fold:
0.493
Sensitivity
ProbKnot:
0.508
Fold:
0.496
Positive Predictive Value
ProbKnot:
0.517
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Fold
Matthews Correlation Coefficient
MXScarna(20):
0.500
Fold:
0.493
Sensitivity
MXScarna(20):
0.353
Fold:
0.496
Positive Predictive Value
MXScarna(20):
0.710
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
1242
UNAFold vs Fold
Matthews Correlation Coefficient
UNAFold:
0.494
Fold:
0.493
Sensitivity
UNAFold:
0.491
Fold:
0.496
Positive Predictive Value
UNAFold:
0.498
Fold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.00238364898467
|
|
+
Fold vs RNAfold
Matthews Correlation Coefficient
Fold:
0.493
RNAfold:
0.485
Sensitivity
Fold:
0.496
RNAfold:
0.486
Positive Predictive Value
Fold:
0.490
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PknotsRG
Matthews Correlation Coefficient
Fold:
0.493
PknotsRG:
0.474
Sensitivity
Fold:
0.496
PknotsRG:
0.440
Positive Predictive Value
Fold:
0.490
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CRWrnafold
Matthews Correlation Coefficient
Fold:
0.493
CRWrnafold:
0.430
Sensitivity
Fold:
0.496
CRWrnafold:
0.379
Positive Predictive Value
Fold:
0.490
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAsubopt
Matthews Correlation Coefficient
Fold:
0.493
RNAsubopt:
0.422
Sensitivity
Fold:
0.496
RNAsubopt:
0.344
Positive Predictive Value
Fold:
0.490
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs TurboFold(20)
Matthews Correlation Coefficient
Fold:
0.493
TurboFold(20):
0.419
Sensitivity
Fold:
0.496
TurboFold(20):
0.210
Positive Predictive Value
Fold:
0.490
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PPfold(20)
Matthews Correlation Coefficient
Fold:
0.493
PPfold(20):
0.416
Sensitivity
Fold:
0.496
PPfold(20):
0.197
Positive Predictive Value
Fold:
0.490
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Afold
Matthews Correlation Coefficient
Fold:
0.493
Afold:
0.408
Sensitivity
Fold:
0.496
Afold:
0.343
Positive Predictive Value
Fold:
0.490
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Carnac(20)
Matthews Correlation Coefficient
Fold:
0.493
Carnac(20):
0.401
Sensitivity
Fold:
0.496
Carnac(20):
0.180
Positive Predictive Value
Fold:
0.490
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs McQFold
Matthews Correlation Coefficient
Fold:
0.493
McQFold:
0.389
Sensitivity
Fold:
0.496
McQFold:
0.308
Positive Predictive Value
Fold:
0.490
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RSpredict(20)
Matthews Correlation Coefficient
Fold:
0.493
RSpredict(20):
0.377
Sensitivity
Fold:
0.496
RSpredict(20):
0.241
Positive Predictive Value
Fold:
0.490
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAshapes
Matthews Correlation Coefficient
Fold:
0.493
RNAshapes:
0.367
Sensitivity
Fold:
0.496
RNAshapes:
0.285
Positive Predictive Value
Fold:
0.490
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNASLOpt
Matthews Correlation Coefficient
Fold:
0.493
RNASLOpt:
0.364
Sensitivity
Fold:
0.496
RNASLOpt:
0.261
Positive Predictive Value
Fold:
0.490
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Multilign(20)
Matthews Correlation Coefficient
Fold:
0.493
Multilign(20):
0.350
Sensitivity
Fold:
0.496
Multilign(20):
0.161
Positive Predictive Value
Fold:
0.490
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Murlet(20)
Matthews Correlation Coefficient
Fold:
0.493
Murlet(20):
0.321
Sensitivity
Fold:
0.496
Murlet(20):
0.131
Positive Predictive Value
Fold:
0.490
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNAwolf
Matthews Correlation Coefficient
Fold:
0.493
RNAwolf:
0.310
Sensitivity
Fold:
0.496
RNAwolf:
0.261
Positive Predictive Value
Fold:
0.490
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RSpredict(seed)
Matthews Correlation Coefficient
Fold:
0.493
RSpredict(seed):
0.295
Sensitivity
Fold:
0.496
RSpredict(seed):
0.146
Positive Predictive Value
Fold:
0.490
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CMfinder(20)
Matthews Correlation Coefficient
Fold:
0.493
CMfinder(20):
0.292
Sensitivity
Fold:
0.496
CMfinder(20):
0.107
Positive Predictive Value
Fold:
0.490
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Vsfold4
Matthews Correlation Coefficient
Fold:
0.493
Vsfold4:
0.270
Sensitivity
Fold:
0.496
Vsfold4:
0.198
Positive Predictive Value
Fold:
0.490
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Vsfold5
Matthews Correlation Coefficient
Fold:
0.493
Vsfold5:
0.260
Sensitivity
Fold:
0.496
Vsfold5:
0.195
Positive Predictive Value
Fold:
0.490
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNASampler(20)
Matthews Correlation Coefficient
Fold:
0.493
RNASampler(20):
0.246
Sensitivity
Fold:
0.496
RNASampler(20):
0.069
Positive Predictive Value
Fold:
0.490
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs HotKnots
Matthews Correlation Coefficient
Fold:
0.493
HotKnots:
0.212
Sensitivity
Fold:
0.496
HotKnots:
0.069
Positive Predictive Value
Fold:
0.490
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Mastr(20)
Matthews Correlation Coefficient
Fold:
0.493
Mastr(20):
0.187
Sensitivity
Fold:
0.496
Mastr(20):
0.044
Positive Predictive Value
Fold:
0.490
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Cylofold
Matthews Correlation Coefficient
Fold:
0.493
Cylofold:
0.178
Sensitivity
Fold:
0.496
Cylofold:
0.053
Positive Predictive Value
Fold:
0.490
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Pknots
Matthews Correlation Coefficient
Fold:
0.493
Pknots:
0.165
Sensitivity
Fold:
0.496
Pknots:
0.041
Positive Predictive Value
Fold:
0.490
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Alterna
Matthews Correlation Coefficient
Fold:
0.493
Alterna:
0.145
Sensitivity
Fold:
0.496
Alterna:
0.036
Positive Predictive Value
Fold:
0.490
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs MCFold
Matthews Correlation Coefficient
Fold:
0.493
MCFold:
0.138
Sensitivity
Fold:
0.496
MCFold:
0.044
Positive Predictive Value
Fold:
0.490
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RDfolder
Matthews Correlation Coefficient
Fold:
0.493
RDfolder:
0.138
Sensitivity
Fold:
0.496
RDfolder:
0.031
Positive Predictive Value
Fold:
0.490
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Murlet(seed)
Matthews Correlation Coefficient
Fold:
0.493
Murlet(seed):
0.110
Sensitivity
Fold:
0.496
Murlet(seed):
0.015
Positive Predictive Value
Fold:
0.490
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs CMfinder(seed)
Matthews Correlation Coefficient
Fold:
0.493
CMfinder(seed):
0.111
Sensitivity
Fold:
0.496
CMfinder(seed):
0.016
Positive Predictive Value
Fold:
0.490
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs TurboFold(seed)
Matthews Correlation Coefficient
Fold:
0.493
TurboFold(seed):
0.106
Sensitivity
Fold:
0.496
TurboFold(seed):
0.014
Positive Predictive Value
Fold:
0.490
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs PPfold(seed)
Matthews Correlation Coefficient
Fold:
0.493
PPfold(seed):
0.097
Sensitivity
Fold:
0.496
PPfold(seed):
0.012
Positive Predictive Value
Fold:
0.490
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Carnac(seed)
Matthews Correlation Coefficient
Fold:
0.493
Carnac(seed):
0.093
Sensitivity
Fold:
0.496
Carnac(seed):
0.010
Positive Predictive Value
Fold:
0.490
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs RNASampler(seed)
Matthews Correlation Coefficient
Fold:
0.493
RNASampler(seed):
0.088
Sensitivity
Fold:
0.496
RNASampler(seed):
0.009
Positive Predictive Value
Fold:
0.490
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Mastr(seed)
Matthews Correlation Coefficient
Fold:
0.493
Mastr(seed):
0.072
Sensitivity
Fold:
0.496
Mastr(seed):
0.006
Positive Predictive Value
Fold:
0.490
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs NanoFolder
Matthews Correlation Coefficient
Fold:
0.493
NanoFolder:
0.068
Sensitivity
Fold:
0.496
NanoFolder:
0.027
Positive Predictive Value
Fold:
0.490
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Fold vs Multilign(seed)
Matthews Correlation Coefficient
Fold:
0.493
Multilign(seed):
0.059
Sensitivity
Fold:
0.496
Multilign(seed):
0.005
Positive Predictive Value
Fold:
0.490
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAfold |
1242
ContextFold vs RNAfold
Matthews Correlation Coefficient
ContextFold:
0.754
RNAfold:
0.485
Sensitivity
ContextFold:
0.717
RNAfold:
0.486
Positive Predictive Value
ContextFold:
0.794
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNAfold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAfold:
0.485
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAfold:
0.486
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNAfold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAfold:
0.485
Sensitivity
CentroidAlifold(seed):
0.414
RNAfold:
0.486
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNAfold
Matthews Correlation Coefficient
IPknot:
0.607
RNAfold:
0.485
Sensitivity
IPknot:
0.551
RNAfold:
0.486
Positive Predictive Value
IPknot:
0.670
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNAfold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAfold:
0.485
Sensitivity
PETfold_pre2.0(20):
0.410
RNAfold:
0.486
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNAfold
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAfold:
0.485
Sensitivity
CentroidFold:
0.515
RNAfold:
0.486
Positive Predictive Value
CentroidFold:
0.633
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNAfold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAfold:
0.485
Sensitivity
CentroidAlifold(20):
0.369
RNAfold:
0.486
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNAfold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAfold:
0.485
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAfold:
0.486
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNAfold
Matthews Correlation Coefficient
Contrafold:
0.556
RNAfold:
0.485
Sensitivity
Contrafold:
0.542
RNAfold:
0.486
Positive Predictive Value
Contrafold:
0.571
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNAfold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAfold:
0.485
Sensitivity
MXScarna(seed):
0.382
RNAfold:
0.486
Positive Predictive Value
MXScarna(seed):
0.753
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNAfold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAfold:
0.485
Sensitivity
RNAalifold(20):
0.359
RNAfold:
0.486
Positive Predictive Value
RNAalifold(20):
0.787
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNAfold
Matthews Correlation Coefficient
Sfold:
0.527
RNAfold:
0.485
Sensitivity
Sfold:
0.477
RNAfold:
0.486
Positive Predictive Value
Sfold:
0.582
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNAfold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAfold:
0.485
Sensitivity
RNAalifold(seed):
0.328
RNAfold:
0.486
Positive Predictive Value
RNAalifold(seed):
0.835
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNAfold
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAfold:
0.485
Sensitivity
MaxExpect:
0.505
RNAfold:
0.486
Positive Predictive Value
MaxExpect:
0.528
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNAfold
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAfold:
0.485
Sensitivity
ProbKnot:
0.508
RNAfold:
0.486
Positive Predictive Value
ProbKnot:
0.517
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNAfold
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAfold:
0.485
Sensitivity
MXScarna(20):
0.353
RNAfold:
0.486
Positive Predictive Value
MXScarna(20):
0.710
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNAfold
Matthews Correlation Coefficient
UNAFold:
0.494
RNAfold:
0.485
Sensitivity
UNAFold:
0.491
RNAfold:
0.486
Positive Predictive Value
UNAFold:
0.498
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNAfold
Matthews Correlation Coefficient
Fold:
0.493
RNAfold:
0.485
Sensitivity
Fold:
0.496
RNAfold:
0.486
Positive Predictive Value
Fold:
0.490
RNAfold:
0.485
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAfold vs PknotsRG
Matthews Correlation Coefficient
RNAfold:
0.485
PknotsRG:
0.474
Sensitivity
RNAfold:
0.486
PknotsRG:
0.440
Positive Predictive Value
RNAfold:
0.485
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CRWrnafold
Matthews Correlation Coefficient
RNAfold:
0.485
CRWrnafold:
0.430
Sensitivity
RNAfold:
0.486
CRWrnafold:
0.379
Positive Predictive Value
RNAfold:
0.485
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAsubopt
Matthews Correlation Coefficient
RNAfold:
0.485
RNAsubopt:
0.422
Sensitivity
RNAfold:
0.486
RNAsubopt:
0.344
Positive Predictive Value
RNAfold:
0.485
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs TurboFold(20)
Matthews Correlation Coefficient
RNAfold:
0.485
TurboFold(20):
0.419
Sensitivity
RNAfold:
0.486
TurboFold(20):
0.210
Positive Predictive Value
RNAfold:
0.485
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs PPfold(20)
Matthews Correlation Coefficient
RNAfold:
0.485
PPfold(20):
0.416
Sensitivity
RNAfold:
0.486
PPfold(20):
0.197
Positive Predictive Value
RNAfold:
0.485
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Afold
Matthews Correlation Coefficient
RNAfold:
0.485
Afold:
0.408
Sensitivity
RNAfold:
0.486
Afold:
0.343
Positive Predictive Value
RNAfold:
0.485
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Carnac(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Carnac(20):
0.401
Sensitivity
RNAfold:
0.486
Carnac(20):
0.180
Positive Predictive Value
RNAfold:
0.485
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs McQFold
Matthews Correlation Coefficient
RNAfold:
0.485
McQFold:
0.389
Sensitivity
RNAfold:
0.486
McQFold:
0.308
Positive Predictive Value
RNAfold:
0.485
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RSpredict(20)
Matthews Correlation Coefficient
RNAfold:
0.485
RSpredict(20):
0.377
Sensitivity
RNAfold:
0.486
RSpredict(20):
0.241
Positive Predictive Value
RNAfold:
0.485
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAshapes
Matthews Correlation Coefficient
RNAfold:
0.485
RNAshapes:
0.367
Sensitivity
RNAfold:
0.486
RNAshapes:
0.285
Positive Predictive Value
RNAfold:
0.485
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNASLOpt
Matthews Correlation Coefficient
RNAfold:
0.485
RNASLOpt:
0.364
Sensitivity
RNAfold:
0.486
RNASLOpt:
0.261
Positive Predictive Value
RNAfold:
0.485
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Multilign(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Multilign(20):
0.350
Sensitivity
RNAfold:
0.486
Multilign(20):
0.161
Positive Predictive Value
RNAfold:
0.485
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Murlet(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Murlet(20):
0.321
Sensitivity
RNAfold:
0.486
Murlet(20):
0.131
Positive Predictive Value
RNAfold:
0.485
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNAwolf
Matthews Correlation Coefficient
RNAfold:
0.485
RNAwolf:
0.310
Sensitivity
RNAfold:
0.486
RNAwolf:
0.261
Positive Predictive Value
RNAfold:
0.485
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RSpredict(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
RSpredict(seed):
0.295
Sensitivity
RNAfold:
0.486
RSpredict(seed):
0.146
Positive Predictive Value
RNAfold:
0.485
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CMfinder(20)
Matthews Correlation Coefficient
RNAfold:
0.485
CMfinder(20):
0.292
Sensitivity
RNAfold:
0.486
CMfinder(20):
0.107
Positive Predictive Value
RNAfold:
0.485
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Vsfold4
Matthews Correlation Coefficient
RNAfold:
0.485
Vsfold4:
0.270
Sensitivity
RNAfold:
0.486
Vsfold4:
0.198
Positive Predictive Value
RNAfold:
0.485
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Vsfold5
Matthews Correlation Coefficient
RNAfold:
0.485
Vsfold5:
0.260
Sensitivity
RNAfold:
0.486
Vsfold5:
0.195
Positive Predictive Value
RNAfold:
0.485
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNASampler(20)
Matthews Correlation Coefficient
RNAfold:
0.485
RNASampler(20):
0.246
Sensitivity
RNAfold:
0.486
RNASampler(20):
0.069
Positive Predictive Value
RNAfold:
0.485
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs HotKnots
Matthews Correlation Coefficient
RNAfold:
0.485
HotKnots:
0.212
Sensitivity
RNAfold:
0.486
HotKnots:
0.069
Positive Predictive Value
RNAfold:
0.485
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Mastr(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Mastr(20):
0.187
Sensitivity
RNAfold:
0.486
Mastr(20):
0.044
Positive Predictive Value
RNAfold:
0.485
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Cylofold
Matthews Correlation Coefficient
RNAfold:
0.485
Cylofold:
0.178
Sensitivity
RNAfold:
0.486
Cylofold:
0.053
Positive Predictive Value
RNAfold:
0.485
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Pknots
Matthews Correlation Coefficient
RNAfold:
0.485
Pknots:
0.165
Sensitivity
RNAfold:
0.486
Pknots:
0.041
Positive Predictive Value
RNAfold:
0.485
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Alterna
Matthews Correlation Coefficient
RNAfold:
0.485
Alterna:
0.145
Sensitivity
RNAfold:
0.486
Alterna:
0.036
Positive Predictive Value
RNAfold:
0.485
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs MCFold
Matthews Correlation Coefficient
RNAfold:
0.485
MCFold:
0.138
Sensitivity
RNAfold:
0.486
MCFold:
0.044
Positive Predictive Value
RNAfold:
0.485
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RDfolder
Matthews Correlation Coefficient
RNAfold:
0.485
RDfolder:
0.138
Sensitivity
RNAfold:
0.486
RDfolder:
0.031
Positive Predictive Value
RNAfold:
0.485
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Murlet(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Murlet(seed):
0.110
Sensitivity
RNAfold:
0.486
Murlet(seed):
0.015
Positive Predictive Value
RNAfold:
0.485
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs CMfinder(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
CMfinder(seed):
0.111
Sensitivity
RNAfold:
0.486
CMfinder(seed):
0.016
Positive Predictive Value
RNAfold:
0.485
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs TurboFold(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
TurboFold(seed):
0.106
Sensitivity
RNAfold:
0.486
TurboFold(seed):
0.014
Positive Predictive Value
RNAfold:
0.485
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs PPfold(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
PPfold(seed):
0.097
Sensitivity
RNAfold:
0.486
PPfold(seed):
0.012
Positive Predictive Value
RNAfold:
0.485
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Carnac(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Carnac(seed):
0.093
Sensitivity
RNAfold:
0.486
Carnac(seed):
0.010
Positive Predictive Value
RNAfold:
0.485
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs RNASampler(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
RNASampler(seed):
0.088
Sensitivity
RNAfold:
0.486
RNASampler(seed):
0.009
Positive Predictive Value
RNAfold:
0.485
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Mastr(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Mastr(seed):
0.072
Sensitivity
RNAfold:
0.486
Mastr(seed):
0.006
Positive Predictive Value
RNAfold:
0.485
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs NanoFolder
Matthews Correlation Coefficient
RNAfold:
0.485
NanoFolder:
0.068
Sensitivity
RNAfold:
0.486
NanoFolder:
0.027
Positive Predictive Value
RNAfold:
0.485
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAfold vs Multilign(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Multilign(seed):
0.059
Sensitivity
RNAfold:
0.486
Multilign(seed):
0.005
Positive Predictive Value
RNAfold:
0.485
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PknotsRG |
1242
ContextFold vs PknotsRG
Matthews Correlation Coefficient
ContextFold:
0.754
PknotsRG:
0.474
Sensitivity
ContextFold:
0.717
PknotsRG:
0.440
Positive Predictive Value
ContextFold:
0.794
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PknotsRG:
0.474
Sensitivity
PETfold_pre2.0(seed):
0.667
PknotsRG:
0.440
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PknotsRG:
0.474
Sensitivity
CentroidAlifold(seed):
0.414
PknotsRG:
0.440
Positive Predictive Value
CentroidAlifold(seed):
0.901
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs PknotsRG
Matthews Correlation Coefficient
IPknot:
0.607
PknotsRG:
0.474
Sensitivity
IPknot:
0.551
PknotsRG:
0.440
Positive Predictive Value
IPknot:
0.670
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs PknotsRG
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
PknotsRG:
0.474
Sensitivity
PETfold_pre2.0(20):
0.410
PknotsRG:
0.440
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs PknotsRG
Matthews Correlation Coefficient
CentroidFold:
0.571
PknotsRG:
0.474
Sensitivity
CentroidFold:
0.515
PknotsRG:
0.440
Positive Predictive Value
CentroidFold:
0.633
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs PknotsRG
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
PknotsRG:
0.474
Sensitivity
CentroidAlifold(20):
0.369
PknotsRG:
0.440
Positive Predictive Value
CentroidAlifold(20):
0.891
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs PknotsRG
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
PknotsRG:
0.474
Sensitivity
CentroidHomfold‑LAST:
0.359
PknotsRG:
0.440
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs PknotsRG
Matthews Correlation Coefficient
Contrafold:
0.556
PknotsRG:
0.474
Sensitivity
Contrafold:
0.542
PknotsRG:
0.440
Positive Predictive Value
Contrafold:
0.571
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs PknotsRG
Matthews Correlation Coefficient
MXScarna(seed):
0.536
PknotsRG:
0.474
Sensitivity
MXScarna(seed):
0.382
PknotsRG:
0.440
Positive Predictive Value
MXScarna(seed):
0.753
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs PknotsRG
Matthews Correlation Coefficient
RNAalifold(20):
0.532
PknotsRG:
0.474
Sensitivity
RNAalifold(20):
0.359
PknotsRG:
0.440
Positive Predictive Value
RNAalifold(20):
0.787
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs PknotsRG
Matthews Correlation Coefficient
Sfold:
0.527
PknotsRG:
0.474
Sensitivity
Sfold:
0.477
PknotsRG:
0.440
Positive Predictive Value
Sfold:
0.582
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs PknotsRG
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
PknotsRG:
0.474
Sensitivity
RNAalifold(seed):
0.328
PknotsRG:
0.440
Positive Predictive Value
RNAalifold(seed):
0.835
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs PknotsRG
Matthews Correlation Coefficient
MaxExpect:
0.516
PknotsRG:
0.474
Sensitivity
MaxExpect:
0.505
PknotsRG:
0.440
Positive Predictive Value
MaxExpect:
0.528
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs PknotsRG
Matthews Correlation Coefficient
ProbKnot:
0.512
PknotsRG:
0.474
Sensitivity
ProbKnot:
0.508
PknotsRG:
0.440
Positive Predictive Value
ProbKnot:
0.517
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs PknotsRG
Matthews Correlation Coefficient
MXScarna(20):
0.500
PknotsRG:
0.474
Sensitivity
MXScarna(20):
0.353
PknotsRG:
0.440
Positive Predictive Value
MXScarna(20):
0.710
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs PknotsRG
Matthews Correlation Coefficient
UNAFold:
0.494
PknotsRG:
0.474
Sensitivity
UNAFold:
0.491
PknotsRG:
0.440
Positive Predictive Value
UNAFold:
0.498
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs PknotsRG
Matthews Correlation Coefficient
Fold:
0.493
PknotsRG:
0.474
Sensitivity
Fold:
0.496
PknotsRG:
0.440
Positive Predictive Value
Fold:
0.490
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs PknotsRG
Matthews Correlation Coefficient
RNAfold:
0.485
PknotsRG:
0.474
Sensitivity
RNAfold:
0.486
PknotsRG:
0.440
Positive Predictive Value
RNAfold:
0.485
PknotsRG:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PknotsRG vs CRWrnafold
Matthews Correlation Coefficient
PknotsRG:
0.474
CRWrnafold:
0.430
Sensitivity
PknotsRG:
0.440
CRWrnafold:
0.379
Positive Predictive Value
PknotsRG:
0.510
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNAsubopt
Matthews Correlation Coefficient
PknotsRG:
0.474
RNAsubopt:
0.422
Sensitivity
PknotsRG:
0.440
RNAsubopt:
0.344
Positive Predictive Value
PknotsRG:
0.510
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs TurboFold(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
TurboFold(20):
0.419
Sensitivity
PknotsRG:
0.440
TurboFold(20):
0.210
Positive Predictive Value
PknotsRG:
0.510
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs PPfold(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
PPfold(20):
0.416
Sensitivity
PknotsRG:
0.440
PPfold(20):
0.197
Positive Predictive Value
PknotsRG:
0.510
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Afold
Matthews Correlation Coefficient
PknotsRG:
0.474
Afold:
0.408
Sensitivity
PknotsRG:
0.440
Afold:
0.343
Positive Predictive Value
PknotsRG:
0.510
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Carnac(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Carnac(20):
0.401
Sensitivity
PknotsRG:
0.440
Carnac(20):
0.180
Positive Predictive Value
PknotsRG:
0.510
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs McQFold
Matthews Correlation Coefficient
PknotsRG:
0.474
McQFold:
0.389
Sensitivity
PknotsRG:
0.440
McQFold:
0.308
Positive Predictive Value
PknotsRG:
0.510
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RSpredict(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
RSpredict(20):
0.377
Sensitivity
PknotsRG:
0.440
RSpredict(20):
0.241
Positive Predictive Value
PknotsRG:
0.510
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNAshapes
Matthews Correlation Coefficient
PknotsRG:
0.474
RNAshapes:
0.367
Sensitivity
PknotsRG:
0.440
RNAshapes:
0.285
Positive Predictive Value
PknotsRG:
0.510
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNASLOpt
Matthews Correlation Coefficient
PknotsRG:
0.474
RNASLOpt:
0.364
Sensitivity
PknotsRG:
0.440
RNASLOpt:
0.261
Positive Predictive Value
PknotsRG:
0.510
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Multilign(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Multilign(20):
0.350
Sensitivity
PknotsRG:
0.440
Multilign(20):
0.161
Positive Predictive Value
PknotsRG:
0.510
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Murlet(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Murlet(20):
0.321
Sensitivity
PknotsRG:
0.440
Murlet(20):
0.131
Positive Predictive Value
PknotsRG:
0.510
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNAwolf
Matthews Correlation Coefficient
PknotsRG:
0.474
RNAwolf:
0.310
Sensitivity
PknotsRG:
0.440
RNAwolf:
0.261
Positive Predictive Value
PknotsRG:
0.510
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RSpredict(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
RSpredict(seed):
0.295
Sensitivity
PknotsRG:
0.440
RSpredict(seed):
0.146
Positive Predictive Value
PknotsRG:
0.510
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs CMfinder(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
CMfinder(20):
0.292
Sensitivity
PknotsRG:
0.440
CMfinder(20):
0.107
Positive Predictive Value
PknotsRG:
0.510
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Vsfold4
Matthews Correlation Coefficient
PknotsRG:
0.474
Vsfold4:
0.270
Sensitivity
PknotsRG:
0.440
Vsfold4:
0.198
Positive Predictive Value
PknotsRG:
0.510
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Vsfold5
Matthews Correlation Coefficient
PknotsRG:
0.474
Vsfold5:
0.260
Sensitivity
PknotsRG:
0.440
Vsfold5:
0.195
Positive Predictive Value
PknotsRG:
0.510
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNASampler(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
RNASampler(20):
0.246
Sensitivity
PknotsRG:
0.440
RNASampler(20):
0.069
Positive Predictive Value
PknotsRG:
0.510
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs HotKnots
Matthews Correlation Coefficient
PknotsRG:
0.474
HotKnots:
0.212
Sensitivity
PknotsRG:
0.440
HotKnots:
0.069
Positive Predictive Value
PknotsRG:
0.510
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Mastr(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Mastr(20):
0.187
Sensitivity
PknotsRG:
0.440
Mastr(20):
0.044
Positive Predictive Value
PknotsRG:
0.510
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Cylofold
Matthews Correlation Coefficient
PknotsRG:
0.474
Cylofold:
0.178
Sensitivity
PknotsRG:
0.440
Cylofold:
0.053
Positive Predictive Value
PknotsRG:
0.510
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Pknots
Matthews Correlation Coefficient
PknotsRG:
0.474
Pknots:
0.165
Sensitivity
PknotsRG:
0.440
Pknots:
0.041
Positive Predictive Value
PknotsRG:
0.510
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Alterna
Matthews Correlation Coefficient
PknotsRG:
0.474
Alterna:
0.145
Sensitivity
PknotsRG:
0.440
Alterna:
0.036
Positive Predictive Value
PknotsRG:
0.510
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs MCFold
Matthews Correlation Coefficient
PknotsRG:
0.474
MCFold:
0.138
Sensitivity
PknotsRG:
0.440
MCFold:
0.044
Positive Predictive Value
PknotsRG:
0.510
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RDfolder
Matthews Correlation Coefficient
PknotsRG:
0.474
RDfolder:
0.138
Sensitivity
PknotsRG:
0.440
RDfolder:
0.031
Positive Predictive Value
PknotsRG:
0.510
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Murlet(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Murlet(seed):
0.110
Sensitivity
PknotsRG:
0.440
Murlet(seed):
0.015
Positive Predictive Value
PknotsRG:
0.510
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs CMfinder(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
CMfinder(seed):
0.111
Sensitivity
PknotsRG:
0.440
CMfinder(seed):
0.016
Positive Predictive Value
PknotsRG:
0.510
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs TurboFold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
TurboFold(seed):
0.106
Sensitivity
PknotsRG:
0.440
TurboFold(seed):
0.014
Positive Predictive Value
PknotsRG:
0.510
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs PPfold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
PPfold(seed):
0.097
Sensitivity
PknotsRG:
0.440
PPfold(seed):
0.012
Positive Predictive Value
PknotsRG:
0.510
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Carnac(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Carnac(seed):
0.093
Sensitivity
PknotsRG:
0.440
Carnac(seed):
0.010
Positive Predictive Value
PknotsRG:
0.510
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs RNASampler(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
RNASampler(seed):
0.088
Sensitivity
PknotsRG:
0.440
RNASampler(seed):
0.009
Positive Predictive Value
PknotsRG:
0.510
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Mastr(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Mastr(seed):
0.072
Sensitivity
PknotsRG:
0.440
Mastr(seed):
0.006
Positive Predictive Value
PknotsRG:
0.510
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs NanoFolder
Matthews Correlation Coefficient
PknotsRG:
0.474
NanoFolder:
0.068
Sensitivity
PknotsRG:
0.440
NanoFolder:
0.027
Positive Predictive Value
PknotsRG:
0.510
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PknotsRG vs Multilign(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Multilign(seed):
0.059
Sensitivity
PknotsRG:
0.440
Multilign(seed):
0.005
Positive Predictive Value
PknotsRG:
0.510
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CRWrnafold |
1242
ContextFold vs CRWrnafold
Matthews Correlation Coefficient
ContextFold:
0.754
CRWrnafold:
0.430
Sensitivity
ContextFold:
0.717
CRWrnafold:
0.379
Positive Predictive Value
ContextFold:
0.794
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CRWrnafold:
0.430
Sensitivity
PETfold_pre2.0(seed):
0.667
CRWrnafold:
0.379
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CRWrnafold:
0.430
Sensitivity
CentroidAlifold(seed):
0.414
CRWrnafold:
0.379
Positive Predictive Value
CentroidAlifold(seed):
0.901
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs CRWrnafold
Matthews Correlation Coefficient
IPknot:
0.607
CRWrnafold:
0.430
Sensitivity
IPknot:
0.551
CRWrnafold:
0.379
Positive Predictive Value
IPknot:
0.670
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs CRWrnafold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CRWrnafold:
0.430
Sensitivity
PETfold_pre2.0(20):
0.410
CRWrnafold:
0.379
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs CRWrnafold
Matthews Correlation Coefficient
CentroidFold:
0.571
CRWrnafold:
0.430
Sensitivity
CentroidFold:
0.515
CRWrnafold:
0.379
Positive Predictive Value
CentroidFold:
0.633
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs CRWrnafold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CRWrnafold:
0.430
Sensitivity
CentroidAlifold(20):
0.369
CRWrnafold:
0.379
Positive Predictive Value
CentroidAlifold(20):
0.891
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs CRWrnafold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
CRWrnafold:
0.430
Sensitivity
CentroidHomfold‑LAST:
0.359
CRWrnafold:
0.379
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs CRWrnafold
Matthews Correlation Coefficient
Contrafold:
0.556
CRWrnafold:
0.430
Sensitivity
Contrafold:
0.542
CRWrnafold:
0.379
Positive Predictive Value
Contrafold:
0.571
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
CRWrnafold:
0.430
Sensitivity
MXScarna(seed):
0.382
CRWrnafold:
0.379
Positive Predictive Value
MXScarna(seed):
0.753
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
CRWrnafold:
0.430
Sensitivity
RNAalifold(20):
0.359
CRWrnafold:
0.379
Positive Predictive Value
RNAalifold(20):
0.787
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs CRWrnafold
Matthews Correlation Coefficient
Sfold:
0.527
CRWrnafold:
0.430
Sensitivity
Sfold:
0.477
CRWrnafold:
0.379
Positive Predictive Value
Sfold:
0.582
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs CRWrnafold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
CRWrnafold:
0.430
Sensitivity
RNAalifold(seed):
0.328
CRWrnafold:
0.379
Positive Predictive Value
RNAalifold(seed):
0.835
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs CRWrnafold
Matthews Correlation Coefficient
MaxExpect:
0.516
CRWrnafold:
0.430
Sensitivity
MaxExpect:
0.505
CRWrnafold:
0.379
Positive Predictive Value
MaxExpect:
0.528
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs CRWrnafold
Matthews Correlation Coefficient
ProbKnot:
0.512
CRWrnafold:
0.430
Sensitivity
ProbKnot:
0.508
CRWrnafold:
0.379
Positive Predictive Value
ProbKnot:
0.517
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs CRWrnafold
Matthews Correlation Coefficient
MXScarna(20):
0.500
CRWrnafold:
0.430
Sensitivity
MXScarna(20):
0.353
CRWrnafold:
0.379
Positive Predictive Value
MXScarna(20):
0.710
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs CRWrnafold
Matthews Correlation Coefficient
UNAFold:
0.494
CRWrnafold:
0.430
Sensitivity
UNAFold:
0.491
CRWrnafold:
0.379
Positive Predictive Value
UNAFold:
0.498
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs CRWrnafold
Matthews Correlation Coefficient
Fold:
0.493
CRWrnafold:
0.430
Sensitivity
Fold:
0.496
CRWrnafold:
0.379
Positive Predictive Value
Fold:
0.490
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs CRWrnafold
Matthews Correlation Coefficient
RNAfold:
0.485
CRWrnafold:
0.430
Sensitivity
RNAfold:
0.486
CRWrnafold:
0.379
Positive Predictive Value
RNAfold:
0.485
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs CRWrnafold
Matthews Correlation Coefficient
PknotsRG:
0.474
CRWrnafold:
0.430
Sensitivity
PknotsRG:
0.440
CRWrnafold:
0.379
Positive Predictive Value
PknotsRG:
0.510
CRWrnafold:
0.490
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
CRWrnafold vs RNAsubopt
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNAsubopt:
0.422
Sensitivity
CRWrnafold:
0.379
RNAsubopt:
0.344
Positive Predictive Value
CRWrnafold:
0.490
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs TurboFold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
TurboFold(20):
0.419
Sensitivity
CRWrnafold:
0.379
TurboFold(20):
0.210
Positive Predictive Value
CRWrnafold:
0.490
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs PPfold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
PPfold(20):
0.416
Sensitivity
CRWrnafold:
0.379
PPfold(20):
0.197
Positive Predictive Value
CRWrnafold:
0.490
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Afold
Matthews Correlation Coefficient
CRWrnafold:
0.430
Afold:
0.408
Sensitivity
CRWrnafold:
0.379
Afold:
0.343
Positive Predictive Value
CRWrnafold:
0.490
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Carnac(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Carnac(20):
0.401
Sensitivity
CRWrnafold:
0.379
Carnac(20):
0.180
Positive Predictive Value
CRWrnafold:
0.490
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs McQFold
Matthews Correlation Coefficient
CRWrnafold:
0.430
McQFold:
0.389
Sensitivity
CRWrnafold:
0.379
McQFold:
0.308
Positive Predictive Value
CRWrnafold:
0.490
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RSpredict(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RSpredict(20):
0.377
Sensitivity
CRWrnafold:
0.379
RSpredict(20):
0.241
Positive Predictive Value
CRWrnafold:
0.490
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNAshapes
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNAshapes:
0.367
Sensitivity
CRWrnafold:
0.379
RNAshapes:
0.285
Positive Predictive Value
CRWrnafold:
0.490
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNASLOpt
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNASLOpt:
0.364
Sensitivity
CRWrnafold:
0.379
RNASLOpt:
0.261
Positive Predictive Value
CRWrnafold:
0.490
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Multilign(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Multilign(20):
0.350
Sensitivity
CRWrnafold:
0.379
Multilign(20):
0.161
Positive Predictive Value
CRWrnafold:
0.490
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Murlet(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Murlet(20):
0.321
Sensitivity
CRWrnafold:
0.379
Murlet(20):
0.131
Positive Predictive Value
CRWrnafold:
0.490
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNAwolf
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNAwolf:
0.310
Sensitivity
CRWrnafold:
0.379
RNAwolf:
0.261
Positive Predictive Value
CRWrnafold:
0.490
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RSpredict(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RSpredict(seed):
0.295
Sensitivity
CRWrnafold:
0.379
RSpredict(seed):
0.146
Positive Predictive Value
CRWrnafold:
0.490
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs CMfinder(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
CMfinder(20):
0.292
Sensitivity
CRWrnafold:
0.379
CMfinder(20):
0.107
Positive Predictive Value
CRWrnafold:
0.490
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Vsfold4
Matthews Correlation Coefficient
CRWrnafold:
0.430
Vsfold4:
0.270
Sensitivity
CRWrnafold:
0.379
Vsfold4:
0.198
Positive Predictive Value
CRWrnafold:
0.490
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Vsfold5
Matthews Correlation Coefficient
CRWrnafold:
0.430
Vsfold5:
0.260
Sensitivity
CRWrnafold:
0.379
Vsfold5:
0.195
Positive Predictive Value
CRWrnafold:
0.490
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNASampler(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNASampler(20):
0.246
Sensitivity
CRWrnafold:
0.379
RNASampler(20):
0.069
Positive Predictive Value
CRWrnafold:
0.490
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs HotKnots
Matthews Correlation Coefficient
CRWrnafold:
0.430
HotKnots:
0.212
Sensitivity
CRWrnafold:
0.379
HotKnots:
0.069
Positive Predictive Value
CRWrnafold:
0.490
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Mastr(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Mastr(20):
0.187
Sensitivity
CRWrnafold:
0.379
Mastr(20):
0.044
Positive Predictive Value
CRWrnafold:
0.490
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Cylofold
Matthews Correlation Coefficient
CRWrnafold:
0.430
Cylofold:
0.178
Sensitivity
CRWrnafold:
0.379
Cylofold:
0.053
Positive Predictive Value
CRWrnafold:
0.490
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Pknots
Matthews Correlation Coefficient
CRWrnafold:
0.430
Pknots:
0.165
Sensitivity
CRWrnafold:
0.379
Pknots:
0.041
Positive Predictive Value
CRWrnafold:
0.490
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Alterna
Matthews Correlation Coefficient
CRWrnafold:
0.430
Alterna:
0.145
Sensitivity
CRWrnafold:
0.379
Alterna:
0.036
Positive Predictive Value
CRWrnafold:
0.490
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs MCFold
Matthews Correlation Coefficient
CRWrnafold:
0.430
MCFold:
0.138
Sensitivity
CRWrnafold:
0.379
MCFold:
0.044
Positive Predictive Value
CRWrnafold:
0.490
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RDfolder
Matthews Correlation Coefficient
CRWrnafold:
0.430
RDfolder:
0.138
Sensitivity
CRWrnafold:
0.379
RDfolder:
0.031
Positive Predictive Value
CRWrnafold:
0.490
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Murlet(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Murlet(seed):
0.110
Sensitivity
CRWrnafold:
0.379
Murlet(seed):
0.015
Positive Predictive Value
CRWrnafold:
0.490
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs CMfinder(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
CMfinder(seed):
0.111
Sensitivity
CRWrnafold:
0.379
CMfinder(seed):
0.016
Positive Predictive Value
CRWrnafold:
0.490
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs TurboFold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
TurboFold(seed):
0.106
Sensitivity
CRWrnafold:
0.379
TurboFold(seed):
0.014
Positive Predictive Value
CRWrnafold:
0.490
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs PPfold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
PPfold(seed):
0.097
Sensitivity
CRWrnafold:
0.379
PPfold(seed):
0.012
Positive Predictive Value
CRWrnafold:
0.490
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Carnac(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Carnac(seed):
0.093
Sensitivity
CRWrnafold:
0.379
Carnac(seed):
0.010
Positive Predictive Value
CRWrnafold:
0.490
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs RNASampler(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNASampler(seed):
0.088
Sensitivity
CRWrnafold:
0.379
RNASampler(seed):
0.009
Positive Predictive Value
CRWrnafold:
0.490
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Mastr(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Mastr(seed):
0.072
Sensitivity
CRWrnafold:
0.379
Mastr(seed):
0.006
Positive Predictive Value
CRWrnafold:
0.490
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs NanoFolder
Matthews Correlation Coefficient
CRWrnafold:
0.430
NanoFolder:
0.068
Sensitivity
CRWrnafold:
0.379
NanoFolder:
0.027
Positive Predictive Value
CRWrnafold:
0.490
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CRWrnafold vs Multilign(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Multilign(seed):
0.059
Sensitivity
CRWrnafold:
0.379
Multilign(seed):
0.005
Positive Predictive Value
CRWrnafold:
0.490
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAsubopt |
1242
ContextFold vs RNAsubopt
Matthews Correlation Coefficient
ContextFold:
0.754
RNAsubopt:
0.422
Sensitivity
ContextFold:
0.717
RNAsubopt:
0.344
Positive Predictive Value
ContextFold:
0.794
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAsubopt:
0.422
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAsubopt:
0.344
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAsubopt:
0.422
Sensitivity
CentroidAlifold(seed):
0.414
RNAsubopt:
0.344
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNAsubopt
Matthews Correlation Coefficient
IPknot:
0.607
RNAsubopt:
0.422
Sensitivity
IPknot:
0.551
RNAsubopt:
0.344
Positive Predictive Value
IPknot:
0.670
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNAsubopt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAsubopt:
0.422
Sensitivity
PETfold_pre2.0(20):
0.410
RNAsubopt:
0.344
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNAsubopt
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAsubopt:
0.422
Sensitivity
CentroidFold:
0.515
RNAsubopt:
0.344
Positive Predictive Value
CentroidFold:
0.633
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNAsubopt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAsubopt:
0.422
Sensitivity
CentroidAlifold(20):
0.369
RNAsubopt:
0.344
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNAsubopt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAsubopt:
0.422
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAsubopt:
0.344
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNAsubopt
Matthews Correlation Coefficient
Contrafold:
0.556
RNAsubopt:
0.422
Sensitivity
Contrafold:
0.542
RNAsubopt:
0.344
Positive Predictive Value
Contrafold:
0.571
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAsubopt:
0.422
Sensitivity
MXScarna(seed):
0.382
RNAsubopt:
0.344
Positive Predictive Value
MXScarna(seed):
0.753
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAsubopt:
0.422
Sensitivity
RNAalifold(20):
0.359
RNAsubopt:
0.344
Positive Predictive Value
RNAalifold(20):
0.787
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNAsubopt
Matthews Correlation Coefficient
Sfold:
0.527
RNAsubopt:
0.422
Sensitivity
Sfold:
0.477
RNAsubopt:
0.344
Positive Predictive Value
Sfold:
0.582
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNAsubopt
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAsubopt:
0.422
Sensitivity
RNAalifold(seed):
0.328
RNAsubopt:
0.344
Positive Predictive Value
RNAalifold(seed):
0.835
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNAsubopt
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAsubopt:
0.422
Sensitivity
MaxExpect:
0.505
RNAsubopt:
0.344
Positive Predictive Value
MaxExpect:
0.528
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNAsubopt
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAsubopt:
0.422
Sensitivity
ProbKnot:
0.508
RNAsubopt:
0.344
Positive Predictive Value
ProbKnot:
0.517
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNAsubopt
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAsubopt:
0.422
Sensitivity
MXScarna(20):
0.353
RNAsubopt:
0.344
Positive Predictive Value
MXScarna(20):
0.710
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNAsubopt
Matthews Correlation Coefficient
UNAFold:
0.494
RNAsubopt:
0.422
Sensitivity
UNAFold:
0.491
RNAsubopt:
0.344
Positive Predictive Value
UNAFold:
0.498
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNAsubopt
Matthews Correlation Coefficient
Fold:
0.493
RNAsubopt:
0.422
Sensitivity
Fold:
0.496
RNAsubopt:
0.344
Positive Predictive Value
Fold:
0.490
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RNAsubopt
Matthews Correlation Coefficient
RNAfold:
0.485
RNAsubopt:
0.422
Sensitivity
RNAfold:
0.486
RNAsubopt:
0.344
Positive Predictive Value
RNAfold:
0.485
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RNAsubopt
Matthews Correlation Coefficient
PknotsRG:
0.474
RNAsubopt:
0.422
Sensitivity
PknotsRG:
0.440
RNAsubopt:
0.344
Positive Predictive Value
PknotsRG:
0.510
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RNAsubopt
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNAsubopt:
0.422
Sensitivity
CRWrnafold:
0.379
RNAsubopt:
0.344
Positive Predictive Value
CRWrnafold:
0.490
RNAsubopt:
0.519
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAsubopt vs TurboFold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
TurboFold(20):
0.419
Sensitivity
RNAsubopt:
0.344
TurboFold(20):
0.210
Positive Predictive Value
RNAsubopt:
0.519
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.80472700427e-05
|
+
RNAsubopt vs PPfold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
PPfold(20):
0.416
Sensitivity
RNAsubopt:
0.344
PPfold(20):
0.197
Positive Predictive Value
RNAsubopt:
0.519
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.48494289621e-08
|
+
RNAsubopt vs Afold
Matthews Correlation Coefficient
RNAsubopt:
0.422
Afold:
0.408
Sensitivity
RNAsubopt:
0.344
Afold:
0.343
Positive Predictive Value
RNAsubopt:
0.519
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Carnac(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Carnac(20):
0.401
Sensitivity
RNAsubopt:
0.344
Carnac(20):
0.180
Positive Predictive Value
RNAsubopt:
0.519
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs McQFold
Matthews Correlation Coefficient
RNAsubopt:
0.422
McQFold:
0.389
Sensitivity
RNAsubopt:
0.344
McQFold:
0.308
Positive Predictive Value
RNAsubopt:
0.519
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RSpredict(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RSpredict(20):
0.377
Sensitivity
RNAsubopt:
0.344
RSpredict(20):
0.241
Positive Predictive Value
RNAsubopt:
0.519
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNAshapes
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNAshapes:
0.367
Sensitivity
RNAsubopt:
0.344
RNAshapes:
0.285
Positive Predictive Value
RNAsubopt:
0.519
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNASLOpt
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNASLOpt:
0.364
Sensitivity
RNAsubopt:
0.344
RNASLOpt:
0.261
Positive Predictive Value
RNAsubopt:
0.519
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Multilign(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Multilign(20):
0.350
Sensitivity
RNAsubopt:
0.344
Multilign(20):
0.161
Positive Predictive Value
RNAsubopt:
0.519
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Murlet(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Murlet(20):
0.321
Sensitivity
RNAsubopt:
0.344
Murlet(20):
0.131
Positive Predictive Value
RNAsubopt:
0.519
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNAwolf
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNAwolf:
0.310
Sensitivity
RNAsubopt:
0.344
RNAwolf:
0.261
Positive Predictive Value
RNAsubopt:
0.519
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RSpredict(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RSpredict(seed):
0.295
Sensitivity
RNAsubopt:
0.344
RSpredict(seed):
0.146
Positive Predictive Value
RNAsubopt:
0.519
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs CMfinder(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
CMfinder(20):
0.292
Sensitivity
RNAsubopt:
0.344
CMfinder(20):
0.107
Positive Predictive Value
RNAsubopt:
0.519
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Vsfold4
Matthews Correlation Coefficient
RNAsubopt:
0.422
Vsfold4:
0.270
Sensitivity
RNAsubopt:
0.344
Vsfold4:
0.198
Positive Predictive Value
RNAsubopt:
0.519
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Vsfold5
Matthews Correlation Coefficient
RNAsubopt:
0.422
Vsfold5:
0.260
Sensitivity
RNAsubopt:
0.344
Vsfold5:
0.195
Positive Predictive Value
RNAsubopt:
0.519
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNASampler(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNASampler(20):
0.246
Sensitivity
RNAsubopt:
0.344
RNASampler(20):
0.069
Positive Predictive Value
RNAsubopt:
0.519
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs HotKnots
Matthews Correlation Coefficient
RNAsubopt:
0.422
HotKnots:
0.212
Sensitivity
RNAsubopt:
0.344
HotKnots:
0.069
Positive Predictive Value
RNAsubopt:
0.519
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Mastr(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Mastr(20):
0.187
Sensitivity
RNAsubopt:
0.344
Mastr(20):
0.044
Positive Predictive Value
RNAsubopt:
0.519
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Cylofold
Matthews Correlation Coefficient
RNAsubopt:
0.422
Cylofold:
0.178
Sensitivity
RNAsubopt:
0.344
Cylofold:
0.053
Positive Predictive Value
RNAsubopt:
0.519
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Pknots
Matthews Correlation Coefficient
RNAsubopt:
0.422
Pknots:
0.165
Sensitivity
RNAsubopt:
0.344
Pknots:
0.041
Positive Predictive Value
RNAsubopt:
0.519
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Alterna
Matthews Correlation Coefficient
RNAsubopt:
0.422
Alterna:
0.145
Sensitivity
RNAsubopt:
0.344
Alterna:
0.036
Positive Predictive Value
RNAsubopt:
0.519
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs MCFold
Matthews Correlation Coefficient
RNAsubopt:
0.422
MCFold:
0.138
Sensitivity
RNAsubopt:
0.344
MCFold:
0.044
Positive Predictive Value
RNAsubopt:
0.519
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RDfolder
Matthews Correlation Coefficient
RNAsubopt:
0.422
RDfolder:
0.138
Sensitivity
RNAsubopt:
0.344
RDfolder:
0.031
Positive Predictive Value
RNAsubopt:
0.519
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Murlet(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Murlet(seed):
0.110
Sensitivity
RNAsubopt:
0.344
Murlet(seed):
0.015
Positive Predictive Value
RNAsubopt:
0.519
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs CMfinder(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
CMfinder(seed):
0.111
Sensitivity
RNAsubopt:
0.344
CMfinder(seed):
0.016
Positive Predictive Value
RNAsubopt:
0.519
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs TurboFold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
TurboFold(seed):
0.106
Sensitivity
RNAsubopt:
0.344
TurboFold(seed):
0.014
Positive Predictive Value
RNAsubopt:
0.519
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs PPfold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
PPfold(seed):
0.097
Sensitivity
RNAsubopt:
0.344
PPfold(seed):
0.012
Positive Predictive Value
RNAsubopt:
0.519
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Carnac(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Carnac(seed):
0.093
Sensitivity
RNAsubopt:
0.344
Carnac(seed):
0.010
Positive Predictive Value
RNAsubopt:
0.519
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs RNASampler(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNASampler(seed):
0.088
Sensitivity
RNAsubopt:
0.344
RNASampler(seed):
0.009
Positive Predictive Value
RNAsubopt:
0.519
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Mastr(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Mastr(seed):
0.072
Sensitivity
RNAsubopt:
0.344
Mastr(seed):
0.006
Positive Predictive Value
RNAsubopt:
0.519
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs NanoFolder
Matthews Correlation Coefficient
RNAsubopt:
0.422
NanoFolder:
0.068
Sensitivity
RNAsubopt:
0.344
NanoFolder:
0.027
Positive Predictive Value
RNAsubopt:
0.519
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAsubopt vs Multilign(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Multilign(seed):
0.059
Sensitivity
RNAsubopt:
0.344
Multilign(seed):
0.005
Positive Predictive Value
RNAsubopt:
0.519
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| TurboFold(20) |
1242
ContextFold vs TurboFold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
TurboFold(20):
0.419
Sensitivity
ContextFold:
0.717
TurboFold(20):
0.210
Positive Predictive Value
ContextFold:
0.794
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
TurboFold(20):
0.419
Sensitivity
PETfold_pre2.0(seed):
0.667
TurboFold(20):
0.210
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
TurboFold(20):
0.419
Sensitivity
CentroidAlifold(seed):
0.414
TurboFold(20):
0.210
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs TurboFold(20)
Matthews Correlation Coefficient
IPknot:
0.607
TurboFold(20):
0.419
Sensitivity
IPknot:
0.551
TurboFold(20):
0.210
Positive Predictive Value
IPknot:
0.670
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs TurboFold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
TurboFold(20):
0.419
Sensitivity
PETfold_pre2.0(20):
0.410
TurboFold(20):
0.210
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs TurboFold(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
TurboFold(20):
0.419
Sensitivity
CentroidFold:
0.515
TurboFold(20):
0.210
Positive Predictive Value
CentroidFold:
0.633
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
TurboFold(20):
0.419
Sensitivity
CentroidAlifold(20):
0.369
TurboFold(20):
0.210
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs TurboFold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
TurboFold(20):
0.419
Sensitivity
CentroidHomfold‑LAST:
0.359
TurboFold(20):
0.210
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs TurboFold(20)
Matthews Correlation Coefficient
Contrafold:
0.556
TurboFold(20):
0.419
Sensitivity
Contrafold:
0.542
TurboFold(20):
0.210
Positive Predictive Value
Contrafold:
0.571
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
TurboFold(20):
0.419
Sensitivity
MXScarna(seed):
0.382
TurboFold(20):
0.210
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
TurboFold(20):
0.419
Sensitivity
RNAalifold(20):
0.359
TurboFold(20):
0.210
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs TurboFold(20)
Matthews Correlation Coefficient
Sfold:
0.527
TurboFold(20):
0.419
Sensitivity
Sfold:
0.477
TurboFold(20):
0.210
Positive Predictive Value
Sfold:
0.582
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs TurboFold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
TurboFold(20):
0.419
Sensitivity
RNAalifold(seed):
0.328
TurboFold(20):
0.210
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs TurboFold(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
TurboFold(20):
0.419
Sensitivity
MaxExpect:
0.505
TurboFold(20):
0.210
Positive Predictive Value
MaxExpect:
0.528
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs TurboFold(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
TurboFold(20):
0.419
Sensitivity
ProbKnot:
0.508
TurboFold(20):
0.210
Positive Predictive Value
ProbKnot:
0.517
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs TurboFold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
TurboFold(20):
0.419
Sensitivity
MXScarna(20):
0.353
TurboFold(20):
0.210
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs TurboFold(20)
Matthews Correlation Coefficient
UNAFold:
0.494
TurboFold(20):
0.419
Sensitivity
UNAFold:
0.491
TurboFold(20):
0.210
Positive Predictive Value
UNAFold:
0.498
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs TurboFold(20)
Matthews Correlation Coefficient
Fold:
0.493
TurboFold(20):
0.419
Sensitivity
Fold:
0.496
TurboFold(20):
0.210
Positive Predictive Value
Fold:
0.490
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs TurboFold(20)
Matthews Correlation Coefficient
RNAfold:
0.485
TurboFold(20):
0.419
Sensitivity
RNAfold:
0.486
TurboFold(20):
0.210
Positive Predictive Value
RNAfold:
0.485
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs TurboFold(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
TurboFold(20):
0.419
Sensitivity
PknotsRG:
0.440
TurboFold(20):
0.210
Positive Predictive Value
PknotsRG:
0.510
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs TurboFold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
TurboFold(20):
0.419
Sensitivity
CRWrnafold:
0.379
TurboFold(20):
0.210
Positive Predictive Value
CRWrnafold:
0.490
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs TurboFold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
TurboFold(20):
0.419
Sensitivity
RNAsubopt:
0.344
TurboFold(20):
0.210
Positive Predictive Value
RNAsubopt:
0.519
TurboFold(20):
0.837
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.80472700427e-05
|
|
+
TurboFold(20) vs PPfold(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
PPfold(20):
0.416
Sensitivity
TurboFold(20):
0.210
PPfold(20):
0.197
Positive Predictive Value
TurboFold(20):
0.837
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
+
TurboFold(20) vs Afold
Matthews Correlation Coefficient
TurboFold(20):
0.419
Afold:
0.408
Sensitivity
TurboFold(20):
0.210
Afold:
0.343
Positive Predictive Value
TurboFold(20):
0.837
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Carnac(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Carnac(20):
0.401
Sensitivity
TurboFold(20):
0.210
Carnac(20):
0.180
Positive Predictive Value
TurboFold(20):
0.837
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs McQFold
Matthews Correlation Coefficient
TurboFold(20):
0.419
McQFold:
0.389
Sensitivity
TurboFold(20):
0.210
McQFold:
0.308
Positive Predictive Value
TurboFold(20):
0.837
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RSpredict(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RSpredict(20):
0.377
Sensitivity
TurboFold(20):
0.210
RSpredict(20):
0.241
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNAshapes
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNAshapes:
0.367
Sensitivity
TurboFold(20):
0.210
RNAshapes:
0.285
Positive Predictive Value
TurboFold(20):
0.837
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNASLOpt
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNASLOpt:
0.364
Sensitivity
TurboFold(20):
0.210
RNASLOpt:
0.261
Positive Predictive Value
TurboFold(20):
0.837
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Multilign(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Multilign(20):
0.350
Sensitivity
TurboFold(20):
0.210
Multilign(20):
0.161
Positive Predictive Value
TurboFold(20):
0.837
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Murlet(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Murlet(20):
0.321
Sensitivity
TurboFold(20):
0.210
Murlet(20):
0.131
Positive Predictive Value
TurboFold(20):
0.837
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNAwolf
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNAwolf:
0.310
Sensitivity
TurboFold(20):
0.210
RNAwolf:
0.261
Positive Predictive Value
TurboFold(20):
0.837
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RSpredict(seed):
0.295
Sensitivity
TurboFold(20):
0.210
RSpredict(seed):
0.146
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs CMfinder(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
CMfinder(20):
0.292
Sensitivity
TurboFold(20):
0.210
CMfinder(20):
0.107
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Vsfold4
Matthews Correlation Coefficient
TurboFold(20):
0.419
Vsfold4:
0.270
Sensitivity
TurboFold(20):
0.210
Vsfold4:
0.198
Positive Predictive Value
TurboFold(20):
0.837
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Vsfold5
Matthews Correlation Coefficient
TurboFold(20):
0.419
Vsfold5:
0.260
Sensitivity
TurboFold(20):
0.210
Vsfold5:
0.195
Positive Predictive Value
TurboFold(20):
0.837
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNASampler(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNASampler(20):
0.246
Sensitivity
TurboFold(20):
0.210
RNASampler(20):
0.069
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs HotKnots
Matthews Correlation Coefficient
TurboFold(20):
0.419
HotKnots:
0.212
Sensitivity
TurboFold(20):
0.210
HotKnots:
0.069
Positive Predictive Value
TurboFold(20):
0.837
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Mastr(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Mastr(20):
0.187
Sensitivity
TurboFold(20):
0.210
Mastr(20):
0.044
Positive Predictive Value
TurboFold(20):
0.837
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Cylofold
Matthews Correlation Coefficient
TurboFold(20):
0.419
Cylofold:
0.178
Sensitivity
TurboFold(20):
0.210
Cylofold:
0.053
Positive Predictive Value
TurboFold(20):
0.837
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Pknots
Matthews Correlation Coefficient
TurboFold(20):
0.419
Pknots:
0.165
Sensitivity
TurboFold(20):
0.210
Pknots:
0.041
Positive Predictive Value
TurboFold(20):
0.837
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Alterna
Matthews Correlation Coefficient
TurboFold(20):
0.419
Alterna:
0.145
Sensitivity
TurboFold(20):
0.210
Alterna:
0.036
Positive Predictive Value
TurboFold(20):
0.837
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs MCFold
Matthews Correlation Coefficient
TurboFold(20):
0.419
MCFold:
0.138
Sensitivity
TurboFold(20):
0.210
MCFold:
0.044
Positive Predictive Value
TurboFold(20):
0.837
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RDfolder
Matthews Correlation Coefficient
TurboFold(20):
0.419
RDfolder:
0.138
Sensitivity
TurboFold(20):
0.210
RDfolder:
0.031
Positive Predictive Value
TurboFold(20):
0.837
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Murlet(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Murlet(seed):
0.110
Sensitivity
TurboFold(20):
0.210
Murlet(seed):
0.015
Positive Predictive Value
TurboFold(20):
0.837
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
CMfinder(seed):
0.111
Sensitivity
TurboFold(20):
0.210
CMfinder(seed):
0.016
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
TurboFold(seed):
0.106
Sensitivity
TurboFold(20):
0.210
TurboFold(seed):
0.014
Positive Predictive Value
TurboFold(20):
0.837
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
PPfold(seed):
0.097
Sensitivity
TurboFold(20):
0.210
PPfold(seed):
0.012
Positive Predictive Value
TurboFold(20):
0.837
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Carnac(seed):
0.093
Sensitivity
TurboFold(20):
0.210
Carnac(seed):
0.010
Positive Predictive Value
TurboFold(20):
0.837
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNASampler(seed):
0.088
Sensitivity
TurboFold(20):
0.210
RNASampler(seed):
0.009
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Mastr(seed):
0.072
Sensitivity
TurboFold(20):
0.210
Mastr(seed):
0.006
Positive Predictive Value
TurboFold(20):
0.837
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(20):
0.419
NanoFolder:
0.068
Sensitivity
TurboFold(20):
0.210
NanoFolder:
0.027
Positive Predictive Value
TurboFold(20):
0.837
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(20) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Multilign(seed):
0.059
Sensitivity
TurboFold(20):
0.210
Multilign(seed):
0.005
Positive Predictive Value
TurboFold(20):
0.837
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PPfold(20) |
1242
ContextFold vs PPfold(20)
Matthews Correlation Coefficient
ContextFold:
0.754
PPfold(20):
0.416
Sensitivity
ContextFold:
0.717
PPfold(20):
0.197
Positive Predictive Value
ContextFold:
0.794
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PPfold(20):
0.416
Sensitivity
PETfold_pre2.0(seed):
0.667
PPfold(20):
0.197
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PPfold(20):
0.416
Sensitivity
CentroidAlifold(seed):
0.414
PPfold(20):
0.197
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs PPfold(20)
Matthews Correlation Coefficient
IPknot:
0.607
PPfold(20):
0.416
Sensitivity
IPknot:
0.551
PPfold(20):
0.197
Positive Predictive Value
IPknot:
0.670
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs PPfold(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
PPfold(20):
0.416
Sensitivity
PETfold_pre2.0(20):
0.410
PPfold(20):
0.197
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs PPfold(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
PPfold(20):
0.416
Sensitivity
CentroidFold:
0.515
PPfold(20):
0.197
Positive Predictive Value
CentroidFold:
0.633
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs PPfold(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
PPfold(20):
0.416
Sensitivity
CentroidAlifold(20):
0.369
PPfold(20):
0.197
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs PPfold(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
PPfold(20):
0.416
Sensitivity
CentroidHomfold‑LAST:
0.359
PPfold(20):
0.197
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs PPfold(20)
Matthews Correlation Coefficient
Contrafold:
0.556
PPfold(20):
0.416
Sensitivity
Contrafold:
0.542
PPfold(20):
0.197
Positive Predictive Value
Contrafold:
0.571
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
PPfold(20):
0.416
Sensitivity
MXScarna(seed):
0.382
PPfold(20):
0.197
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
PPfold(20):
0.416
Sensitivity
RNAalifold(20):
0.359
PPfold(20):
0.197
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs PPfold(20)
Matthews Correlation Coefficient
Sfold:
0.527
PPfold(20):
0.416
Sensitivity
Sfold:
0.477
PPfold(20):
0.197
Positive Predictive Value
Sfold:
0.582
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs PPfold(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
PPfold(20):
0.416
Sensitivity
RNAalifold(seed):
0.328
PPfold(20):
0.197
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs PPfold(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
PPfold(20):
0.416
Sensitivity
MaxExpect:
0.505
PPfold(20):
0.197
Positive Predictive Value
MaxExpect:
0.528
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs PPfold(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
PPfold(20):
0.416
Sensitivity
ProbKnot:
0.508
PPfold(20):
0.197
Positive Predictive Value
ProbKnot:
0.517
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs PPfold(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
PPfold(20):
0.416
Sensitivity
MXScarna(20):
0.353
PPfold(20):
0.197
Positive Predictive Value
MXScarna(20):
0.710
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs PPfold(20)
Matthews Correlation Coefficient
UNAFold:
0.494
PPfold(20):
0.416
Sensitivity
UNAFold:
0.491
PPfold(20):
0.197
Positive Predictive Value
UNAFold:
0.498
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs PPfold(20)
Matthews Correlation Coefficient
Fold:
0.493
PPfold(20):
0.416
Sensitivity
Fold:
0.496
PPfold(20):
0.197
Positive Predictive Value
Fold:
0.490
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs PPfold(20)
Matthews Correlation Coefficient
RNAfold:
0.485
PPfold(20):
0.416
Sensitivity
RNAfold:
0.486
PPfold(20):
0.197
Positive Predictive Value
RNAfold:
0.485
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs PPfold(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
PPfold(20):
0.416
Sensitivity
PknotsRG:
0.440
PPfold(20):
0.197
Positive Predictive Value
PknotsRG:
0.510
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs PPfold(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
PPfold(20):
0.416
Sensitivity
CRWrnafold:
0.379
PPfold(20):
0.197
Positive Predictive Value
CRWrnafold:
0.490
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs PPfold(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
PPfold(20):
0.416
Sensitivity
RNAsubopt:
0.344
PPfold(20):
0.197
Positive Predictive Value
RNAsubopt:
0.519
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.48494289621e-08
|
1242
TurboFold(20) vs PPfold(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
PPfold(20):
0.416
Sensitivity
TurboFold(20):
0.210
PPfold(20):
0.197
Positive Predictive Value
TurboFold(20):
0.837
PPfold(20):
0.879
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.85233844192e-08
|
|
+
PPfold(20) vs Afold
Matthews Correlation Coefficient
PPfold(20):
0.416
Afold:
0.408
Sensitivity
PPfold(20):
0.197
Afold:
0.343
Positive Predictive Value
PPfold(20):
0.879
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 7.5939721025e-08
|
+
PPfold(20) vs Carnac(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Carnac(20):
0.401
Sensitivity
PPfold(20):
0.197
Carnac(20):
0.180
Positive Predictive Value
PPfold(20):
0.879
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs McQFold
Matthews Correlation Coefficient
PPfold(20):
0.416
McQFold:
0.389
Sensitivity
PPfold(20):
0.197
McQFold:
0.308
Positive Predictive Value
PPfold(20):
0.879
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RSpredict(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
RSpredict(20):
0.377
Sensitivity
PPfold(20):
0.197
RSpredict(20):
0.241
Positive Predictive Value
PPfold(20):
0.879
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNAshapes
Matthews Correlation Coefficient
PPfold(20):
0.416
RNAshapes:
0.367
Sensitivity
PPfold(20):
0.197
RNAshapes:
0.285
Positive Predictive Value
PPfold(20):
0.879
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNASLOpt
Matthews Correlation Coefficient
PPfold(20):
0.416
RNASLOpt:
0.364
Sensitivity
PPfold(20):
0.197
RNASLOpt:
0.261
Positive Predictive Value
PPfold(20):
0.879
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Multilign(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Multilign(20):
0.350
Sensitivity
PPfold(20):
0.197
Multilign(20):
0.161
Positive Predictive Value
PPfold(20):
0.879
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Murlet(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Murlet(20):
0.321
Sensitivity
PPfold(20):
0.197
Murlet(20):
0.131
Positive Predictive Value
PPfold(20):
0.879
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNAwolf
Matthews Correlation Coefficient
PPfold(20):
0.416
RNAwolf:
0.310
Sensitivity
PPfold(20):
0.197
RNAwolf:
0.261
Positive Predictive Value
PPfold(20):
0.879
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
RSpredict(seed):
0.295
Sensitivity
PPfold(20):
0.197
RSpredict(seed):
0.146
Positive Predictive Value
PPfold(20):
0.879
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs CMfinder(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
CMfinder(20):
0.292
Sensitivity
PPfold(20):
0.197
CMfinder(20):
0.107
Positive Predictive Value
PPfold(20):
0.879
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Vsfold4
Matthews Correlation Coefficient
PPfold(20):
0.416
Vsfold4:
0.270
Sensitivity
PPfold(20):
0.197
Vsfold4:
0.198
Positive Predictive Value
PPfold(20):
0.879
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Vsfold5
Matthews Correlation Coefficient
PPfold(20):
0.416
Vsfold5:
0.260
Sensitivity
PPfold(20):
0.197
Vsfold5:
0.195
Positive Predictive Value
PPfold(20):
0.879
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNASampler(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
RNASampler(20):
0.246
Sensitivity
PPfold(20):
0.197
RNASampler(20):
0.069
Positive Predictive Value
PPfold(20):
0.879
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs HotKnots
Matthews Correlation Coefficient
PPfold(20):
0.416
HotKnots:
0.212
Sensitivity
PPfold(20):
0.197
HotKnots:
0.069
Positive Predictive Value
PPfold(20):
0.879
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Mastr(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Mastr(20):
0.187
Sensitivity
PPfold(20):
0.197
Mastr(20):
0.044
Positive Predictive Value
PPfold(20):
0.879
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Cylofold
Matthews Correlation Coefficient
PPfold(20):
0.416
Cylofold:
0.178
Sensitivity
PPfold(20):
0.197
Cylofold:
0.053
Positive Predictive Value
PPfold(20):
0.879
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Pknots
Matthews Correlation Coefficient
PPfold(20):
0.416
Pknots:
0.165
Sensitivity
PPfold(20):
0.197
Pknots:
0.041
Positive Predictive Value
PPfold(20):
0.879
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Alterna
Matthews Correlation Coefficient
PPfold(20):
0.416
Alterna:
0.145
Sensitivity
PPfold(20):
0.197
Alterna:
0.036
Positive Predictive Value
PPfold(20):
0.879
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs MCFold
Matthews Correlation Coefficient
PPfold(20):
0.416
MCFold:
0.138
Sensitivity
PPfold(20):
0.197
MCFold:
0.044
Positive Predictive Value
PPfold(20):
0.879
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RDfolder
Matthews Correlation Coefficient
PPfold(20):
0.416
RDfolder:
0.138
Sensitivity
PPfold(20):
0.197
RDfolder:
0.031
Positive Predictive Value
PPfold(20):
0.879
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Murlet(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Murlet(seed):
0.110
Sensitivity
PPfold(20):
0.197
Murlet(seed):
0.015
Positive Predictive Value
PPfold(20):
0.879
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
CMfinder(seed):
0.111
Sensitivity
PPfold(20):
0.197
CMfinder(seed):
0.016
Positive Predictive Value
PPfold(20):
0.879
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
TurboFold(seed):
0.106
Sensitivity
PPfold(20):
0.197
TurboFold(seed):
0.014
Positive Predictive Value
PPfold(20):
0.879
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs PPfold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
PPfold(seed):
0.097
Sensitivity
PPfold(20):
0.197
PPfold(seed):
0.012
Positive Predictive Value
PPfold(20):
0.879
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Carnac(seed):
0.093
Sensitivity
PPfold(20):
0.197
Carnac(seed):
0.010
Positive Predictive Value
PPfold(20):
0.879
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
RNASampler(seed):
0.088
Sensitivity
PPfold(20):
0.197
RNASampler(seed):
0.009
Positive Predictive Value
PPfold(20):
0.879
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Mastr(seed):
0.072
Sensitivity
PPfold(20):
0.197
Mastr(seed):
0.006
Positive Predictive Value
PPfold(20):
0.879
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs NanoFolder
Matthews Correlation Coefficient
PPfold(20):
0.416
NanoFolder:
0.068
Sensitivity
PPfold(20):
0.197
NanoFolder:
0.027
Positive Predictive Value
PPfold(20):
0.879
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(20) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Multilign(seed):
0.059
Sensitivity
PPfold(20):
0.197
Multilign(seed):
0.005
Positive Predictive Value
PPfold(20):
0.879
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Afold |
1242
ContextFold vs Afold
Matthews Correlation Coefficient
ContextFold:
0.754
Afold:
0.408
Sensitivity
ContextFold:
0.717
Afold:
0.343
Positive Predictive Value
ContextFold:
0.794
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Afold:
0.408
Sensitivity
PETfold_pre2.0(seed):
0.667
Afold:
0.343
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Afold:
0.408
Sensitivity
CentroidAlifold(seed):
0.414
Afold:
0.343
Positive Predictive Value
CentroidAlifold(seed):
0.901
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Afold
Matthews Correlation Coefficient
IPknot:
0.607
Afold:
0.408
Sensitivity
IPknot:
0.551
Afold:
0.343
Positive Predictive Value
IPknot:
0.670
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Afold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Afold:
0.408
Sensitivity
PETfold_pre2.0(20):
0.410
Afold:
0.343
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Afold
Matthews Correlation Coefficient
CentroidFold:
0.571
Afold:
0.408
Sensitivity
CentroidFold:
0.515
Afold:
0.343
Positive Predictive Value
CentroidFold:
0.633
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Afold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Afold:
0.408
Sensitivity
CentroidAlifold(20):
0.369
Afold:
0.343
Positive Predictive Value
CentroidAlifold(20):
0.891
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Afold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Afold:
0.408
Sensitivity
CentroidHomfold‑LAST:
0.359
Afold:
0.343
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Afold
Matthews Correlation Coefficient
Contrafold:
0.556
Afold:
0.408
Sensitivity
Contrafold:
0.542
Afold:
0.343
Positive Predictive Value
Contrafold:
0.571
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Afold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Afold:
0.408
Sensitivity
MXScarna(seed):
0.382
Afold:
0.343
Positive Predictive Value
MXScarna(seed):
0.753
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Afold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Afold:
0.408
Sensitivity
RNAalifold(20):
0.359
Afold:
0.343
Positive Predictive Value
RNAalifold(20):
0.787
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Afold
Matthews Correlation Coefficient
Sfold:
0.527
Afold:
0.408
Sensitivity
Sfold:
0.477
Afold:
0.343
Positive Predictive Value
Sfold:
0.582
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Afold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Afold:
0.408
Sensitivity
RNAalifold(seed):
0.328
Afold:
0.343
Positive Predictive Value
RNAalifold(seed):
0.835
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Afold
Matthews Correlation Coefficient
MaxExpect:
0.516
Afold:
0.408
Sensitivity
MaxExpect:
0.505
Afold:
0.343
Positive Predictive Value
MaxExpect:
0.528
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Afold
Matthews Correlation Coefficient
ProbKnot:
0.512
Afold:
0.408
Sensitivity
ProbKnot:
0.508
Afold:
0.343
Positive Predictive Value
ProbKnot:
0.517
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Afold
Matthews Correlation Coefficient
MXScarna(20):
0.500
Afold:
0.408
Sensitivity
MXScarna(20):
0.353
Afold:
0.343
Positive Predictive Value
MXScarna(20):
0.710
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Afold
Matthews Correlation Coefficient
UNAFold:
0.494
Afold:
0.408
Sensitivity
UNAFold:
0.491
Afold:
0.343
Positive Predictive Value
UNAFold:
0.498
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Afold
Matthews Correlation Coefficient
Fold:
0.493
Afold:
0.408
Sensitivity
Fold:
0.496
Afold:
0.343
Positive Predictive Value
Fold:
0.490
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Afold
Matthews Correlation Coefficient
RNAfold:
0.485
Afold:
0.408
Sensitivity
RNAfold:
0.486
Afold:
0.343
Positive Predictive Value
RNAfold:
0.485
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Afold
Matthews Correlation Coefficient
PknotsRG:
0.474
Afold:
0.408
Sensitivity
PknotsRG:
0.440
Afold:
0.343
Positive Predictive Value
PknotsRG:
0.510
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Afold
Matthews Correlation Coefficient
CRWrnafold:
0.430
Afold:
0.408
Sensitivity
CRWrnafold:
0.379
Afold:
0.343
Positive Predictive Value
CRWrnafold:
0.490
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Afold
Matthews Correlation Coefficient
RNAsubopt:
0.422
Afold:
0.408
Sensitivity
RNAsubopt:
0.344
Afold:
0.343
Positive Predictive Value
RNAsubopt:
0.519
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Afold
Matthews Correlation Coefficient
TurboFold(20):
0.419
Afold:
0.408
Sensitivity
TurboFold(20):
0.210
Afold:
0.343
Positive Predictive Value
TurboFold(20):
0.837
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Afold
Matthews Correlation Coefficient
PPfold(20):
0.416
Afold:
0.408
Sensitivity
PPfold(20):
0.197
Afold:
0.343
Positive Predictive Value
PPfold(20):
0.879
Afold:
0.487
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 7.5939721025e-08
|
|
+
Afold vs Carnac(20)
Matthews Correlation Coefficient
Afold:
0.408
Carnac(20):
0.401
Sensitivity
Afold:
0.343
Carnac(20):
0.180
Positive Predictive Value
Afold:
0.487
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.1569883686e-08
|
+
Afold vs McQFold
Matthews Correlation Coefficient
Afold:
0.408
McQFold:
0.389
Sensitivity
Afold:
0.343
McQFold:
0.308
Positive Predictive Value
Afold:
0.487
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RSpredict(20)
Matthews Correlation Coefficient
Afold:
0.408
RSpredict(20):
0.377
Sensitivity
Afold:
0.343
RSpredict(20):
0.241
Positive Predictive Value
Afold:
0.487
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNAshapes
Matthews Correlation Coefficient
Afold:
0.408
RNAshapes:
0.367
Sensitivity
Afold:
0.343
RNAshapes:
0.285
Positive Predictive Value
Afold:
0.487
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNASLOpt
Matthews Correlation Coefficient
Afold:
0.408
RNASLOpt:
0.364
Sensitivity
Afold:
0.343
RNASLOpt:
0.261
Positive Predictive Value
Afold:
0.487
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Multilign(20)
Matthews Correlation Coefficient
Afold:
0.408
Multilign(20):
0.350
Sensitivity
Afold:
0.343
Multilign(20):
0.161
Positive Predictive Value
Afold:
0.487
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Murlet(20)
Matthews Correlation Coefficient
Afold:
0.408
Murlet(20):
0.321
Sensitivity
Afold:
0.343
Murlet(20):
0.131
Positive Predictive Value
Afold:
0.487
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNAwolf
Matthews Correlation Coefficient
Afold:
0.408
RNAwolf:
0.310
Sensitivity
Afold:
0.343
RNAwolf:
0.261
Positive Predictive Value
Afold:
0.487
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RSpredict(seed)
Matthews Correlation Coefficient
Afold:
0.408
RSpredict(seed):
0.295
Sensitivity
Afold:
0.343
RSpredict(seed):
0.146
Positive Predictive Value
Afold:
0.487
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs CMfinder(20)
Matthews Correlation Coefficient
Afold:
0.408
CMfinder(20):
0.292
Sensitivity
Afold:
0.343
CMfinder(20):
0.107
Positive Predictive Value
Afold:
0.487
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Vsfold4
Matthews Correlation Coefficient
Afold:
0.408
Vsfold4:
0.270
Sensitivity
Afold:
0.343
Vsfold4:
0.198
Positive Predictive Value
Afold:
0.487
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Vsfold5
Matthews Correlation Coefficient
Afold:
0.408
Vsfold5:
0.260
Sensitivity
Afold:
0.343
Vsfold5:
0.195
Positive Predictive Value
Afold:
0.487
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNASampler(20)
Matthews Correlation Coefficient
Afold:
0.408
RNASampler(20):
0.246
Sensitivity
Afold:
0.343
RNASampler(20):
0.069
Positive Predictive Value
Afold:
0.487
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs HotKnots
Matthews Correlation Coefficient
Afold:
0.408
HotKnots:
0.212
Sensitivity
Afold:
0.343
HotKnots:
0.069
Positive Predictive Value
Afold:
0.487
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Mastr(20)
Matthews Correlation Coefficient
Afold:
0.408
Mastr(20):
0.187
Sensitivity
Afold:
0.343
Mastr(20):
0.044
Positive Predictive Value
Afold:
0.487
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Cylofold
Matthews Correlation Coefficient
Afold:
0.408
Cylofold:
0.178
Sensitivity
Afold:
0.343
Cylofold:
0.053
Positive Predictive Value
Afold:
0.487
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Pknots
Matthews Correlation Coefficient
Afold:
0.408
Pknots:
0.165
Sensitivity
Afold:
0.343
Pknots:
0.041
Positive Predictive Value
Afold:
0.487
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Alterna
Matthews Correlation Coefficient
Afold:
0.408
Alterna:
0.145
Sensitivity
Afold:
0.343
Alterna:
0.036
Positive Predictive Value
Afold:
0.487
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs MCFold
Matthews Correlation Coefficient
Afold:
0.408
MCFold:
0.138
Sensitivity
Afold:
0.343
MCFold:
0.044
Positive Predictive Value
Afold:
0.487
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RDfolder
Matthews Correlation Coefficient
Afold:
0.408
RDfolder:
0.138
Sensitivity
Afold:
0.343
RDfolder:
0.031
Positive Predictive Value
Afold:
0.487
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Murlet(seed)
Matthews Correlation Coefficient
Afold:
0.408
Murlet(seed):
0.110
Sensitivity
Afold:
0.343
Murlet(seed):
0.015
Positive Predictive Value
Afold:
0.487
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs CMfinder(seed)
Matthews Correlation Coefficient
Afold:
0.408
CMfinder(seed):
0.111
Sensitivity
Afold:
0.343
CMfinder(seed):
0.016
Positive Predictive Value
Afold:
0.487
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs TurboFold(seed)
Matthews Correlation Coefficient
Afold:
0.408
TurboFold(seed):
0.106
Sensitivity
Afold:
0.343
TurboFold(seed):
0.014
Positive Predictive Value
Afold:
0.487
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs PPfold(seed)
Matthews Correlation Coefficient
Afold:
0.408
PPfold(seed):
0.097
Sensitivity
Afold:
0.343
PPfold(seed):
0.012
Positive Predictive Value
Afold:
0.487
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Carnac(seed)
Matthews Correlation Coefficient
Afold:
0.408
Carnac(seed):
0.093
Sensitivity
Afold:
0.343
Carnac(seed):
0.010
Positive Predictive Value
Afold:
0.487
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs RNASampler(seed)
Matthews Correlation Coefficient
Afold:
0.408
RNASampler(seed):
0.088
Sensitivity
Afold:
0.343
RNASampler(seed):
0.009
Positive Predictive Value
Afold:
0.487
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Mastr(seed)
Matthews Correlation Coefficient
Afold:
0.408
Mastr(seed):
0.072
Sensitivity
Afold:
0.343
Mastr(seed):
0.006
Positive Predictive Value
Afold:
0.487
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs NanoFolder
Matthews Correlation Coefficient
Afold:
0.408
NanoFolder:
0.068
Sensitivity
Afold:
0.343
NanoFolder:
0.027
Positive Predictive Value
Afold:
0.487
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Afold vs Multilign(seed)
Matthews Correlation Coefficient
Afold:
0.408
Multilign(seed):
0.059
Sensitivity
Afold:
0.343
Multilign(seed):
0.005
Positive Predictive Value
Afold:
0.487
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Carnac(20) |
1242
ContextFold vs Carnac(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Carnac(20):
0.401
Sensitivity
ContextFold:
0.717
Carnac(20):
0.180
Positive Predictive Value
ContextFold:
0.794
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Carnac(20):
0.401
Sensitivity
PETfold_pre2.0(seed):
0.667
Carnac(20):
0.180
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Carnac(20):
0.401
Sensitivity
CentroidAlifold(seed):
0.414
Carnac(20):
0.180
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Carnac(20)
Matthews Correlation Coefficient
IPknot:
0.607
Carnac(20):
0.401
Sensitivity
IPknot:
0.551
Carnac(20):
0.180
Positive Predictive Value
IPknot:
0.670
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Carnac(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Carnac(20):
0.401
Sensitivity
PETfold_pre2.0(20):
0.410
Carnac(20):
0.180
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Carnac(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Carnac(20):
0.401
Sensitivity
CentroidFold:
0.515
Carnac(20):
0.180
Positive Predictive Value
CentroidFold:
0.633
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Carnac(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Carnac(20):
0.401
Sensitivity
CentroidAlifold(20):
0.369
Carnac(20):
0.180
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Carnac(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Carnac(20):
0.401
Sensitivity
CentroidHomfold‑LAST:
0.359
Carnac(20):
0.180
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Carnac(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Carnac(20):
0.401
Sensitivity
Contrafold:
0.542
Carnac(20):
0.180
Positive Predictive Value
Contrafold:
0.571
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Carnac(20):
0.401
Sensitivity
MXScarna(seed):
0.382
Carnac(20):
0.180
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Carnac(20):
0.401
Sensitivity
RNAalifold(20):
0.359
Carnac(20):
0.180
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Carnac(20)
Matthews Correlation Coefficient
Sfold:
0.527
Carnac(20):
0.401
Sensitivity
Sfold:
0.477
Carnac(20):
0.180
Positive Predictive Value
Sfold:
0.582
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Carnac(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Carnac(20):
0.401
Sensitivity
RNAalifold(seed):
0.328
Carnac(20):
0.180
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Carnac(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Carnac(20):
0.401
Sensitivity
MaxExpect:
0.505
Carnac(20):
0.180
Positive Predictive Value
MaxExpect:
0.528
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Carnac(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Carnac(20):
0.401
Sensitivity
ProbKnot:
0.508
Carnac(20):
0.180
Positive Predictive Value
ProbKnot:
0.517
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Carnac(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Carnac(20):
0.401
Sensitivity
MXScarna(20):
0.353
Carnac(20):
0.180
Positive Predictive Value
MXScarna(20):
0.710
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Carnac(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Carnac(20):
0.401
Sensitivity
UNAFold:
0.491
Carnac(20):
0.180
Positive Predictive Value
UNAFold:
0.498
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Carnac(20)
Matthews Correlation Coefficient
Fold:
0.493
Carnac(20):
0.401
Sensitivity
Fold:
0.496
Carnac(20):
0.180
Positive Predictive Value
Fold:
0.490
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Carnac(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Carnac(20):
0.401
Sensitivity
RNAfold:
0.486
Carnac(20):
0.180
Positive Predictive Value
RNAfold:
0.485
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Carnac(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Carnac(20):
0.401
Sensitivity
PknotsRG:
0.440
Carnac(20):
0.180
Positive Predictive Value
PknotsRG:
0.510
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Carnac(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Carnac(20):
0.401
Sensitivity
CRWrnafold:
0.379
Carnac(20):
0.180
Positive Predictive Value
CRWrnafold:
0.490
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Carnac(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Carnac(20):
0.401
Sensitivity
RNAsubopt:
0.344
Carnac(20):
0.180
Positive Predictive Value
RNAsubopt:
0.519
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Carnac(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Carnac(20):
0.401
Sensitivity
TurboFold(20):
0.210
Carnac(20):
0.180
Positive Predictive Value
TurboFold(20):
0.837
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Carnac(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Carnac(20):
0.401
Sensitivity
PPfold(20):
0.197
Carnac(20):
0.180
Positive Predictive Value
PPfold(20):
0.879
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Carnac(20)
Matthews Correlation Coefficient
Afold:
0.408
Carnac(20):
0.401
Sensitivity
Afold:
0.343
Carnac(20):
0.180
Positive Predictive Value
Afold:
0.487
Carnac(20):
0.892
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.1569883686e-08
|
|
+
Carnac(20) vs McQFold
Matthews Correlation Coefficient
Carnac(20):
0.401
McQFold:
0.389
Sensitivity
Carnac(20):
0.180
McQFold:
0.308
Positive Predictive Value
Carnac(20):
0.892
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.1569883686e-08
|
+
Carnac(20) vs RSpredict(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
RSpredict(20):
0.377
Sensitivity
Carnac(20):
0.180
RSpredict(20):
0.241
Positive Predictive Value
Carnac(20):
0.892
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNAshapes
Matthews Correlation Coefficient
Carnac(20):
0.401
RNAshapes:
0.367
Sensitivity
Carnac(20):
0.180
RNAshapes:
0.285
Positive Predictive Value
Carnac(20):
0.892
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNASLOpt
Matthews Correlation Coefficient
Carnac(20):
0.401
RNASLOpt:
0.364
Sensitivity
Carnac(20):
0.180
RNASLOpt:
0.261
Positive Predictive Value
Carnac(20):
0.892
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Multilign(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
Multilign(20):
0.350
Sensitivity
Carnac(20):
0.180
Multilign(20):
0.161
Positive Predictive Value
Carnac(20):
0.892
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Murlet(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
Murlet(20):
0.321
Sensitivity
Carnac(20):
0.180
Murlet(20):
0.131
Positive Predictive Value
Carnac(20):
0.892
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNAwolf
Matthews Correlation Coefficient
Carnac(20):
0.401
RNAwolf:
0.310
Sensitivity
Carnac(20):
0.180
RNAwolf:
0.261
Positive Predictive Value
Carnac(20):
0.892
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
RSpredict(seed):
0.295
Sensitivity
Carnac(20):
0.180
RSpredict(seed):
0.146
Positive Predictive Value
Carnac(20):
0.892
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs CMfinder(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
CMfinder(20):
0.292
Sensitivity
Carnac(20):
0.180
CMfinder(20):
0.107
Positive Predictive Value
Carnac(20):
0.892
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Vsfold4
Matthews Correlation Coefficient
Carnac(20):
0.401
Vsfold4:
0.270
Sensitivity
Carnac(20):
0.180
Vsfold4:
0.198
Positive Predictive Value
Carnac(20):
0.892
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Vsfold5
Matthews Correlation Coefficient
Carnac(20):
0.401
Vsfold5:
0.260
Sensitivity
Carnac(20):
0.180
Vsfold5:
0.195
Positive Predictive Value
Carnac(20):
0.892
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNASampler(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
RNASampler(20):
0.246
Sensitivity
Carnac(20):
0.180
RNASampler(20):
0.069
Positive Predictive Value
Carnac(20):
0.892
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs HotKnots
Matthews Correlation Coefficient
Carnac(20):
0.401
HotKnots:
0.212
Sensitivity
Carnac(20):
0.180
HotKnots:
0.069
Positive Predictive Value
Carnac(20):
0.892
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Mastr(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
Mastr(20):
0.187
Sensitivity
Carnac(20):
0.180
Mastr(20):
0.044
Positive Predictive Value
Carnac(20):
0.892
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Cylofold
Matthews Correlation Coefficient
Carnac(20):
0.401
Cylofold:
0.178
Sensitivity
Carnac(20):
0.180
Cylofold:
0.053
Positive Predictive Value
Carnac(20):
0.892
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Pknots
Matthews Correlation Coefficient
Carnac(20):
0.401
Pknots:
0.165
Sensitivity
Carnac(20):
0.180
Pknots:
0.041
Positive Predictive Value
Carnac(20):
0.892
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Alterna
Matthews Correlation Coefficient
Carnac(20):
0.401
Alterna:
0.145
Sensitivity
Carnac(20):
0.180
Alterna:
0.036
Positive Predictive Value
Carnac(20):
0.892
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs MCFold
Matthews Correlation Coefficient
Carnac(20):
0.401
MCFold:
0.138
Sensitivity
Carnac(20):
0.180
MCFold:
0.044
Positive Predictive Value
Carnac(20):
0.892
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RDfolder
Matthews Correlation Coefficient
Carnac(20):
0.401
RDfolder:
0.138
Sensitivity
Carnac(20):
0.180
RDfolder:
0.031
Positive Predictive Value
Carnac(20):
0.892
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Murlet(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Murlet(seed):
0.110
Sensitivity
Carnac(20):
0.180
Murlet(seed):
0.015
Positive Predictive Value
Carnac(20):
0.892
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
CMfinder(seed):
0.111
Sensitivity
Carnac(20):
0.180
CMfinder(seed):
0.016
Positive Predictive Value
Carnac(20):
0.892
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
TurboFold(seed):
0.106
Sensitivity
Carnac(20):
0.180
TurboFold(seed):
0.014
Positive Predictive Value
Carnac(20):
0.892
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs PPfold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
PPfold(seed):
0.097
Sensitivity
Carnac(20):
0.180
PPfold(seed):
0.012
Positive Predictive Value
Carnac(20):
0.892
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Carnac(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Carnac(seed):
0.093
Sensitivity
Carnac(20):
0.180
Carnac(seed):
0.010
Positive Predictive Value
Carnac(20):
0.892
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
RNASampler(seed):
0.088
Sensitivity
Carnac(20):
0.180
RNASampler(seed):
0.009
Positive Predictive Value
Carnac(20):
0.892
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Mastr(seed):
0.072
Sensitivity
Carnac(20):
0.180
Mastr(seed):
0.006
Positive Predictive Value
Carnac(20):
0.892
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs NanoFolder
Matthews Correlation Coefficient
Carnac(20):
0.401
NanoFolder:
0.068
Sensitivity
Carnac(20):
0.180
NanoFolder:
0.027
Positive Predictive Value
Carnac(20):
0.892
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(20) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Multilign(seed):
0.059
Sensitivity
Carnac(20):
0.180
Multilign(seed):
0.005
Positive Predictive Value
Carnac(20):
0.892
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| McQFold |
1242
ContextFold vs McQFold
Matthews Correlation Coefficient
ContextFold:
0.754
McQFold:
0.389
Sensitivity
ContextFold:
0.717
McQFold:
0.308
Positive Predictive Value
ContextFold:
0.794
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
McQFold:
0.389
Sensitivity
PETfold_pre2.0(seed):
0.667
McQFold:
0.308
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
McQFold:
0.389
Sensitivity
CentroidAlifold(seed):
0.414
McQFold:
0.308
Positive Predictive Value
CentroidAlifold(seed):
0.901
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs McQFold
Matthews Correlation Coefficient
IPknot:
0.607
McQFold:
0.389
Sensitivity
IPknot:
0.551
McQFold:
0.308
Positive Predictive Value
IPknot:
0.670
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs McQFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
McQFold:
0.389
Sensitivity
PETfold_pre2.0(20):
0.410
McQFold:
0.308
Positive Predictive Value
PETfold_pre2.0(20):
0.807
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs McQFold
Matthews Correlation Coefficient
CentroidFold:
0.571
McQFold:
0.389
Sensitivity
CentroidFold:
0.515
McQFold:
0.308
Positive Predictive Value
CentroidFold:
0.633
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs McQFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
McQFold:
0.389
Sensitivity
CentroidAlifold(20):
0.369
McQFold:
0.308
Positive Predictive Value
CentroidAlifold(20):
0.891
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs McQFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
McQFold:
0.389
Sensitivity
CentroidHomfold‑LAST:
0.359
McQFold:
0.308
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs McQFold
Matthews Correlation Coefficient
Contrafold:
0.556
McQFold:
0.389
Sensitivity
Contrafold:
0.542
McQFold:
0.308
Positive Predictive Value
Contrafold:
0.571
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs McQFold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
McQFold:
0.389
Sensitivity
MXScarna(seed):
0.382
McQFold:
0.308
Positive Predictive Value
MXScarna(seed):
0.753
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs McQFold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
McQFold:
0.389
Sensitivity
RNAalifold(20):
0.359
McQFold:
0.308
Positive Predictive Value
RNAalifold(20):
0.787
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs McQFold
Matthews Correlation Coefficient
Sfold:
0.527
McQFold:
0.389
Sensitivity
Sfold:
0.477
McQFold:
0.308
Positive Predictive Value
Sfold:
0.582
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs McQFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
McQFold:
0.389
Sensitivity
RNAalifold(seed):
0.328
McQFold:
0.308
Positive Predictive Value
RNAalifold(seed):
0.835
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs McQFold
Matthews Correlation Coefficient
MaxExpect:
0.516
McQFold:
0.389
Sensitivity
MaxExpect:
0.505
McQFold:
0.308
Positive Predictive Value
MaxExpect:
0.528
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs McQFold
Matthews Correlation Coefficient
ProbKnot:
0.512
McQFold:
0.389
Sensitivity
ProbKnot:
0.508
McQFold:
0.308
Positive Predictive Value
ProbKnot:
0.517
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs McQFold
Matthews Correlation Coefficient
MXScarna(20):
0.500
McQFold:
0.389
Sensitivity
MXScarna(20):
0.353
McQFold:
0.308
Positive Predictive Value
MXScarna(20):
0.710
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs McQFold
Matthews Correlation Coefficient
UNAFold:
0.494
McQFold:
0.389
Sensitivity
UNAFold:
0.491
McQFold:
0.308
Positive Predictive Value
UNAFold:
0.498
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs McQFold
Matthews Correlation Coefficient
Fold:
0.493
McQFold:
0.389
Sensitivity
Fold:
0.496
McQFold:
0.308
Positive Predictive Value
Fold:
0.490
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs McQFold
Matthews Correlation Coefficient
RNAfold:
0.485
McQFold:
0.389
Sensitivity
RNAfold:
0.486
McQFold:
0.308
Positive Predictive Value
RNAfold:
0.485
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs McQFold
Matthews Correlation Coefficient
PknotsRG:
0.474
McQFold:
0.389
Sensitivity
PknotsRG:
0.440
McQFold:
0.308
Positive Predictive Value
PknotsRG:
0.510
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs McQFold
Matthews Correlation Coefficient
CRWrnafold:
0.430
McQFold:
0.389
Sensitivity
CRWrnafold:
0.379
McQFold:
0.308
Positive Predictive Value
CRWrnafold:
0.490
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs McQFold
Matthews Correlation Coefficient
RNAsubopt:
0.422
McQFold:
0.389
Sensitivity
RNAsubopt:
0.344
McQFold:
0.308
Positive Predictive Value
RNAsubopt:
0.519
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs McQFold
Matthews Correlation Coefficient
TurboFold(20):
0.419
McQFold:
0.389
Sensitivity
TurboFold(20):
0.210
McQFold:
0.308
Positive Predictive Value
TurboFold(20):
0.837
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs McQFold
Matthews Correlation Coefficient
PPfold(20):
0.416
McQFold:
0.389
Sensitivity
PPfold(20):
0.197
McQFold:
0.308
Positive Predictive Value
PPfold(20):
0.879
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs McQFold
Matthews Correlation Coefficient
Afold:
0.408
McQFold:
0.389
Sensitivity
Afold:
0.343
McQFold:
0.308
Positive Predictive Value
Afold:
0.487
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs McQFold
Matthews Correlation Coefficient
Carnac(20):
0.401
McQFold:
0.389
Sensitivity
Carnac(20):
0.180
McQFold:
0.308
Positive Predictive Value
Carnac(20):
0.892
McQFold:
0.492
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 4.1569883686e-08
|
|
+
McQFold vs RSpredict(20)
Matthews Correlation Coefficient
McQFold:
0.389
RSpredict(20):
0.377
Sensitivity
McQFold:
0.308
RSpredict(20):
0.241
Positive Predictive Value
McQFold:
0.492
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNAshapes
Matthews Correlation Coefficient
McQFold:
0.389
RNAshapes:
0.367
Sensitivity
McQFold:
0.308
RNAshapes:
0.285
Positive Predictive Value
McQFold:
0.492
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNASLOpt
Matthews Correlation Coefficient
McQFold:
0.389
RNASLOpt:
0.364
Sensitivity
McQFold:
0.308
RNASLOpt:
0.261
Positive Predictive Value
McQFold:
0.492
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Multilign(20)
Matthews Correlation Coefficient
McQFold:
0.389
Multilign(20):
0.350
Sensitivity
McQFold:
0.308
Multilign(20):
0.161
Positive Predictive Value
McQFold:
0.492
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Murlet(20)
Matthews Correlation Coefficient
McQFold:
0.389
Murlet(20):
0.321
Sensitivity
McQFold:
0.308
Murlet(20):
0.131
Positive Predictive Value
McQFold:
0.492
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNAwolf
Matthews Correlation Coefficient
McQFold:
0.389
RNAwolf:
0.310
Sensitivity
McQFold:
0.308
RNAwolf:
0.261
Positive Predictive Value
McQFold:
0.492
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RSpredict(seed)
Matthews Correlation Coefficient
McQFold:
0.389
RSpredict(seed):
0.295
Sensitivity
McQFold:
0.308
RSpredict(seed):
0.146
Positive Predictive Value
McQFold:
0.492
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs CMfinder(20)
Matthews Correlation Coefficient
McQFold:
0.389
CMfinder(20):
0.292
Sensitivity
McQFold:
0.308
CMfinder(20):
0.107
Positive Predictive Value
McQFold:
0.492
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Vsfold4
Matthews Correlation Coefficient
McQFold:
0.389
Vsfold4:
0.270
Sensitivity
McQFold:
0.308
Vsfold4:
0.198
Positive Predictive Value
McQFold:
0.492
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Vsfold5
Matthews Correlation Coefficient
McQFold:
0.389
Vsfold5:
0.260
Sensitivity
McQFold:
0.308
Vsfold5:
0.195
Positive Predictive Value
McQFold:
0.492
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNASampler(20)
Matthews Correlation Coefficient
McQFold:
0.389
RNASampler(20):
0.246
Sensitivity
McQFold:
0.308
RNASampler(20):
0.069
Positive Predictive Value
McQFold:
0.492
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs HotKnots
Matthews Correlation Coefficient
McQFold:
0.389
HotKnots:
0.212
Sensitivity
McQFold:
0.308
HotKnots:
0.069
Positive Predictive Value
McQFold:
0.492
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Mastr(20)
Matthews Correlation Coefficient
McQFold:
0.389
Mastr(20):
0.187
Sensitivity
McQFold:
0.308
Mastr(20):
0.044
Positive Predictive Value
McQFold:
0.492
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Cylofold
Matthews Correlation Coefficient
McQFold:
0.389
Cylofold:
0.178
Sensitivity
McQFold:
0.308
Cylofold:
0.053
Positive Predictive Value
McQFold:
0.492
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Pknots
Matthews Correlation Coefficient
McQFold:
0.389
Pknots:
0.165
Sensitivity
McQFold:
0.308
Pknots:
0.041
Positive Predictive Value
McQFold:
0.492
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Alterna
Matthews Correlation Coefficient
McQFold:
0.389
Alterna:
0.145
Sensitivity
McQFold:
0.308
Alterna:
0.036
Positive Predictive Value
McQFold:
0.492
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs MCFold
Matthews Correlation Coefficient
McQFold:
0.389
MCFold:
0.138
Sensitivity
McQFold:
0.308
MCFold:
0.044
Positive Predictive Value
McQFold:
0.492
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RDfolder
Matthews Correlation Coefficient
McQFold:
0.389
RDfolder:
0.138
Sensitivity
McQFold:
0.308
RDfolder:
0.031
Positive Predictive Value
McQFold:
0.492
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Murlet(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Murlet(seed):
0.110
Sensitivity
McQFold:
0.308
Murlet(seed):
0.015
Positive Predictive Value
McQFold:
0.492
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs CMfinder(seed)
Matthews Correlation Coefficient
McQFold:
0.389
CMfinder(seed):
0.111
Sensitivity
McQFold:
0.308
CMfinder(seed):
0.016
Positive Predictive Value
McQFold:
0.492
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs TurboFold(seed)
Matthews Correlation Coefficient
McQFold:
0.389
TurboFold(seed):
0.106
Sensitivity
McQFold:
0.308
TurboFold(seed):
0.014
Positive Predictive Value
McQFold:
0.492
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs PPfold(seed)
Matthews Correlation Coefficient
McQFold:
0.389
PPfold(seed):
0.097
Sensitivity
McQFold:
0.308
PPfold(seed):
0.012
Positive Predictive Value
McQFold:
0.492
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Carnac(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Carnac(seed):
0.093
Sensitivity
McQFold:
0.308
Carnac(seed):
0.010
Positive Predictive Value
McQFold:
0.492
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs RNASampler(seed)
Matthews Correlation Coefficient
McQFold:
0.389
RNASampler(seed):
0.088
Sensitivity
McQFold:
0.308
RNASampler(seed):
0.009
Positive Predictive Value
McQFold:
0.492
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Mastr(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Mastr(seed):
0.072
Sensitivity
McQFold:
0.308
Mastr(seed):
0.006
Positive Predictive Value
McQFold:
0.492
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs NanoFolder
Matthews Correlation Coefficient
McQFold:
0.389
NanoFolder:
0.068
Sensitivity
McQFold:
0.308
NanoFolder:
0.027
Positive Predictive Value
McQFold:
0.492
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
McQFold vs Multilign(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Multilign(seed):
0.059
Sensitivity
McQFold:
0.308
Multilign(seed):
0.005
Positive Predictive Value
McQFold:
0.492
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RSpredict(20) |
1242
ContextFold vs RSpredict(20)
Matthews Correlation Coefficient
ContextFold:
0.754
RSpredict(20):
0.377
Sensitivity
ContextFold:
0.717
RSpredict(20):
0.241
Positive Predictive Value
ContextFold:
0.794
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RSpredict(20):
0.377
Sensitivity
PETfold_pre2.0(seed):
0.667
RSpredict(20):
0.241
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RSpredict(20):
0.377
Sensitivity
CentroidAlifold(seed):
0.414
RSpredict(20):
0.241
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RSpredict(20)
Matthews Correlation Coefficient
IPknot:
0.607
RSpredict(20):
0.377
Sensitivity
IPknot:
0.551
RSpredict(20):
0.241
Positive Predictive Value
IPknot:
0.670
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RSpredict(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RSpredict(20):
0.377
Sensitivity
PETfold_pre2.0(20):
0.410
RSpredict(20):
0.241
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RSpredict(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
RSpredict(20):
0.377
Sensitivity
CentroidFold:
0.515
RSpredict(20):
0.241
Positive Predictive Value
CentroidFold:
0.633
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RSpredict(20):
0.377
Sensitivity
CentroidAlifold(20):
0.369
RSpredict(20):
0.241
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RSpredict(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RSpredict(20):
0.377
Sensitivity
CentroidHomfold‑LAST:
0.359
RSpredict(20):
0.241
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RSpredict(20)
Matthews Correlation Coefficient
Contrafold:
0.556
RSpredict(20):
0.377
Sensitivity
Contrafold:
0.542
RSpredict(20):
0.241
Positive Predictive Value
Contrafold:
0.571
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RSpredict(20):
0.377
Sensitivity
MXScarna(seed):
0.382
RSpredict(20):
0.241
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RSpredict(20):
0.377
Sensitivity
RNAalifold(20):
0.359
RSpredict(20):
0.241
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RSpredict(20)
Matthews Correlation Coefficient
Sfold:
0.527
RSpredict(20):
0.377
Sensitivity
Sfold:
0.477
RSpredict(20):
0.241
Positive Predictive Value
Sfold:
0.582
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RSpredict(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RSpredict(20):
0.377
Sensitivity
RNAalifold(seed):
0.328
RSpredict(20):
0.241
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RSpredict(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
RSpredict(20):
0.377
Sensitivity
MaxExpect:
0.505
RSpredict(20):
0.241
Positive Predictive Value
MaxExpect:
0.528
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RSpredict(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
RSpredict(20):
0.377
Sensitivity
ProbKnot:
0.508
RSpredict(20):
0.241
Positive Predictive Value
ProbKnot:
0.517
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RSpredict(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RSpredict(20):
0.377
Sensitivity
MXScarna(20):
0.353
RSpredict(20):
0.241
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RSpredict(20)
Matthews Correlation Coefficient
UNAFold:
0.494
RSpredict(20):
0.377
Sensitivity
UNAFold:
0.491
RSpredict(20):
0.241
Positive Predictive Value
UNAFold:
0.498
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RSpredict(20)
Matthews Correlation Coefficient
Fold:
0.493
RSpredict(20):
0.377
Sensitivity
Fold:
0.496
RSpredict(20):
0.241
Positive Predictive Value
Fold:
0.490
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RSpredict(20)
Matthews Correlation Coefficient
RNAfold:
0.485
RSpredict(20):
0.377
Sensitivity
RNAfold:
0.486
RSpredict(20):
0.241
Positive Predictive Value
RNAfold:
0.485
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RSpredict(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
RSpredict(20):
0.377
Sensitivity
PknotsRG:
0.440
RSpredict(20):
0.241
Positive Predictive Value
PknotsRG:
0.510
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RSpredict(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RSpredict(20):
0.377
Sensitivity
CRWrnafold:
0.379
RSpredict(20):
0.241
Positive Predictive Value
CRWrnafold:
0.490
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RSpredict(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RSpredict(20):
0.377
Sensitivity
RNAsubopt:
0.344
RSpredict(20):
0.241
Positive Predictive Value
RNAsubopt:
0.519
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RSpredict(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RSpredict(20):
0.377
Sensitivity
TurboFold(20):
0.210
RSpredict(20):
0.241
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RSpredict(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
RSpredict(20):
0.377
Sensitivity
PPfold(20):
0.197
RSpredict(20):
0.241
Positive Predictive Value
PPfold(20):
0.879
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RSpredict(20)
Matthews Correlation Coefficient
Afold:
0.408
RSpredict(20):
0.377
Sensitivity
Afold:
0.343
RSpredict(20):
0.241
Positive Predictive Value
Afold:
0.487
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RSpredict(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
RSpredict(20):
0.377
Sensitivity
Carnac(20):
0.180
RSpredict(20):
0.241
Positive Predictive Value
Carnac(20):
0.892
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RSpredict(20)
Matthews Correlation Coefficient
McQFold:
0.389
RSpredict(20):
0.377
Sensitivity
McQFold:
0.308
RSpredict(20):
0.241
Positive Predictive Value
McQFold:
0.492
RSpredict(20):
0.590
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RSpredict(20) vs RNAshapes
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNAshapes:
0.367
Sensitivity
RSpredict(20):
0.241
RNAshapes:
0.285
Positive Predictive Value
RSpredict(20):
0.590
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.18450348589e-07
|
+
RSpredict(20) vs RNASLOpt
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNASLOpt:
0.364
Sensitivity
RSpredict(20):
0.241
RNASLOpt:
0.261
Positive Predictive Value
RSpredict(20):
0.590
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Multilign(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Multilign(20):
0.350
Sensitivity
RSpredict(20):
0.241
Multilign(20):
0.161
Positive Predictive Value
RSpredict(20):
0.590
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Murlet(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Murlet(20):
0.321
Sensitivity
RSpredict(20):
0.241
Murlet(20):
0.131
Positive Predictive Value
RSpredict(20):
0.590
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RNAwolf
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNAwolf:
0.310
Sensitivity
RSpredict(20):
0.241
RNAwolf:
0.261
Positive Predictive Value
RSpredict(20):
0.590
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
RSpredict(seed):
0.295
Sensitivity
RSpredict(20):
0.241
RSpredict(seed):
0.146
Positive Predictive Value
RSpredict(20):
0.590
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
CMfinder(20):
0.292
Sensitivity
RSpredict(20):
0.241
CMfinder(20):
0.107
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Vsfold4
Matthews Correlation Coefficient
RSpredict(20):
0.377
Vsfold4:
0.270
Sensitivity
RSpredict(20):
0.241
Vsfold4:
0.198
Positive Predictive Value
RSpredict(20):
0.590
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(20):
0.377
Vsfold5:
0.260
Sensitivity
RSpredict(20):
0.241
Vsfold5:
0.195
Positive Predictive Value
RSpredict(20):
0.590
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNASampler(20):
0.246
Sensitivity
RSpredict(20):
0.241
RNASampler(20):
0.069
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs HotKnots
Matthews Correlation Coefficient
RSpredict(20):
0.377
HotKnots:
0.212
Sensitivity
RSpredict(20):
0.241
HotKnots:
0.069
Positive Predictive Value
RSpredict(20):
0.590
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Mastr(20):
0.187
Sensitivity
RSpredict(20):
0.241
Mastr(20):
0.044
Positive Predictive Value
RSpredict(20):
0.590
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Cylofold
Matthews Correlation Coefficient
RSpredict(20):
0.377
Cylofold:
0.178
Sensitivity
RSpredict(20):
0.241
Cylofold:
0.053
Positive Predictive Value
RSpredict(20):
0.590
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Pknots
Matthews Correlation Coefficient
RSpredict(20):
0.377
Pknots:
0.165
Sensitivity
RSpredict(20):
0.241
Pknots:
0.041
Positive Predictive Value
RSpredict(20):
0.590
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Alterna
Matthews Correlation Coefficient
RSpredict(20):
0.377
Alterna:
0.145
Sensitivity
RSpredict(20):
0.241
Alterna:
0.036
Positive Predictive Value
RSpredict(20):
0.590
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs MCFold
Matthews Correlation Coefficient
RSpredict(20):
0.377
MCFold:
0.138
Sensitivity
RSpredict(20):
0.241
MCFold:
0.044
Positive Predictive Value
RSpredict(20):
0.590
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RDfolder
Matthews Correlation Coefficient
RSpredict(20):
0.377
RDfolder:
0.138
Sensitivity
RSpredict(20):
0.241
RDfolder:
0.031
Positive Predictive Value
RSpredict(20):
0.590
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Murlet(seed):
0.110
Sensitivity
RSpredict(20):
0.241
Murlet(seed):
0.015
Positive Predictive Value
RSpredict(20):
0.590
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
CMfinder(seed):
0.111
Sensitivity
RSpredict(20):
0.241
CMfinder(seed):
0.016
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
TurboFold(seed):
0.106
Sensitivity
RSpredict(20):
0.241
TurboFold(seed):
0.014
Positive Predictive Value
RSpredict(20):
0.590
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
PPfold(seed):
0.097
Sensitivity
RSpredict(20):
0.241
PPfold(seed):
0.012
Positive Predictive Value
RSpredict(20):
0.590
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Carnac(seed):
0.093
Sensitivity
RSpredict(20):
0.241
Carnac(seed):
0.010
Positive Predictive Value
RSpredict(20):
0.590
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNASampler(seed):
0.088
Sensitivity
RSpredict(20):
0.241
RNASampler(seed):
0.009
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Mastr(seed):
0.072
Sensitivity
RSpredict(20):
0.241
Mastr(seed):
0.006
Positive Predictive Value
RSpredict(20):
0.590
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(20):
0.377
NanoFolder:
0.068
Sensitivity
RSpredict(20):
0.241
NanoFolder:
0.027
Positive Predictive Value
RSpredict(20):
0.590
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(20) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Multilign(seed):
0.059
Sensitivity
RSpredict(20):
0.241
Multilign(seed):
0.005
Positive Predictive Value
RSpredict(20):
0.590
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAshapes |
1242
ContextFold vs RNAshapes
Matthews Correlation Coefficient
ContextFold:
0.754
RNAshapes:
0.367
Sensitivity
ContextFold:
0.717
RNAshapes:
0.285
Positive Predictive Value
ContextFold:
0.794
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAshapes:
0.367
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAshapes:
0.285
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAshapes:
0.367
Sensitivity
CentroidAlifold(seed):
0.414
RNAshapes:
0.285
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNAshapes
Matthews Correlation Coefficient
IPknot:
0.607
RNAshapes:
0.367
Sensitivity
IPknot:
0.551
RNAshapes:
0.285
Positive Predictive Value
IPknot:
0.670
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNAshapes
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAshapes:
0.367
Sensitivity
PETfold_pre2.0(20):
0.410
RNAshapes:
0.285
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNAshapes
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAshapes:
0.367
Sensitivity
CentroidFold:
0.515
RNAshapes:
0.285
Positive Predictive Value
CentroidFold:
0.633
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNAshapes
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAshapes:
0.367
Sensitivity
CentroidAlifold(20):
0.369
RNAshapes:
0.285
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNAshapes
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAshapes:
0.367
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAshapes:
0.285
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNAshapes
Matthews Correlation Coefficient
Contrafold:
0.556
RNAshapes:
0.367
Sensitivity
Contrafold:
0.542
RNAshapes:
0.285
Positive Predictive Value
Contrafold:
0.571
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAshapes:
0.367
Sensitivity
MXScarna(seed):
0.382
RNAshapes:
0.285
Positive Predictive Value
MXScarna(seed):
0.753
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAshapes:
0.367
Sensitivity
RNAalifold(20):
0.359
RNAshapes:
0.285
Positive Predictive Value
RNAalifold(20):
0.787
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNAshapes
Matthews Correlation Coefficient
Sfold:
0.527
RNAshapes:
0.367
Sensitivity
Sfold:
0.477
RNAshapes:
0.285
Positive Predictive Value
Sfold:
0.582
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNAshapes
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAshapes:
0.367
Sensitivity
RNAalifold(seed):
0.328
RNAshapes:
0.285
Positive Predictive Value
RNAalifold(seed):
0.835
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNAshapes
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAshapes:
0.367
Sensitivity
MaxExpect:
0.505
RNAshapes:
0.285
Positive Predictive Value
MaxExpect:
0.528
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNAshapes
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAshapes:
0.367
Sensitivity
ProbKnot:
0.508
RNAshapes:
0.285
Positive Predictive Value
ProbKnot:
0.517
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNAshapes
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAshapes:
0.367
Sensitivity
MXScarna(20):
0.353
RNAshapes:
0.285
Positive Predictive Value
MXScarna(20):
0.710
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNAshapes
Matthews Correlation Coefficient
UNAFold:
0.494
RNAshapes:
0.367
Sensitivity
UNAFold:
0.491
RNAshapes:
0.285
Positive Predictive Value
UNAFold:
0.498
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNAshapes
Matthews Correlation Coefficient
Fold:
0.493
RNAshapes:
0.367
Sensitivity
Fold:
0.496
RNAshapes:
0.285
Positive Predictive Value
Fold:
0.490
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RNAshapes
Matthews Correlation Coefficient
RNAfold:
0.485
RNAshapes:
0.367
Sensitivity
RNAfold:
0.486
RNAshapes:
0.285
Positive Predictive Value
RNAfold:
0.485
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RNAshapes
Matthews Correlation Coefficient
PknotsRG:
0.474
RNAshapes:
0.367
Sensitivity
PknotsRG:
0.440
RNAshapes:
0.285
Positive Predictive Value
PknotsRG:
0.510
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RNAshapes
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNAshapes:
0.367
Sensitivity
CRWrnafold:
0.379
RNAshapes:
0.285
Positive Predictive Value
CRWrnafold:
0.490
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RNAshapes
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNAshapes:
0.367
Sensitivity
RNAsubopt:
0.344
RNAshapes:
0.285
Positive Predictive Value
RNAsubopt:
0.519
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RNAshapes
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNAshapes:
0.367
Sensitivity
TurboFold(20):
0.210
RNAshapes:
0.285
Positive Predictive Value
TurboFold(20):
0.837
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RNAshapes
Matthews Correlation Coefficient
PPfold(20):
0.416
RNAshapes:
0.367
Sensitivity
PPfold(20):
0.197
RNAshapes:
0.285
Positive Predictive Value
PPfold(20):
0.879
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RNAshapes
Matthews Correlation Coefficient
Afold:
0.408
RNAshapes:
0.367
Sensitivity
Afold:
0.343
RNAshapes:
0.285
Positive Predictive Value
Afold:
0.487
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RNAshapes
Matthews Correlation Coefficient
Carnac(20):
0.401
RNAshapes:
0.367
Sensitivity
Carnac(20):
0.180
RNAshapes:
0.285
Positive Predictive Value
Carnac(20):
0.892
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RNAshapes
Matthews Correlation Coefficient
McQFold:
0.389
RNAshapes:
0.367
Sensitivity
McQFold:
0.308
RNAshapes:
0.285
Positive Predictive Value
McQFold:
0.492
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RNAshapes
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNAshapes:
0.367
Sensitivity
RSpredict(20):
0.241
RNAshapes:
0.285
Positive Predictive Value
RSpredict(20):
0.590
RNAshapes:
0.474
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.18450348589e-07
|
|
+
RNAshapes vs RNASLOpt
Matthews Correlation Coefficient
RNAshapes:
0.367
RNASLOpt:
0.364
Sensitivity
RNAshapes:
0.285
RNASLOpt:
0.261
Positive Predictive Value
RNAshapes:
0.474
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 7.04732813174e-08
|
+
RNAshapes vs Multilign(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
Multilign(20):
0.350
Sensitivity
RNAshapes:
0.285
Multilign(20):
0.161
Positive Predictive Value
RNAshapes:
0.474
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Murlet(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
Murlet(20):
0.321
Sensitivity
RNAshapes:
0.285
Murlet(20):
0.131
Positive Predictive Value
RNAshapes:
0.474
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RNAwolf
Matthews Correlation Coefficient
RNAshapes:
0.367
RNAwolf:
0.310
Sensitivity
RNAshapes:
0.285
RNAwolf:
0.261
Positive Predictive Value
RNAshapes:
0.474
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RSpredict(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
RSpredict(seed):
0.295
Sensitivity
RNAshapes:
0.285
RSpredict(seed):
0.146
Positive Predictive Value
RNAshapes:
0.474
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs CMfinder(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
CMfinder(20):
0.292
Sensitivity
RNAshapes:
0.285
CMfinder(20):
0.107
Positive Predictive Value
RNAshapes:
0.474
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Vsfold4
Matthews Correlation Coefficient
RNAshapes:
0.367
Vsfold4:
0.270
Sensitivity
RNAshapes:
0.285
Vsfold4:
0.198
Positive Predictive Value
RNAshapes:
0.474
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Vsfold5
Matthews Correlation Coefficient
RNAshapes:
0.367
Vsfold5:
0.260
Sensitivity
RNAshapes:
0.285
Vsfold5:
0.195
Positive Predictive Value
RNAshapes:
0.474
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RNASampler(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
RNASampler(20):
0.246
Sensitivity
RNAshapes:
0.285
RNASampler(20):
0.069
Positive Predictive Value
RNAshapes:
0.474
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs HotKnots
Matthews Correlation Coefficient
RNAshapes:
0.367
HotKnots:
0.212
Sensitivity
RNAshapes:
0.285
HotKnots:
0.069
Positive Predictive Value
RNAshapes:
0.474
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Mastr(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
Mastr(20):
0.187
Sensitivity
RNAshapes:
0.285
Mastr(20):
0.044
Positive Predictive Value
RNAshapes:
0.474
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Cylofold
Matthews Correlation Coefficient
RNAshapes:
0.367
Cylofold:
0.178
Sensitivity
RNAshapes:
0.285
Cylofold:
0.053
Positive Predictive Value
RNAshapes:
0.474
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Pknots
Matthews Correlation Coefficient
RNAshapes:
0.367
Pknots:
0.165
Sensitivity
RNAshapes:
0.285
Pknots:
0.041
Positive Predictive Value
RNAshapes:
0.474
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Alterna
Matthews Correlation Coefficient
RNAshapes:
0.367
Alterna:
0.145
Sensitivity
RNAshapes:
0.285
Alterna:
0.036
Positive Predictive Value
RNAshapes:
0.474
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs MCFold
Matthews Correlation Coefficient
RNAshapes:
0.367
MCFold:
0.138
Sensitivity
RNAshapes:
0.285
MCFold:
0.044
Positive Predictive Value
RNAshapes:
0.474
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RDfolder
Matthews Correlation Coefficient
RNAshapes:
0.367
RDfolder:
0.138
Sensitivity
RNAshapes:
0.285
RDfolder:
0.031
Positive Predictive Value
RNAshapes:
0.474
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Murlet(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Murlet(seed):
0.110
Sensitivity
RNAshapes:
0.285
Murlet(seed):
0.015
Positive Predictive Value
RNAshapes:
0.474
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs CMfinder(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
CMfinder(seed):
0.111
Sensitivity
RNAshapes:
0.285
CMfinder(seed):
0.016
Positive Predictive Value
RNAshapes:
0.474
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs TurboFold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
TurboFold(seed):
0.106
Sensitivity
RNAshapes:
0.285
TurboFold(seed):
0.014
Positive Predictive Value
RNAshapes:
0.474
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs PPfold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
PPfold(seed):
0.097
Sensitivity
RNAshapes:
0.285
PPfold(seed):
0.012
Positive Predictive Value
RNAshapes:
0.474
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Carnac(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Carnac(seed):
0.093
Sensitivity
RNAshapes:
0.285
Carnac(seed):
0.010
Positive Predictive Value
RNAshapes:
0.474
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs RNASampler(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
RNASampler(seed):
0.088
Sensitivity
RNAshapes:
0.285
RNASampler(seed):
0.009
Positive Predictive Value
RNAshapes:
0.474
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Mastr(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Mastr(seed):
0.072
Sensitivity
RNAshapes:
0.285
Mastr(seed):
0.006
Positive Predictive Value
RNAshapes:
0.474
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs NanoFolder
Matthews Correlation Coefficient
RNAshapes:
0.367
NanoFolder:
0.068
Sensitivity
RNAshapes:
0.285
NanoFolder:
0.027
Positive Predictive Value
RNAshapes:
0.474
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAshapes vs Multilign(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Multilign(seed):
0.059
Sensitivity
RNAshapes:
0.285
Multilign(seed):
0.005
Positive Predictive Value
RNAshapes:
0.474
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNASLOpt |
1242
ContextFold vs RNASLOpt
Matthews Correlation Coefficient
ContextFold:
0.754
RNASLOpt:
0.364
Sensitivity
ContextFold:
0.717
RNASLOpt:
0.261
Positive Predictive Value
ContextFold:
0.794
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNASLOpt:
0.364
Sensitivity
PETfold_pre2.0(seed):
0.667
RNASLOpt:
0.261
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNASLOpt:
0.364
Sensitivity
CentroidAlifold(seed):
0.414
RNASLOpt:
0.261
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNASLOpt
Matthews Correlation Coefficient
IPknot:
0.607
RNASLOpt:
0.364
Sensitivity
IPknot:
0.551
RNASLOpt:
0.261
Positive Predictive Value
IPknot:
0.670
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNASLOpt
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNASLOpt:
0.364
Sensitivity
PETfold_pre2.0(20):
0.410
RNASLOpt:
0.261
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNASLOpt
Matthews Correlation Coefficient
CentroidFold:
0.571
RNASLOpt:
0.364
Sensitivity
CentroidFold:
0.515
RNASLOpt:
0.261
Positive Predictive Value
CentroidFold:
0.633
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNASLOpt
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNASLOpt:
0.364
Sensitivity
CentroidAlifold(20):
0.369
RNASLOpt:
0.261
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNASLOpt
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNASLOpt:
0.364
Sensitivity
CentroidHomfold‑LAST:
0.359
RNASLOpt:
0.261
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNASLOpt
Matthews Correlation Coefficient
Contrafold:
0.556
RNASLOpt:
0.364
Sensitivity
Contrafold:
0.542
RNASLOpt:
0.261
Positive Predictive Value
Contrafold:
0.571
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNASLOpt:
0.364
Sensitivity
MXScarna(seed):
0.382
RNASLOpt:
0.261
Positive Predictive Value
MXScarna(seed):
0.753
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNASLOpt:
0.364
Sensitivity
RNAalifold(20):
0.359
RNASLOpt:
0.261
Positive Predictive Value
RNAalifold(20):
0.787
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNASLOpt
Matthews Correlation Coefficient
Sfold:
0.527
RNASLOpt:
0.364
Sensitivity
Sfold:
0.477
RNASLOpt:
0.261
Positive Predictive Value
Sfold:
0.582
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNASLOpt
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNASLOpt:
0.364
Sensitivity
RNAalifold(seed):
0.328
RNASLOpt:
0.261
Positive Predictive Value
RNAalifold(seed):
0.835
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNASLOpt
Matthews Correlation Coefficient
MaxExpect:
0.516
RNASLOpt:
0.364
Sensitivity
MaxExpect:
0.505
RNASLOpt:
0.261
Positive Predictive Value
MaxExpect:
0.528
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNASLOpt
Matthews Correlation Coefficient
ProbKnot:
0.512
RNASLOpt:
0.364
Sensitivity
ProbKnot:
0.508
RNASLOpt:
0.261
Positive Predictive Value
ProbKnot:
0.517
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNASLOpt
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNASLOpt:
0.364
Sensitivity
MXScarna(20):
0.353
RNASLOpt:
0.261
Positive Predictive Value
MXScarna(20):
0.710
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNASLOpt
Matthews Correlation Coefficient
UNAFold:
0.494
RNASLOpt:
0.364
Sensitivity
UNAFold:
0.491
RNASLOpt:
0.261
Positive Predictive Value
UNAFold:
0.498
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNASLOpt
Matthews Correlation Coefficient
Fold:
0.493
RNASLOpt:
0.364
Sensitivity
Fold:
0.496
RNASLOpt:
0.261
Positive Predictive Value
Fold:
0.490
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RNASLOpt
Matthews Correlation Coefficient
RNAfold:
0.485
RNASLOpt:
0.364
Sensitivity
RNAfold:
0.486
RNASLOpt:
0.261
Positive Predictive Value
RNAfold:
0.485
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RNASLOpt
Matthews Correlation Coefficient
PknotsRG:
0.474
RNASLOpt:
0.364
Sensitivity
PknotsRG:
0.440
RNASLOpt:
0.261
Positive Predictive Value
PknotsRG:
0.510
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RNASLOpt
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNASLOpt:
0.364
Sensitivity
CRWrnafold:
0.379
RNASLOpt:
0.261
Positive Predictive Value
CRWrnafold:
0.490
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RNASLOpt
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNASLOpt:
0.364
Sensitivity
RNAsubopt:
0.344
RNASLOpt:
0.261
Positive Predictive Value
RNAsubopt:
0.519
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RNASLOpt
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNASLOpt:
0.364
Sensitivity
TurboFold(20):
0.210
RNASLOpt:
0.261
Positive Predictive Value
TurboFold(20):
0.837
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RNASLOpt
Matthews Correlation Coefficient
PPfold(20):
0.416
RNASLOpt:
0.364
Sensitivity
PPfold(20):
0.197
RNASLOpt:
0.261
Positive Predictive Value
PPfold(20):
0.879
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RNASLOpt
Matthews Correlation Coefficient
Afold:
0.408
RNASLOpt:
0.364
Sensitivity
Afold:
0.343
RNASLOpt:
0.261
Positive Predictive Value
Afold:
0.487
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RNASLOpt
Matthews Correlation Coefficient
Carnac(20):
0.401
RNASLOpt:
0.364
Sensitivity
Carnac(20):
0.180
RNASLOpt:
0.261
Positive Predictive Value
Carnac(20):
0.892
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RNASLOpt
Matthews Correlation Coefficient
McQFold:
0.389
RNASLOpt:
0.364
Sensitivity
McQFold:
0.308
RNASLOpt:
0.261
Positive Predictive Value
McQFold:
0.492
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RNASLOpt
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNASLOpt:
0.364
Sensitivity
RSpredict(20):
0.241
RNASLOpt:
0.261
Positive Predictive Value
RSpredict(20):
0.590
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs RNASLOpt
Matthews Correlation Coefficient
RNAshapes:
0.367
RNASLOpt:
0.364
Sensitivity
RNAshapes:
0.285
RNASLOpt:
0.261
Positive Predictive Value
RNAshapes:
0.474
RNASLOpt:
0.510
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 7.04732813174e-08
|
|
+
RNASLOpt vs Multilign(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Multilign(20):
0.350
Sensitivity
RNASLOpt:
0.261
Multilign(20):
0.161
Positive Predictive Value
RNASLOpt:
0.510
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Murlet(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Murlet(20):
0.321
Sensitivity
RNASLOpt:
0.261
Murlet(20):
0.131
Positive Predictive Value
RNASLOpt:
0.510
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RNAwolf
Matthews Correlation Coefficient
RNASLOpt:
0.364
RNAwolf:
0.310
Sensitivity
RNASLOpt:
0.261
RNAwolf:
0.261
Positive Predictive Value
RNASLOpt:
0.510
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RSpredict(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
RSpredict(seed):
0.295
Sensitivity
RNASLOpt:
0.261
RSpredict(seed):
0.146
Positive Predictive Value
RNASLOpt:
0.510
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs CMfinder(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
CMfinder(20):
0.292
Sensitivity
RNASLOpt:
0.261
CMfinder(20):
0.107
Positive Predictive Value
RNASLOpt:
0.510
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Vsfold4
Matthews Correlation Coefficient
RNASLOpt:
0.364
Vsfold4:
0.270
Sensitivity
RNASLOpt:
0.261
Vsfold4:
0.198
Positive Predictive Value
RNASLOpt:
0.510
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Vsfold5
Matthews Correlation Coefficient
RNASLOpt:
0.364
Vsfold5:
0.260
Sensitivity
RNASLOpt:
0.261
Vsfold5:
0.195
Positive Predictive Value
RNASLOpt:
0.510
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RNASampler(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
RNASampler(20):
0.246
Sensitivity
RNASLOpt:
0.261
RNASampler(20):
0.069
Positive Predictive Value
RNASLOpt:
0.510
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs HotKnots
Matthews Correlation Coefficient
RNASLOpt:
0.364
HotKnots:
0.212
Sensitivity
RNASLOpt:
0.261
HotKnots:
0.069
Positive Predictive Value
RNASLOpt:
0.510
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Mastr(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Mastr(20):
0.187
Sensitivity
RNASLOpt:
0.261
Mastr(20):
0.044
Positive Predictive Value
RNASLOpt:
0.510
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Cylofold
Matthews Correlation Coefficient
RNASLOpt:
0.364
Cylofold:
0.178
Sensitivity
RNASLOpt:
0.261
Cylofold:
0.053
Positive Predictive Value
RNASLOpt:
0.510
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Pknots
Matthews Correlation Coefficient
RNASLOpt:
0.364
Pknots:
0.165
Sensitivity
RNASLOpt:
0.261
Pknots:
0.041
Positive Predictive Value
RNASLOpt:
0.510
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Alterna
Matthews Correlation Coefficient
RNASLOpt:
0.364
Alterna:
0.145
Sensitivity
RNASLOpt:
0.261
Alterna:
0.036
Positive Predictive Value
RNASLOpt:
0.510
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs MCFold
Matthews Correlation Coefficient
RNASLOpt:
0.364
MCFold:
0.138
Sensitivity
RNASLOpt:
0.261
MCFold:
0.044
Positive Predictive Value
RNASLOpt:
0.510
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RDfolder
Matthews Correlation Coefficient
RNASLOpt:
0.364
RDfolder:
0.138
Sensitivity
RNASLOpt:
0.261
RDfolder:
0.031
Positive Predictive Value
RNASLOpt:
0.510
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Murlet(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Murlet(seed):
0.110
Sensitivity
RNASLOpt:
0.261
Murlet(seed):
0.015
Positive Predictive Value
RNASLOpt:
0.510
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs CMfinder(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
CMfinder(seed):
0.111
Sensitivity
RNASLOpt:
0.261
CMfinder(seed):
0.016
Positive Predictive Value
RNASLOpt:
0.510
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs TurboFold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
TurboFold(seed):
0.106
Sensitivity
RNASLOpt:
0.261
TurboFold(seed):
0.014
Positive Predictive Value
RNASLOpt:
0.510
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs PPfold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
PPfold(seed):
0.097
Sensitivity
RNASLOpt:
0.261
PPfold(seed):
0.012
Positive Predictive Value
RNASLOpt:
0.510
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Carnac(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Carnac(seed):
0.093
Sensitivity
RNASLOpt:
0.261
Carnac(seed):
0.010
Positive Predictive Value
RNASLOpt:
0.510
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs RNASampler(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
RNASampler(seed):
0.088
Sensitivity
RNASLOpt:
0.261
RNASampler(seed):
0.009
Positive Predictive Value
RNASLOpt:
0.510
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Mastr(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Mastr(seed):
0.072
Sensitivity
RNASLOpt:
0.261
Mastr(seed):
0.006
Positive Predictive Value
RNASLOpt:
0.510
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs NanoFolder
Matthews Correlation Coefficient
RNASLOpt:
0.364
NanoFolder:
0.068
Sensitivity
RNASLOpt:
0.261
NanoFolder:
0.027
Positive Predictive Value
RNASLOpt:
0.510
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASLOpt vs Multilign(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Multilign(seed):
0.059
Sensitivity
RNASLOpt:
0.261
Multilign(seed):
0.005
Positive Predictive Value
RNASLOpt:
0.510
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Multilign(20) |
1242
ContextFold vs Multilign(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Multilign(20):
0.350
Sensitivity
ContextFold:
0.717
Multilign(20):
0.161
Positive Predictive Value
ContextFold:
0.794
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Multilign(20):
0.350
Sensitivity
PETfold_pre2.0(seed):
0.667
Multilign(20):
0.161
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Multilign(20):
0.350
Sensitivity
CentroidAlifold(seed):
0.414
Multilign(20):
0.161
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Multilign(20)
Matthews Correlation Coefficient
IPknot:
0.607
Multilign(20):
0.350
Sensitivity
IPknot:
0.551
Multilign(20):
0.161
Positive Predictive Value
IPknot:
0.670
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Multilign(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Multilign(20):
0.350
Sensitivity
PETfold_pre2.0(20):
0.410
Multilign(20):
0.161
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Multilign(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Multilign(20):
0.350
Sensitivity
CentroidFold:
0.515
Multilign(20):
0.161
Positive Predictive Value
CentroidFold:
0.633
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Multilign(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Multilign(20):
0.350
Sensitivity
CentroidAlifold(20):
0.369
Multilign(20):
0.161
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Multilign(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Multilign(20):
0.350
Sensitivity
CentroidHomfold‑LAST:
0.359
Multilign(20):
0.161
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Multilign(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Multilign(20):
0.350
Sensitivity
Contrafold:
0.542
Multilign(20):
0.161
Positive Predictive Value
Contrafold:
0.571
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Multilign(20):
0.350
Sensitivity
MXScarna(seed):
0.382
Multilign(20):
0.161
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Multilign(20):
0.350
Sensitivity
RNAalifold(20):
0.359
Multilign(20):
0.161
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Multilign(20)
Matthews Correlation Coefficient
Sfold:
0.527
Multilign(20):
0.350
Sensitivity
Sfold:
0.477
Multilign(20):
0.161
Positive Predictive Value
Sfold:
0.582
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Multilign(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Multilign(20):
0.350
Sensitivity
RNAalifold(seed):
0.328
Multilign(20):
0.161
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Multilign(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Multilign(20):
0.350
Sensitivity
MaxExpect:
0.505
Multilign(20):
0.161
Positive Predictive Value
MaxExpect:
0.528
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Multilign(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Multilign(20):
0.350
Sensitivity
ProbKnot:
0.508
Multilign(20):
0.161
Positive Predictive Value
ProbKnot:
0.517
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Multilign(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Multilign(20):
0.350
Sensitivity
MXScarna(20):
0.353
Multilign(20):
0.161
Positive Predictive Value
MXScarna(20):
0.710
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Multilign(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Multilign(20):
0.350
Sensitivity
UNAFold:
0.491
Multilign(20):
0.161
Positive Predictive Value
UNAFold:
0.498
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Multilign(20)
Matthews Correlation Coefficient
Fold:
0.493
Multilign(20):
0.350
Sensitivity
Fold:
0.496
Multilign(20):
0.161
Positive Predictive Value
Fold:
0.490
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Multilign(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Multilign(20):
0.350
Sensitivity
RNAfold:
0.486
Multilign(20):
0.161
Positive Predictive Value
RNAfold:
0.485
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Multilign(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Multilign(20):
0.350
Sensitivity
PknotsRG:
0.440
Multilign(20):
0.161
Positive Predictive Value
PknotsRG:
0.510
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Multilign(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Multilign(20):
0.350
Sensitivity
CRWrnafold:
0.379
Multilign(20):
0.161
Positive Predictive Value
CRWrnafold:
0.490
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Multilign(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Multilign(20):
0.350
Sensitivity
RNAsubopt:
0.344
Multilign(20):
0.161
Positive Predictive Value
RNAsubopt:
0.519
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Multilign(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Multilign(20):
0.350
Sensitivity
TurboFold(20):
0.210
Multilign(20):
0.161
Positive Predictive Value
TurboFold(20):
0.837
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Multilign(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Multilign(20):
0.350
Sensitivity
PPfold(20):
0.197
Multilign(20):
0.161
Positive Predictive Value
PPfold(20):
0.879
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Multilign(20)
Matthews Correlation Coefficient
Afold:
0.408
Multilign(20):
0.350
Sensitivity
Afold:
0.343
Multilign(20):
0.161
Positive Predictive Value
Afold:
0.487
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Multilign(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
Multilign(20):
0.350
Sensitivity
Carnac(20):
0.180
Multilign(20):
0.161
Positive Predictive Value
Carnac(20):
0.892
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Multilign(20)
Matthews Correlation Coefficient
McQFold:
0.389
Multilign(20):
0.350
Sensitivity
McQFold:
0.308
Multilign(20):
0.161
Positive Predictive Value
McQFold:
0.492
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Multilign(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Multilign(20):
0.350
Sensitivity
RSpredict(20):
0.241
Multilign(20):
0.161
Positive Predictive Value
RSpredict(20):
0.590
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Multilign(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
Multilign(20):
0.350
Sensitivity
RNAshapes:
0.285
Multilign(20):
0.161
Positive Predictive Value
RNAshapes:
0.474
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Multilign(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Multilign(20):
0.350
Sensitivity
RNASLOpt:
0.261
Multilign(20):
0.161
Positive Predictive Value
RNASLOpt:
0.510
Multilign(20):
0.762
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Multilign(20) vs Murlet(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
Murlet(20):
0.321
Sensitivity
Multilign(20):
0.161
Murlet(20):
0.131
Positive Predictive Value
Multilign(20):
0.762
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RNAwolf
Matthews Correlation Coefficient
Multilign(20):
0.350
RNAwolf:
0.310
Sensitivity
Multilign(20):
0.161
RNAwolf:
0.261
Positive Predictive Value
Multilign(20):
0.762
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
RSpredict(seed):
0.295
Sensitivity
Multilign(20):
0.161
RSpredict(seed):
0.146
Positive Predictive Value
Multilign(20):
0.762
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs CMfinder(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
CMfinder(20):
0.292
Sensitivity
Multilign(20):
0.161
CMfinder(20):
0.107
Positive Predictive Value
Multilign(20):
0.762
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Vsfold4
Matthews Correlation Coefficient
Multilign(20):
0.350
Vsfold4:
0.270
Sensitivity
Multilign(20):
0.161
Vsfold4:
0.198
Positive Predictive Value
Multilign(20):
0.762
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Vsfold5
Matthews Correlation Coefficient
Multilign(20):
0.350
Vsfold5:
0.260
Sensitivity
Multilign(20):
0.161
Vsfold5:
0.195
Positive Predictive Value
Multilign(20):
0.762
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RNASampler(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
RNASampler(20):
0.246
Sensitivity
Multilign(20):
0.161
RNASampler(20):
0.069
Positive Predictive Value
Multilign(20):
0.762
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs HotKnots
Matthews Correlation Coefficient
Multilign(20):
0.350
HotKnots:
0.212
Sensitivity
Multilign(20):
0.161
HotKnots:
0.069
Positive Predictive Value
Multilign(20):
0.762
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Mastr(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
Mastr(20):
0.187
Sensitivity
Multilign(20):
0.161
Mastr(20):
0.044
Positive Predictive Value
Multilign(20):
0.762
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Cylofold
Matthews Correlation Coefficient
Multilign(20):
0.350
Cylofold:
0.178
Sensitivity
Multilign(20):
0.161
Cylofold:
0.053
Positive Predictive Value
Multilign(20):
0.762
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Pknots
Matthews Correlation Coefficient
Multilign(20):
0.350
Pknots:
0.165
Sensitivity
Multilign(20):
0.161
Pknots:
0.041
Positive Predictive Value
Multilign(20):
0.762
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Alterna
Matthews Correlation Coefficient
Multilign(20):
0.350
Alterna:
0.145
Sensitivity
Multilign(20):
0.161
Alterna:
0.036
Positive Predictive Value
Multilign(20):
0.762
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs MCFold
Matthews Correlation Coefficient
Multilign(20):
0.350
MCFold:
0.138
Sensitivity
Multilign(20):
0.161
MCFold:
0.044
Positive Predictive Value
Multilign(20):
0.762
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RDfolder
Matthews Correlation Coefficient
Multilign(20):
0.350
RDfolder:
0.138
Sensitivity
Multilign(20):
0.161
RDfolder:
0.031
Positive Predictive Value
Multilign(20):
0.762
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Murlet(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Murlet(seed):
0.110
Sensitivity
Multilign(20):
0.161
Murlet(seed):
0.015
Positive Predictive Value
Multilign(20):
0.762
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
CMfinder(seed):
0.111
Sensitivity
Multilign(20):
0.161
CMfinder(seed):
0.016
Positive Predictive Value
Multilign(20):
0.762
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
TurboFold(seed):
0.106
Sensitivity
Multilign(20):
0.161
TurboFold(seed):
0.014
Positive Predictive Value
Multilign(20):
0.762
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs PPfold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
PPfold(seed):
0.097
Sensitivity
Multilign(20):
0.161
PPfold(seed):
0.012
Positive Predictive Value
Multilign(20):
0.762
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Carnac(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Carnac(seed):
0.093
Sensitivity
Multilign(20):
0.161
Carnac(seed):
0.010
Positive Predictive Value
Multilign(20):
0.762
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
RNASampler(seed):
0.088
Sensitivity
Multilign(20):
0.161
RNASampler(seed):
0.009
Positive Predictive Value
Multilign(20):
0.762
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Mastr(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Mastr(seed):
0.072
Sensitivity
Multilign(20):
0.161
Mastr(seed):
0.006
Positive Predictive Value
Multilign(20):
0.762
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs NanoFolder
Matthews Correlation Coefficient
Multilign(20):
0.350
NanoFolder:
0.068
Sensitivity
Multilign(20):
0.161
NanoFolder:
0.027
Positive Predictive Value
Multilign(20):
0.762
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Multilign(20) vs Multilign(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Multilign(seed):
0.059
Sensitivity
Multilign(20):
0.161
Multilign(seed):
0.005
Positive Predictive Value
Multilign(20):
0.762
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Murlet(20) |
1242
ContextFold vs Murlet(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Murlet(20):
0.321
Sensitivity
ContextFold:
0.717
Murlet(20):
0.131
Positive Predictive Value
ContextFold:
0.794
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Murlet(20):
0.321
Sensitivity
PETfold_pre2.0(seed):
0.667
Murlet(20):
0.131
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Murlet(20):
0.321
Sensitivity
CentroidAlifold(seed):
0.414
Murlet(20):
0.131
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Murlet(20)
Matthews Correlation Coefficient
IPknot:
0.607
Murlet(20):
0.321
Sensitivity
IPknot:
0.551
Murlet(20):
0.131
Positive Predictive Value
IPknot:
0.670
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Murlet(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Murlet(20):
0.321
Sensitivity
PETfold_pre2.0(20):
0.410
Murlet(20):
0.131
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Murlet(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Murlet(20):
0.321
Sensitivity
CentroidFold:
0.515
Murlet(20):
0.131
Positive Predictive Value
CentroidFold:
0.633
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Murlet(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Murlet(20):
0.321
Sensitivity
CentroidAlifold(20):
0.369
Murlet(20):
0.131
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Murlet(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Murlet(20):
0.321
Sensitivity
CentroidHomfold‑LAST:
0.359
Murlet(20):
0.131
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Murlet(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Murlet(20):
0.321
Sensitivity
Contrafold:
0.542
Murlet(20):
0.131
Positive Predictive Value
Contrafold:
0.571
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Murlet(20):
0.321
Sensitivity
MXScarna(seed):
0.382
Murlet(20):
0.131
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Murlet(20):
0.321
Sensitivity
RNAalifold(20):
0.359
Murlet(20):
0.131
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Murlet(20)
Matthews Correlation Coefficient
Sfold:
0.527
Murlet(20):
0.321
Sensitivity
Sfold:
0.477
Murlet(20):
0.131
Positive Predictive Value
Sfold:
0.582
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Murlet(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Murlet(20):
0.321
Sensitivity
RNAalifold(seed):
0.328
Murlet(20):
0.131
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Murlet(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Murlet(20):
0.321
Sensitivity
MaxExpect:
0.505
Murlet(20):
0.131
Positive Predictive Value
MaxExpect:
0.528
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Murlet(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Murlet(20):
0.321
Sensitivity
ProbKnot:
0.508
Murlet(20):
0.131
Positive Predictive Value
ProbKnot:
0.517
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Murlet(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Murlet(20):
0.321
Sensitivity
MXScarna(20):
0.353
Murlet(20):
0.131
Positive Predictive Value
MXScarna(20):
0.710
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Murlet(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Murlet(20):
0.321
Sensitivity
UNAFold:
0.491
Murlet(20):
0.131
Positive Predictive Value
UNAFold:
0.498
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Murlet(20)
Matthews Correlation Coefficient
Fold:
0.493
Murlet(20):
0.321
Sensitivity
Fold:
0.496
Murlet(20):
0.131
Positive Predictive Value
Fold:
0.490
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Murlet(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Murlet(20):
0.321
Sensitivity
RNAfold:
0.486
Murlet(20):
0.131
Positive Predictive Value
RNAfold:
0.485
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Murlet(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Murlet(20):
0.321
Sensitivity
PknotsRG:
0.440
Murlet(20):
0.131
Positive Predictive Value
PknotsRG:
0.510
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Murlet(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Murlet(20):
0.321
Sensitivity
CRWrnafold:
0.379
Murlet(20):
0.131
Positive Predictive Value
CRWrnafold:
0.490
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Murlet(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Murlet(20):
0.321
Sensitivity
RNAsubopt:
0.344
Murlet(20):
0.131
Positive Predictive Value
RNAsubopt:
0.519
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Murlet(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Murlet(20):
0.321
Sensitivity
TurboFold(20):
0.210
Murlet(20):
0.131
Positive Predictive Value
TurboFold(20):
0.837
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Murlet(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Murlet(20):
0.321
Sensitivity
PPfold(20):
0.197
Murlet(20):
0.131
Positive Predictive Value
PPfold(20):
0.879
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Murlet(20)
Matthews Correlation Coefficient
Afold:
0.408
Murlet(20):
0.321
Sensitivity
Afold:
0.343
Murlet(20):
0.131
Positive Predictive Value
Afold:
0.487
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Murlet(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
Murlet(20):
0.321
Sensitivity
Carnac(20):
0.180
Murlet(20):
0.131
Positive Predictive Value
Carnac(20):
0.892
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Murlet(20)
Matthews Correlation Coefficient
McQFold:
0.389
Murlet(20):
0.321
Sensitivity
McQFold:
0.308
Murlet(20):
0.131
Positive Predictive Value
McQFold:
0.492
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Murlet(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Murlet(20):
0.321
Sensitivity
RSpredict(20):
0.241
Murlet(20):
0.131
Positive Predictive Value
RSpredict(20):
0.590
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Murlet(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
Murlet(20):
0.321
Sensitivity
RNAshapes:
0.285
Murlet(20):
0.131
Positive Predictive Value
RNAshapes:
0.474
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Murlet(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Murlet(20):
0.321
Sensitivity
RNASLOpt:
0.261
Murlet(20):
0.131
Positive Predictive Value
RNASLOpt:
0.510
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Murlet(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
Murlet(20):
0.321
Sensitivity
Multilign(20):
0.161
Murlet(20):
0.131
Positive Predictive Value
Multilign(20):
0.762
Murlet(20):
0.787
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Murlet(20) vs RNAwolf
Matthews Correlation Coefficient
Murlet(20):
0.321
RNAwolf:
0.310
Sensitivity
Murlet(20):
0.131
RNAwolf:
0.261
Positive Predictive Value
Murlet(20):
0.787
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
RSpredict(seed):
0.295
Sensitivity
Murlet(20):
0.131
RSpredict(seed):
0.146
Positive Predictive Value
Murlet(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs CMfinder(20)
Matthews Correlation Coefficient
Murlet(20):
0.321
CMfinder(20):
0.292
Sensitivity
Murlet(20):
0.131
CMfinder(20):
0.107
Positive Predictive Value
Murlet(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Vsfold4
Matthews Correlation Coefficient
Murlet(20):
0.321
Vsfold4:
0.270
Sensitivity
Murlet(20):
0.131
Vsfold4:
0.198
Positive Predictive Value
Murlet(20):
0.787
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Vsfold5
Matthews Correlation Coefficient
Murlet(20):
0.321
Vsfold5:
0.260
Sensitivity
Murlet(20):
0.131
Vsfold5:
0.195
Positive Predictive Value
Murlet(20):
0.787
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RNASampler(20)
Matthews Correlation Coefficient
Murlet(20):
0.321
RNASampler(20):
0.246
Sensitivity
Murlet(20):
0.131
RNASampler(20):
0.069
Positive Predictive Value
Murlet(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs HotKnots
Matthews Correlation Coefficient
Murlet(20):
0.321
HotKnots:
0.212
Sensitivity
Murlet(20):
0.131
HotKnots:
0.069
Positive Predictive Value
Murlet(20):
0.787
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Mastr(20)
Matthews Correlation Coefficient
Murlet(20):
0.321
Mastr(20):
0.187
Sensitivity
Murlet(20):
0.131
Mastr(20):
0.044
Positive Predictive Value
Murlet(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Cylofold
Matthews Correlation Coefficient
Murlet(20):
0.321
Cylofold:
0.178
Sensitivity
Murlet(20):
0.131
Cylofold:
0.053
Positive Predictive Value
Murlet(20):
0.787
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Pknots
Matthews Correlation Coefficient
Murlet(20):
0.321
Pknots:
0.165
Sensitivity
Murlet(20):
0.131
Pknots:
0.041
Positive Predictive Value
Murlet(20):
0.787
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Alterna
Matthews Correlation Coefficient
Murlet(20):
0.321
Alterna:
0.145
Sensitivity
Murlet(20):
0.131
Alterna:
0.036
Positive Predictive Value
Murlet(20):
0.787
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs MCFold
Matthews Correlation Coefficient
Murlet(20):
0.321
MCFold:
0.138
Sensitivity
Murlet(20):
0.131
MCFold:
0.044
Positive Predictive Value
Murlet(20):
0.787
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RDfolder
Matthews Correlation Coefficient
Murlet(20):
0.321
RDfolder:
0.138
Sensitivity
Murlet(20):
0.131
RDfolder:
0.031
Positive Predictive Value
Murlet(20):
0.787
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Murlet(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Murlet(seed):
0.110
Sensitivity
Murlet(20):
0.131
Murlet(seed):
0.015
Positive Predictive Value
Murlet(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
CMfinder(seed):
0.111
Sensitivity
Murlet(20):
0.131
CMfinder(seed):
0.016
Positive Predictive Value
Murlet(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
TurboFold(seed):
0.106
Sensitivity
Murlet(20):
0.131
TurboFold(seed):
0.014
Positive Predictive Value
Murlet(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
PPfold(seed):
0.097
Sensitivity
Murlet(20):
0.131
PPfold(seed):
0.012
Positive Predictive Value
Murlet(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Carnac(seed):
0.093
Sensitivity
Murlet(20):
0.131
Carnac(seed):
0.010
Positive Predictive Value
Murlet(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
RNASampler(seed):
0.088
Sensitivity
Murlet(20):
0.131
RNASampler(seed):
0.009
Positive Predictive Value
Murlet(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Mastr(seed):
0.072
Sensitivity
Murlet(20):
0.131
Mastr(seed):
0.006
Positive Predictive Value
Murlet(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs NanoFolder
Matthews Correlation Coefficient
Murlet(20):
0.321
NanoFolder:
0.068
Sensitivity
Murlet(20):
0.131
NanoFolder:
0.027
Positive Predictive Value
Murlet(20):
0.787
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(20) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Multilign(seed):
0.059
Sensitivity
Murlet(20):
0.131
Multilign(seed):
0.005
Positive Predictive Value
Murlet(20):
0.787
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNAwolf |
1242
ContextFold vs RNAwolf
Matthews Correlation Coefficient
ContextFold:
0.754
RNAwolf:
0.310
Sensitivity
ContextFold:
0.717
RNAwolf:
0.261
Positive Predictive Value
ContextFold:
0.794
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNAwolf:
0.310
Sensitivity
PETfold_pre2.0(seed):
0.667
RNAwolf:
0.261
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNAwolf:
0.310
Sensitivity
CentroidAlifold(seed):
0.414
RNAwolf:
0.261
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNAwolf
Matthews Correlation Coefficient
IPknot:
0.607
RNAwolf:
0.310
Sensitivity
IPknot:
0.551
RNAwolf:
0.261
Positive Predictive Value
IPknot:
0.670
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNAwolf
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNAwolf:
0.310
Sensitivity
PETfold_pre2.0(20):
0.410
RNAwolf:
0.261
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNAwolf
Matthews Correlation Coefficient
CentroidFold:
0.571
RNAwolf:
0.310
Sensitivity
CentroidFold:
0.515
RNAwolf:
0.261
Positive Predictive Value
CentroidFold:
0.633
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNAwolf
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNAwolf:
0.310
Sensitivity
CentroidAlifold(20):
0.369
RNAwolf:
0.261
Positive Predictive Value
CentroidAlifold(20):
0.891
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNAwolf
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNAwolf:
0.310
Sensitivity
CentroidHomfold‑LAST:
0.359
RNAwolf:
0.261
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNAwolf
Matthews Correlation Coefficient
Contrafold:
0.556
RNAwolf:
0.310
Sensitivity
Contrafold:
0.542
RNAwolf:
0.261
Positive Predictive Value
Contrafold:
0.571
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNAwolf:
0.310
Sensitivity
MXScarna(seed):
0.382
RNAwolf:
0.261
Positive Predictive Value
MXScarna(seed):
0.753
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNAwolf:
0.310
Sensitivity
RNAalifold(20):
0.359
RNAwolf:
0.261
Positive Predictive Value
RNAalifold(20):
0.787
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNAwolf
Matthews Correlation Coefficient
Sfold:
0.527
RNAwolf:
0.310
Sensitivity
Sfold:
0.477
RNAwolf:
0.261
Positive Predictive Value
Sfold:
0.582
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNAwolf
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNAwolf:
0.310
Sensitivity
RNAalifold(seed):
0.328
RNAwolf:
0.261
Positive Predictive Value
RNAalifold(seed):
0.835
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNAwolf
Matthews Correlation Coefficient
MaxExpect:
0.516
RNAwolf:
0.310
Sensitivity
MaxExpect:
0.505
RNAwolf:
0.261
Positive Predictive Value
MaxExpect:
0.528
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNAwolf
Matthews Correlation Coefficient
ProbKnot:
0.512
RNAwolf:
0.310
Sensitivity
ProbKnot:
0.508
RNAwolf:
0.261
Positive Predictive Value
ProbKnot:
0.517
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNAwolf
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNAwolf:
0.310
Sensitivity
MXScarna(20):
0.353
RNAwolf:
0.261
Positive Predictive Value
MXScarna(20):
0.710
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNAwolf
Matthews Correlation Coefficient
UNAFold:
0.494
RNAwolf:
0.310
Sensitivity
UNAFold:
0.491
RNAwolf:
0.261
Positive Predictive Value
UNAFold:
0.498
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNAwolf
Matthews Correlation Coefficient
Fold:
0.493
RNAwolf:
0.310
Sensitivity
Fold:
0.496
RNAwolf:
0.261
Positive Predictive Value
Fold:
0.490
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RNAwolf
Matthews Correlation Coefficient
RNAfold:
0.485
RNAwolf:
0.310
Sensitivity
RNAfold:
0.486
RNAwolf:
0.261
Positive Predictive Value
RNAfold:
0.485
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RNAwolf
Matthews Correlation Coefficient
PknotsRG:
0.474
RNAwolf:
0.310
Sensitivity
PknotsRG:
0.440
RNAwolf:
0.261
Positive Predictive Value
PknotsRG:
0.510
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RNAwolf
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNAwolf:
0.310
Sensitivity
CRWrnafold:
0.379
RNAwolf:
0.261
Positive Predictive Value
CRWrnafold:
0.490
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RNAwolf
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNAwolf:
0.310
Sensitivity
RNAsubopt:
0.344
RNAwolf:
0.261
Positive Predictive Value
RNAsubopt:
0.519
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RNAwolf
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNAwolf:
0.310
Sensitivity
TurboFold(20):
0.210
RNAwolf:
0.261
Positive Predictive Value
TurboFold(20):
0.837
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RNAwolf
Matthews Correlation Coefficient
PPfold(20):
0.416
RNAwolf:
0.310
Sensitivity
PPfold(20):
0.197
RNAwolf:
0.261
Positive Predictive Value
PPfold(20):
0.879
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RNAwolf
Matthews Correlation Coefficient
Afold:
0.408
RNAwolf:
0.310
Sensitivity
Afold:
0.343
RNAwolf:
0.261
Positive Predictive Value
Afold:
0.487
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RNAwolf
Matthews Correlation Coefficient
Carnac(20):
0.401
RNAwolf:
0.310
Sensitivity
Carnac(20):
0.180
RNAwolf:
0.261
Positive Predictive Value
Carnac(20):
0.892
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RNAwolf
Matthews Correlation Coefficient
McQFold:
0.389
RNAwolf:
0.310
Sensitivity
McQFold:
0.308
RNAwolf:
0.261
Positive Predictive Value
McQFold:
0.492
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RNAwolf
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNAwolf:
0.310
Sensitivity
RSpredict(20):
0.241
RNAwolf:
0.261
Positive Predictive Value
RSpredict(20):
0.590
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs RNAwolf
Matthews Correlation Coefficient
RNAshapes:
0.367
RNAwolf:
0.310
Sensitivity
RNAshapes:
0.285
RNAwolf:
0.261
Positive Predictive Value
RNAshapes:
0.474
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs RNAwolf
Matthews Correlation Coefficient
RNASLOpt:
0.364
RNAwolf:
0.310
Sensitivity
RNASLOpt:
0.261
RNAwolf:
0.261
Positive Predictive Value
RNASLOpt:
0.510
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs RNAwolf
Matthews Correlation Coefficient
Multilign(20):
0.350
RNAwolf:
0.310
Sensitivity
Multilign(20):
0.161
RNAwolf:
0.261
Positive Predictive Value
Multilign(20):
0.762
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs RNAwolf
Matthews Correlation Coefficient
Murlet(20):
0.321
RNAwolf:
0.310
Sensitivity
Murlet(20):
0.131
RNAwolf:
0.261
Positive Predictive Value
Murlet(20):
0.787
RNAwolf:
0.369
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNAwolf vs RSpredict(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
RSpredict(seed):
0.295
Sensitivity
RNAwolf:
0.261
RSpredict(seed):
0.146
Positive Predictive Value
RNAwolf:
0.369
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs CMfinder(20)
Matthews Correlation Coefficient
RNAwolf:
0.310
CMfinder(20):
0.292
Sensitivity
RNAwolf:
0.261
CMfinder(20):
0.107
Positive Predictive Value
RNAwolf:
0.369
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Vsfold4
Matthews Correlation Coefficient
RNAwolf:
0.310
Vsfold4:
0.270
Sensitivity
RNAwolf:
0.261
Vsfold4:
0.198
Positive Predictive Value
RNAwolf:
0.369
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Vsfold5
Matthews Correlation Coefficient
RNAwolf:
0.310
Vsfold5:
0.260
Sensitivity
RNAwolf:
0.261
Vsfold5:
0.195
Positive Predictive Value
RNAwolf:
0.369
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RNASampler(20)
Matthews Correlation Coefficient
RNAwolf:
0.310
RNASampler(20):
0.246
Sensitivity
RNAwolf:
0.261
RNASampler(20):
0.069
Positive Predictive Value
RNAwolf:
0.369
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs HotKnots
Matthews Correlation Coefficient
RNAwolf:
0.310
HotKnots:
0.212
Sensitivity
RNAwolf:
0.261
HotKnots:
0.069
Positive Predictive Value
RNAwolf:
0.369
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Mastr(20)
Matthews Correlation Coefficient
RNAwolf:
0.310
Mastr(20):
0.187
Sensitivity
RNAwolf:
0.261
Mastr(20):
0.044
Positive Predictive Value
RNAwolf:
0.369
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Cylofold
Matthews Correlation Coefficient
RNAwolf:
0.310
Cylofold:
0.178
Sensitivity
RNAwolf:
0.261
Cylofold:
0.053
Positive Predictive Value
RNAwolf:
0.369
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Pknots
Matthews Correlation Coefficient
RNAwolf:
0.310
Pknots:
0.165
Sensitivity
RNAwolf:
0.261
Pknots:
0.041
Positive Predictive Value
RNAwolf:
0.369
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Alterna
Matthews Correlation Coefficient
RNAwolf:
0.310
Alterna:
0.145
Sensitivity
RNAwolf:
0.261
Alterna:
0.036
Positive Predictive Value
RNAwolf:
0.369
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs MCFold
Matthews Correlation Coefficient
RNAwolf:
0.310
MCFold:
0.138
Sensitivity
RNAwolf:
0.261
MCFold:
0.044
Positive Predictive Value
RNAwolf:
0.369
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RDfolder
Matthews Correlation Coefficient
RNAwolf:
0.310
RDfolder:
0.138
Sensitivity
RNAwolf:
0.261
RDfolder:
0.031
Positive Predictive Value
RNAwolf:
0.369
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Murlet(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Murlet(seed):
0.110
Sensitivity
RNAwolf:
0.261
Murlet(seed):
0.015
Positive Predictive Value
RNAwolf:
0.369
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs CMfinder(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
CMfinder(seed):
0.111
Sensitivity
RNAwolf:
0.261
CMfinder(seed):
0.016
Positive Predictive Value
RNAwolf:
0.369
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs TurboFold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
TurboFold(seed):
0.106
Sensitivity
RNAwolf:
0.261
TurboFold(seed):
0.014
Positive Predictive Value
RNAwolf:
0.369
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs PPfold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
PPfold(seed):
0.097
Sensitivity
RNAwolf:
0.261
PPfold(seed):
0.012
Positive Predictive Value
RNAwolf:
0.369
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Carnac(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Carnac(seed):
0.093
Sensitivity
RNAwolf:
0.261
Carnac(seed):
0.010
Positive Predictive Value
RNAwolf:
0.369
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs RNASampler(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
RNASampler(seed):
0.088
Sensitivity
RNAwolf:
0.261
RNASampler(seed):
0.009
Positive Predictive Value
RNAwolf:
0.369
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Mastr(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Mastr(seed):
0.072
Sensitivity
RNAwolf:
0.261
Mastr(seed):
0.006
Positive Predictive Value
RNAwolf:
0.369
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs NanoFolder
Matthews Correlation Coefficient
RNAwolf:
0.310
NanoFolder:
0.068
Sensitivity
RNAwolf:
0.261
NanoFolder:
0.027
Positive Predictive Value
RNAwolf:
0.369
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNAwolf vs Multilign(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Multilign(seed):
0.059
Sensitivity
RNAwolf:
0.261
Multilign(seed):
0.005
Positive Predictive Value
RNAwolf:
0.369
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RSpredict(seed) |
1242
ContextFold vs RSpredict(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
RSpredict(seed):
0.295
Sensitivity
ContextFold:
0.717
RSpredict(seed):
0.146
Positive Predictive Value
ContextFold:
0.794
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RSpredict(seed):
0.295
Sensitivity
PETfold_pre2.0(seed):
0.667
RSpredict(seed):
0.146
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RSpredict(seed):
0.295
Sensitivity
CentroidAlifold(seed):
0.414
RSpredict(seed):
0.146
Positive Predictive Value
CentroidAlifold(seed):
0.901
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RSpredict(seed)
Matthews Correlation Coefficient
IPknot:
0.607
RSpredict(seed):
0.295
Sensitivity
IPknot:
0.551
RSpredict(seed):
0.146
Positive Predictive Value
IPknot:
0.670
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RSpredict(seed):
0.295
Sensitivity
PETfold_pre2.0(20):
0.410
RSpredict(seed):
0.146
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
RSpredict(seed):
0.295
Sensitivity
CentroidFold:
0.515
RSpredict(seed):
0.146
Positive Predictive Value
CentroidFold:
0.633
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RSpredict(seed):
0.295
Sensitivity
CentroidAlifold(20):
0.369
RSpredict(seed):
0.146
Positive Predictive Value
CentroidAlifold(20):
0.891
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RSpredict(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RSpredict(seed):
0.295
Sensitivity
CentroidHomfold‑LAST:
0.359
RSpredict(seed):
0.146
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RSpredict(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
RSpredict(seed):
0.295
Sensitivity
Contrafold:
0.542
RSpredict(seed):
0.146
Positive Predictive Value
Contrafold:
0.571
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RSpredict(seed):
0.295
Sensitivity
MXScarna(seed):
0.382
RSpredict(seed):
0.146
Positive Predictive Value
MXScarna(seed):
0.753
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RSpredict(seed):
0.295
Sensitivity
RNAalifold(20):
0.359
RSpredict(seed):
0.146
Positive Predictive Value
RNAalifold(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RSpredict(seed)
Matthews Correlation Coefficient
Sfold:
0.527
RSpredict(seed):
0.295
Sensitivity
Sfold:
0.477
RSpredict(seed):
0.146
Positive Predictive Value
Sfold:
0.582
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RSpredict(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RSpredict(seed):
0.295
Sensitivity
RNAalifold(seed):
0.328
RSpredict(seed):
0.146
Positive Predictive Value
RNAalifold(seed):
0.835
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RSpredict(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
RSpredict(seed):
0.295
Sensitivity
MaxExpect:
0.505
RSpredict(seed):
0.146
Positive Predictive Value
MaxExpect:
0.528
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RSpredict(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
RSpredict(seed):
0.295
Sensitivity
ProbKnot:
0.508
RSpredict(seed):
0.146
Positive Predictive Value
ProbKnot:
0.517
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RSpredict(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RSpredict(seed):
0.295
Sensitivity
MXScarna(20):
0.353
RSpredict(seed):
0.146
Positive Predictive Value
MXScarna(20):
0.710
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RSpredict(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
RSpredict(seed):
0.295
Sensitivity
UNAFold:
0.491
RSpredict(seed):
0.146
Positive Predictive Value
UNAFold:
0.498
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RSpredict(seed)
Matthews Correlation Coefficient
Fold:
0.493
RSpredict(seed):
0.295
Sensitivity
Fold:
0.496
RSpredict(seed):
0.146
Positive Predictive Value
Fold:
0.490
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RSpredict(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
RSpredict(seed):
0.295
Sensitivity
RNAfold:
0.486
RSpredict(seed):
0.146
Positive Predictive Value
RNAfold:
0.485
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RSpredict(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
RSpredict(seed):
0.295
Sensitivity
PknotsRG:
0.440
RSpredict(seed):
0.146
Positive Predictive Value
PknotsRG:
0.510
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RSpredict(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RSpredict(seed):
0.295
Sensitivity
CRWrnafold:
0.379
RSpredict(seed):
0.146
Positive Predictive Value
CRWrnafold:
0.490
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RSpredict(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RSpredict(seed):
0.295
Sensitivity
RNAsubopt:
0.344
RSpredict(seed):
0.146
Positive Predictive Value
RNAsubopt:
0.519
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RSpredict(seed):
0.295
Sensitivity
TurboFold(20):
0.210
RSpredict(seed):
0.146
Positive Predictive Value
TurboFold(20):
0.837
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RSpredict(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
RSpredict(seed):
0.295
Sensitivity
PPfold(20):
0.197
RSpredict(seed):
0.146
Positive Predictive Value
PPfold(20):
0.879
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RSpredict(seed)
Matthews Correlation Coefficient
Afold:
0.408
RSpredict(seed):
0.295
Sensitivity
Afold:
0.343
RSpredict(seed):
0.146
Positive Predictive Value
Afold:
0.487
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
RSpredict(seed):
0.295
Sensitivity
Carnac(20):
0.180
RSpredict(seed):
0.146
Positive Predictive Value
Carnac(20):
0.892
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RSpredict(seed)
Matthews Correlation Coefficient
McQFold:
0.389
RSpredict(seed):
0.295
Sensitivity
McQFold:
0.308
RSpredict(seed):
0.146
Positive Predictive Value
McQFold:
0.492
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RSpredict(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
RSpredict(seed):
0.295
Sensitivity
RSpredict(20):
0.241
RSpredict(seed):
0.146
Positive Predictive Value
RSpredict(20):
0.590
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs RSpredict(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
RSpredict(seed):
0.295
Sensitivity
RNAshapes:
0.285
RSpredict(seed):
0.146
Positive Predictive Value
RNAshapes:
0.474
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs RSpredict(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
RSpredict(seed):
0.295
Sensitivity
RNASLOpt:
0.261
RSpredict(seed):
0.146
Positive Predictive Value
RNASLOpt:
0.510
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
RSpredict(seed):
0.295
Sensitivity
Multilign(20):
0.161
RSpredict(seed):
0.146
Positive Predictive Value
Multilign(20):
0.762
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs RSpredict(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
RSpredict(seed):
0.295
Sensitivity
Murlet(20):
0.131
RSpredict(seed):
0.146
Positive Predictive Value
Murlet(20):
0.787
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs RSpredict(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
RSpredict(seed):
0.295
Sensitivity
RNAwolf:
0.261
RSpredict(seed):
0.146
Positive Predictive Value
RNAwolf:
0.369
RSpredict(seed):
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RSpredict(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
CMfinder(20):
0.292
Sensitivity
RSpredict(seed):
0.146
CMfinder(20):
0.107
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.000108348164592
|
+
RSpredict(seed) vs Vsfold4
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Vsfold4:
0.270
Sensitivity
RSpredict(seed):
0.146
Vsfold4:
0.198
Positive Predictive Value
RSpredict(seed):
0.597
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Vsfold5:
0.260
Sensitivity
RSpredict(seed):
0.146
Vsfold5:
0.195
Positive Predictive Value
RSpredict(seed):
0.597
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
RNASampler(20):
0.246
Sensitivity
RSpredict(seed):
0.146
RNASampler(20):
0.069
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs HotKnots
Matthews Correlation Coefficient
RSpredict(seed):
0.295
HotKnots:
0.212
Sensitivity
RSpredict(seed):
0.146
HotKnots:
0.069
Positive Predictive Value
RSpredict(seed):
0.597
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Mastr(20):
0.187
Sensitivity
RSpredict(seed):
0.146
Mastr(20):
0.044
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Cylofold
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Cylofold:
0.178
Sensitivity
RSpredict(seed):
0.146
Cylofold:
0.053
Positive Predictive Value
RSpredict(seed):
0.597
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Pknots
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Pknots:
0.165
Sensitivity
RSpredict(seed):
0.146
Pknots:
0.041
Positive Predictive Value
RSpredict(seed):
0.597
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Alterna
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Alterna:
0.145
Sensitivity
RSpredict(seed):
0.146
Alterna:
0.036
Positive Predictive Value
RSpredict(seed):
0.597
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs MCFold
Matthews Correlation Coefficient
RSpredict(seed):
0.295
MCFold:
0.138
Sensitivity
RSpredict(seed):
0.146
MCFold:
0.044
Positive Predictive Value
RSpredict(seed):
0.597
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs RDfolder
Matthews Correlation Coefficient
RSpredict(seed):
0.295
RDfolder:
0.138
Sensitivity
RSpredict(seed):
0.146
RDfolder:
0.031
Positive Predictive Value
RSpredict(seed):
0.597
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Murlet(seed):
0.110
Sensitivity
RSpredict(seed):
0.146
Murlet(seed):
0.015
Positive Predictive Value
RSpredict(seed):
0.597
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
CMfinder(seed):
0.111
Sensitivity
RSpredict(seed):
0.146
CMfinder(seed):
0.016
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
TurboFold(seed):
0.106
Sensitivity
RSpredict(seed):
0.146
TurboFold(seed):
0.014
Positive Predictive Value
RSpredict(seed):
0.597
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
PPfold(seed):
0.097
Sensitivity
RSpredict(seed):
0.146
PPfold(seed):
0.012
Positive Predictive Value
RSpredict(seed):
0.597
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Carnac(seed):
0.093
Sensitivity
RSpredict(seed):
0.146
Carnac(seed):
0.010
Positive Predictive Value
RSpredict(seed):
0.597
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
RNASampler(seed):
0.088
Sensitivity
RSpredict(seed):
0.146
RNASampler(seed):
0.009
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Mastr(seed):
0.072
Sensitivity
RSpredict(seed):
0.146
Mastr(seed):
0.006
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(seed):
0.295
NanoFolder:
0.068
Sensitivity
RSpredict(seed):
0.146
NanoFolder:
0.027
Positive Predictive Value
RSpredict(seed):
0.597
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RSpredict(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Multilign(seed):
0.059
Sensitivity
RSpredict(seed):
0.146
Multilign(seed):
0.005
Positive Predictive Value
RSpredict(seed):
0.597
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CMfinder(20) |
1242
ContextFold vs CMfinder(20)
Matthews Correlation Coefficient
ContextFold:
0.754
CMfinder(20):
0.292
Sensitivity
ContextFold:
0.717
CMfinder(20):
0.107
Positive Predictive Value
ContextFold:
0.794
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CMfinder(20):
0.292
Sensitivity
PETfold_pre2.0(seed):
0.667
CMfinder(20):
0.107
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CMfinder(20):
0.292
Sensitivity
CentroidAlifold(seed):
0.414
CMfinder(20):
0.107
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs CMfinder(20)
Matthews Correlation Coefficient
IPknot:
0.607
CMfinder(20):
0.292
Sensitivity
IPknot:
0.551
CMfinder(20):
0.107
Positive Predictive Value
IPknot:
0.670
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs CMfinder(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CMfinder(20):
0.292
Sensitivity
PETfold_pre2.0(20):
0.410
CMfinder(20):
0.107
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs CMfinder(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
CMfinder(20):
0.292
Sensitivity
CentroidFold:
0.515
CMfinder(20):
0.107
Positive Predictive Value
CentroidFold:
0.633
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CMfinder(20):
0.292
Sensitivity
CentroidAlifold(20):
0.369
CMfinder(20):
0.107
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs CMfinder(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
CMfinder(20):
0.292
Sensitivity
CentroidHomfold‑LAST:
0.359
CMfinder(20):
0.107
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs CMfinder(20)
Matthews Correlation Coefficient
Contrafold:
0.556
CMfinder(20):
0.292
Sensitivity
Contrafold:
0.542
CMfinder(20):
0.107
Positive Predictive Value
Contrafold:
0.571
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
CMfinder(20):
0.292
Sensitivity
MXScarna(seed):
0.382
CMfinder(20):
0.107
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
CMfinder(20):
0.292
Sensitivity
RNAalifold(20):
0.359
CMfinder(20):
0.107
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs CMfinder(20)
Matthews Correlation Coefficient
Sfold:
0.527
CMfinder(20):
0.292
Sensitivity
Sfold:
0.477
CMfinder(20):
0.107
Positive Predictive Value
Sfold:
0.582
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
CMfinder(20):
0.292
Sensitivity
RNAalifold(seed):
0.328
CMfinder(20):
0.107
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs CMfinder(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
CMfinder(20):
0.292
Sensitivity
MaxExpect:
0.505
CMfinder(20):
0.107
Positive Predictive Value
MaxExpect:
0.528
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs CMfinder(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
CMfinder(20):
0.292
Sensitivity
ProbKnot:
0.508
CMfinder(20):
0.107
Positive Predictive Value
ProbKnot:
0.517
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs CMfinder(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
CMfinder(20):
0.292
Sensitivity
MXScarna(20):
0.353
CMfinder(20):
0.107
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs CMfinder(20)
Matthews Correlation Coefficient
UNAFold:
0.494
CMfinder(20):
0.292
Sensitivity
UNAFold:
0.491
CMfinder(20):
0.107
Positive Predictive Value
UNAFold:
0.498
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs CMfinder(20)
Matthews Correlation Coefficient
Fold:
0.493
CMfinder(20):
0.292
Sensitivity
Fold:
0.496
CMfinder(20):
0.107
Positive Predictive Value
Fold:
0.490
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs CMfinder(20)
Matthews Correlation Coefficient
RNAfold:
0.485
CMfinder(20):
0.292
Sensitivity
RNAfold:
0.486
CMfinder(20):
0.107
Positive Predictive Value
RNAfold:
0.485
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs CMfinder(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
CMfinder(20):
0.292
Sensitivity
PknotsRG:
0.440
CMfinder(20):
0.107
Positive Predictive Value
PknotsRG:
0.510
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs CMfinder(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
CMfinder(20):
0.292
Sensitivity
CRWrnafold:
0.379
CMfinder(20):
0.107
Positive Predictive Value
CRWrnafold:
0.490
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs CMfinder(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
CMfinder(20):
0.292
Sensitivity
RNAsubopt:
0.344
CMfinder(20):
0.107
Positive Predictive Value
RNAsubopt:
0.519
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs CMfinder(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
CMfinder(20):
0.292
Sensitivity
TurboFold(20):
0.210
CMfinder(20):
0.107
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs CMfinder(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
CMfinder(20):
0.292
Sensitivity
PPfold(20):
0.197
CMfinder(20):
0.107
Positive Predictive Value
PPfold(20):
0.879
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs CMfinder(20)
Matthews Correlation Coefficient
Afold:
0.408
CMfinder(20):
0.292
Sensitivity
Afold:
0.343
CMfinder(20):
0.107
Positive Predictive Value
Afold:
0.487
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs CMfinder(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
CMfinder(20):
0.292
Sensitivity
Carnac(20):
0.180
CMfinder(20):
0.107
Positive Predictive Value
Carnac(20):
0.892
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs CMfinder(20)
Matthews Correlation Coefficient
McQFold:
0.389
CMfinder(20):
0.292
Sensitivity
McQFold:
0.308
CMfinder(20):
0.107
Positive Predictive Value
McQFold:
0.492
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
CMfinder(20):
0.292
Sensitivity
RSpredict(20):
0.241
CMfinder(20):
0.107
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs CMfinder(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
CMfinder(20):
0.292
Sensitivity
RNAshapes:
0.285
CMfinder(20):
0.107
Positive Predictive Value
RNAshapes:
0.474
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs CMfinder(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
CMfinder(20):
0.292
Sensitivity
RNASLOpt:
0.261
CMfinder(20):
0.107
Positive Predictive Value
RNASLOpt:
0.510
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs CMfinder(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
CMfinder(20):
0.292
Sensitivity
Multilign(20):
0.161
CMfinder(20):
0.107
Positive Predictive Value
Multilign(20):
0.762
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs CMfinder(20)
Matthews Correlation Coefficient
Murlet(20):
0.321
CMfinder(20):
0.292
Sensitivity
Murlet(20):
0.131
CMfinder(20):
0.107
Positive Predictive Value
Murlet(20):
0.787
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs CMfinder(20)
Matthews Correlation Coefficient
RNAwolf:
0.310
CMfinder(20):
0.292
Sensitivity
RNAwolf:
0.261
CMfinder(20):
0.107
Positive Predictive Value
RNAwolf:
0.369
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs CMfinder(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
CMfinder(20):
0.292
Sensitivity
RSpredict(seed):
0.146
CMfinder(20):
0.107
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(20):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.000108348164592
|
|
+
CMfinder(20) vs Vsfold4
Matthews Correlation Coefficient
CMfinder(20):
0.292
Vsfold4:
0.270
Sensitivity
CMfinder(20):
0.107
Vsfold4:
0.198
Positive Predictive Value
CMfinder(20):
0.794
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Vsfold5
Matthews Correlation Coefficient
CMfinder(20):
0.292
Vsfold5:
0.260
Sensitivity
CMfinder(20):
0.107
Vsfold5:
0.195
Positive Predictive Value
CMfinder(20):
0.794
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs RNASampler(20)
Matthews Correlation Coefficient
CMfinder(20):
0.292
RNASampler(20):
0.246
Sensitivity
CMfinder(20):
0.107
RNASampler(20):
0.069
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs HotKnots
Matthews Correlation Coefficient
CMfinder(20):
0.292
HotKnots:
0.212
Sensitivity
CMfinder(20):
0.107
HotKnots:
0.069
Positive Predictive Value
CMfinder(20):
0.794
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Mastr(20)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Mastr(20):
0.187
Sensitivity
CMfinder(20):
0.107
Mastr(20):
0.044
Positive Predictive Value
CMfinder(20):
0.794
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Cylofold
Matthews Correlation Coefficient
CMfinder(20):
0.292
Cylofold:
0.178
Sensitivity
CMfinder(20):
0.107
Cylofold:
0.053
Positive Predictive Value
CMfinder(20):
0.794
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Pknots
Matthews Correlation Coefficient
CMfinder(20):
0.292
Pknots:
0.165
Sensitivity
CMfinder(20):
0.107
Pknots:
0.041
Positive Predictive Value
CMfinder(20):
0.794
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Alterna
Matthews Correlation Coefficient
CMfinder(20):
0.292
Alterna:
0.145
Sensitivity
CMfinder(20):
0.107
Alterna:
0.036
Positive Predictive Value
CMfinder(20):
0.794
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs MCFold
Matthews Correlation Coefficient
CMfinder(20):
0.292
MCFold:
0.138
Sensitivity
CMfinder(20):
0.107
MCFold:
0.044
Positive Predictive Value
CMfinder(20):
0.794
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs RDfolder
Matthews Correlation Coefficient
CMfinder(20):
0.292
RDfolder:
0.138
Sensitivity
CMfinder(20):
0.107
RDfolder:
0.031
Positive Predictive Value
CMfinder(20):
0.794
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Murlet(seed):
0.110
Sensitivity
CMfinder(20):
0.107
Murlet(seed):
0.015
Positive Predictive Value
CMfinder(20):
0.794
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
CMfinder(seed):
0.111
Sensitivity
CMfinder(20):
0.107
CMfinder(seed):
0.016
Positive Predictive Value
CMfinder(20):
0.794
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
TurboFold(seed):
0.106
Sensitivity
CMfinder(20):
0.107
TurboFold(seed):
0.014
Positive Predictive Value
CMfinder(20):
0.794
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
PPfold(seed):
0.097
Sensitivity
CMfinder(20):
0.107
PPfold(seed):
0.012
Positive Predictive Value
CMfinder(20):
0.794
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Carnac(seed):
0.093
Sensitivity
CMfinder(20):
0.107
Carnac(seed):
0.010
Positive Predictive Value
CMfinder(20):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
RNASampler(seed):
0.088
Sensitivity
CMfinder(20):
0.107
RNASampler(seed):
0.009
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Mastr(seed):
0.072
Sensitivity
CMfinder(20):
0.107
Mastr(seed):
0.006
Positive Predictive Value
CMfinder(20):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(20):
0.292
NanoFolder:
0.068
Sensitivity
CMfinder(20):
0.107
NanoFolder:
0.027
Positive Predictive Value
CMfinder(20):
0.794
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(20) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Multilign(seed):
0.059
Sensitivity
CMfinder(20):
0.107
Multilign(seed):
0.005
Positive Predictive Value
CMfinder(20):
0.794
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Vsfold4 |
1242
ContextFold vs Vsfold4
Matthews Correlation Coefficient
ContextFold:
0.754
Vsfold4:
0.270
Sensitivity
ContextFold:
0.717
Vsfold4:
0.198
Positive Predictive Value
ContextFold:
0.794
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Vsfold4:
0.270
Sensitivity
PETfold_pre2.0(seed):
0.667
Vsfold4:
0.198
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Vsfold4:
0.270
Sensitivity
CentroidAlifold(seed):
0.414
Vsfold4:
0.198
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Vsfold4
Matthews Correlation Coefficient
IPknot:
0.607
Vsfold4:
0.270
Sensitivity
IPknot:
0.551
Vsfold4:
0.198
Positive Predictive Value
IPknot:
0.670
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Vsfold4
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Vsfold4:
0.270
Sensitivity
PETfold_pre2.0(20):
0.410
Vsfold4:
0.198
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Vsfold4
Matthews Correlation Coefficient
CentroidFold:
0.571
Vsfold4:
0.270
Sensitivity
CentroidFold:
0.515
Vsfold4:
0.198
Positive Predictive Value
CentroidFold:
0.633
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Vsfold4
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Vsfold4:
0.270
Sensitivity
CentroidAlifold(20):
0.369
Vsfold4:
0.198
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Vsfold4
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Vsfold4:
0.270
Sensitivity
CentroidHomfold‑LAST:
0.359
Vsfold4:
0.198
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Vsfold4
Matthews Correlation Coefficient
Contrafold:
0.556
Vsfold4:
0.270
Sensitivity
Contrafold:
0.542
Vsfold4:
0.198
Positive Predictive Value
Contrafold:
0.571
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Vsfold4:
0.270
Sensitivity
MXScarna(seed):
0.382
Vsfold4:
0.198
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Vsfold4:
0.270
Sensitivity
RNAalifold(20):
0.359
Vsfold4:
0.198
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Vsfold4
Matthews Correlation Coefficient
Sfold:
0.527
Vsfold4:
0.270
Sensitivity
Sfold:
0.477
Vsfold4:
0.198
Positive Predictive Value
Sfold:
0.582
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Vsfold4
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Vsfold4:
0.270
Sensitivity
RNAalifold(seed):
0.328
Vsfold4:
0.198
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Vsfold4
Matthews Correlation Coefficient
MaxExpect:
0.516
Vsfold4:
0.270
Sensitivity
MaxExpect:
0.505
Vsfold4:
0.198
Positive Predictive Value
MaxExpect:
0.528
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Vsfold4
Matthews Correlation Coefficient
ProbKnot:
0.512
Vsfold4:
0.270
Sensitivity
ProbKnot:
0.508
Vsfold4:
0.198
Positive Predictive Value
ProbKnot:
0.517
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Vsfold4
Matthews Correlation Coefficient
MXScarna(20):
0.500
Vsfold4:
0.270
Sensitivity
MXScarna(20):
0.353
Vsfold4:
0.198
Positive Predictive Value
MXScarna(20):
0.710
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Vsfold4
Matthews Correlation Coefficient
UNAFold:
0.494
Vsfold4:
0.270
Sensitivity
UNAFold:
0.491
Vsfold4:
0.198
Positive Predictive Value
UNAFold:
0.498
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Vsfold4
Matthews Correlation Coefficient
Fold:
0.493
Vsfold4:
0.270
Sensitivity
Fold:
0.496
Vsfold4:
0.198
Positive Predictive Value
Fold:
0.490
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Vsfold4
Matthews Correlation Coefficient
RNAfold:
0.485
Vsfold4:
0.270
Sensitivity
RNAfold:
0.486
Vsfold4:
0.198
Positive Predictive Value
RNAfold:
0.485
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Vsfold4
Matthews Correlation Coefficient
PknotsRG:
0.474
Vsfold4:
0.270
Sensitivity
PknotsRG:
0.440
Vsfold4:
0.198
Positive Predictive Value
PknotsRG:
0.510
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Vsfold4
Matthews Correlation Coefficient
CRWrnafold:
0.430
Vsfold4:
0.270
Sensitivity
CRWrnafold:
0.379
Vsfold4:
0.198
Positive Predictive Value
CRWrnafold:
0.490
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Vsfold4
Matthews Correlation Coefficient
RNAsubopt:
0.422
Vsfold4:
0.270
Sensitivity
RNAsubopt:
0.344
Vsfold4:
0.198
Positive Predictive Value
RNAsubopt:
0.519
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Vsfold4
Matthews Correlation Coefficient
TurboFold(20):
0.419
Vsfold4:
0.270
Sensitivity
TurboFold(20):
0.210
Vsfold4:
0.198
Positive Predictive Value
TurboFold(20):
0.837
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Vsfold4
Matthews Correlation Coefficient
PPfold(20):
0.416
Vsfold4:
0.270
Sensitivity
PPfold(20):
0.197
Vsfold4:
0.198
Positive Predictive Value
PPfold(20):
0.879
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Vsfold4
Matthews Correlation Coefficient
Afold:
0.408
Vsfold4:
0.270
Sensitivity
Afold:
0.343
Vsfold4:
0.198
Positive Predictive Value
Afold:
0.487
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Vsfold4
Matthews Correlation Coefficient
Carnac(20):
0.401
Vsfold4:
0.270
Sensitivity
Carnac(20):
0.180
Vsfold4:
0.198
Positive Predictive Value
Carnac(20):
0.892
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Vsfold4
Matthews Correlation Coefficient
McQFold:
0.389
Vsfold4:
0.270
Sensitivity
McQFold:
0.308
Vsfold4:
0.198
Positive Predictive Value
McQFold:
0.492
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Vsfold4
Matthews Correlation Coefficient
RSpredict(20):
0.377
Vsfold4:
0.270
Sensitivity
RSpredict(20):
0.241
Vsfold4:
0.198
Positive Predictive Value
RSpredict(20):
0.590
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Vsfold4
Matthews Correlation Coefficient
RNAshapes:
0.367
Vsfold4:
0.270
Sensitivity
RNAshapes:
0.285
Vsfold4:
0.198
Positive Predictive Value
RNAshapes:
0.474
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Vsfold4
Matthews Correlation Coefficient
RNASLOpt:
0.364
Vsfold4:
0.270
Sensitivity
RNASLOpt:
0.261
Vsfold4:
0.198
Positive Predictive Value
RNASLOpt:
0.510
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Vsfold4
Matthews Correlation Coefficient
Multilign(20):
0.350
Vsfold4:
0.270
Sensitivity
Multilign(20):
0.161
Vsfold4:
0.198
Positive Predictive Value
Multilign(20):
0.762
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Vsfold4
Matthews Correlation Coefficient
Murlet(20):
0.321
Vsfold4:
0.270
Sensitivity
Murlet(20):
0.131
Vsfold4:
0.198
Positive Predictive Value
Murlet(20):
0.787
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Vsfold4
Matthews Correlation Coefficient
RNAwolf:
0.310
Vsfold4:
0.270
Sensitivity
RNAwolf:
0.261
Vsfold4:
0.198
Positive Predictive Value
RNAwolf:
0.369
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Vsfold4
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Vsfold4:
0.270
Sensitivity
RSpredict(seed):
0.146
Vsfold4:
0.198
Positive Predictive Value
RSpredict(seed):
0.597
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Vsfold4
Matthews Correlation Coefficient
CMfinder(20):
0.292
Vsfold4:
0.270
Sensitivity
CMfinder(20):
0.107
Vsfold4:
0.198
Positive Predictive Value
CMfinder(20):
0.794
Vsfold4:
0.371
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Vsfold4 vs Vsfold5
Matthews Correlation Coefficient
Vsfold4:
0.270
Vsfold5:
0.260
Sensitivity
Vsfold4:
0.198
Vsfold5:
0.195
Positive Predictive Value
Vsfold4:
0.371
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold4:
0.270
RNASampler(20):
0.246
Sensitivity
Vsfold4:
0.198
RNASampler(20):
0.069
Positive Predictive Value
Vsfold4:
0.371
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs HotKnots
Matthews Correlation Coefficient
Vsfold4:
0.270
HotKnots:
0.212
Sensitivity
Vsfold4:
0.198
HotKnots:
0.069
Positive Predictive Value
Vsfold4:
0.371
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold4:
0.270
Mastr(20):
0.187
Sensitivity
Vsfold4:
0.198
Mastr(20):
0.044
Positive Predictive Value
Vsfold4:
0.371
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Cylofold
Matthews Correlation Coefficient
Vsfold4:
0.270
Cylofold:
0.178
Sensitivity
Vsfold4:
0.198
Cylofold:
0.053
Positive Predictive Value
Vsfold4:
0.371
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Pknots
Matthews Correlation Coefficient
Vsfold4:
0.270
Pknots:
0.165
Sensitivity
Vsfold4:
0.198
Pknots:
0.041
Positive Predictive Value
Vsfold4:
0.371
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Alterna
Matthews Correlation Coefficient
Vsfold4:
0.270
Alterna:
0.145
Sensitivity
Vsfold4:
0.198
Alterna:
0.036
Positive Predictive Value
Vsfold4:
0.371
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs MCFold
Matthews Correlation Coefficient
Vsfold4:
0.270
MCFold:
0.138
Sensitivity
Vsfold4:
0.198
MCFold:
0.044
Positive Predictive Value
Vsfold4:
0.371
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RDfolder
Matthews Correlation Coefficient
Vsfold4:
0.270
RDfolder:
0.138
Sensitivity
Vsfold4:
0.198
RDfolder:
0.031
Positive Predictive Value
Vsfold4:
0.371
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Murlet(seed):
0.110
Sensitivity
Vsfold4:
0.198
Murlet(seed):
0.015
Positive Predictive Value
Vsfold4:
0.371
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
CMfinder(seed):
0.111
Sensitivity
Vsfold4:
0.198
CMfinder(seed):
0.016
Positive Predictive Value
Vsfold4:
0.371
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
TurboFold(seed):
0.106
Sensitivity
Vsfold4:
0.198
TurboFold(seed):
0.014
Positive Predictive Value
Vsfold4:
0.371
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
PPfold(seed):
0.097
Sensitivity
Vsfold4:
0.198
PPfold(seed):
0.012
Positive Predictive Value
Vsfold4:
0.371
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Carnac(seed):
0.093
Sensitivity
Vsfold4:
0.198
Carnac(seed):
0.010
Positive Predictive Value
Vsfold4:
0.371
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
RNASampler(seed):
0.088
Sensitivity
Vsfold4:
0.198
RNASampler(seed):
0.009
Positive Predictive Value
Vsfold4:
0.371
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Mastr(seed):
0.072
Sensitivity
Vsfold4:
0.198
Mastr(seed):
0.006
Positive Predictive Value
Vsfold4:
0.371
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs NanoFolder
Matthews Correlation Coefficient
Vsfold4:
0.270
NanoFolder:
0.068
Sensitivity
Vsfold4:
0.198
NanoFolder:
0.027
Positive Predictive Value
Vsfold4:
0.371
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold4 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Multilign(seed):
0.059
Sensitivity
Vsfold4:
0.198
Multilign(seed):
0.005
Positive Predictive Value
Vsfold4:
0.371
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Vsfold5 |
1242
ContextFold vs Vsfold5
Matthews Correlation Coefficient
ContextFold:
0.754
Vsfold5:
0.260
Sensitivity
ContextFold:
0.717
Vsfold5:
0.195
Positive Predictive Value
ContextFold:
0.794
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Vsfold5:
0.260
Sensitivity
PETfold_pre2.0(seed):
0.667
Vsfold5:
0.195
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Vsfold5:
0.260
Sensitivity
CentroidAlifold(seed):
0.414
Vsfold5:
0.195
Positive Predictive Value
CentroidAlifold(seed):
0.901
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Vsfold5
Matthews Correlation Coefficient
IPknot:
0.607
Vsfold5:
0.260
Sensitivity
IPknot:
0.551
Vsfold5:
0.195
Positive Predictive Value
IPknot:
0.670
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Vsfold5
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Vsfold5:
0.260
Sensitivity
PETfold_pre2.0(20):
0.410
Vsfold5:
0.195
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Vsfold5
Matthews Correlation Coefficient
CentroidFold:
0.571
Vsfold5:
0.260
Sensitivity
CentroidFold:
0.515
Vsfold5:
0.195
Positive Predictive Value
CentroidFold:
0.633
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Vsfold5
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Vsfold5:
0.260
Sensitivity
CentroidAlifold(20):
0.369
Vsfold5:
0.195
Positive Predictive Value
CentroidAlifold(20):
0.891
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Vsfold5
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Vsfold5:
0.260
Sensitivity
CentroidHomfold‑LAST:
0.359
Vsfold5:
0.195
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Vsfold5
Matthews Correlation Coefficient
Contrafold:
0.556
Vsfold5:
0.260
Sensitivity
Contrafold:
0.542
Vsfold5:
0.195
Positive Predictive Value
Contrafold:
0.571
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Vsfold5:
0.260
Sensitivity
MXScarna(seed):
0.382
Vsfold5:
0.195
Positive Predictive Value
MXScarna(seed):
0.753
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Vsfold5:
0.260
Sensitivity
RNAalifold(20):
0.359
Vsfold5:
0.195
Positive Predictive Value
RNAalifold(20):
0.787
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Vsfold5
Matthews Correlation Coefficient
Sfold:
0.527
Vsfold5:
0.260
Sensitivity
Sfold:
0.477
Vsfold5:
0.195
Positive Predictive Value
Sfold:
0.582
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Vsfold5
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Vsfold5:
0.260
Sensitivity
RNAalifold(seed):
0.328
Vsfold5:
0.195
Positive Predictive Value
RNAalifold(seed):
0.835
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Vsfold5
Matthews Correlation Coefficient
MaxExpect:
0.516
Vsfold5:
0.260
Sensitivity
MaxExpect:
0.505
Vsfold5:
0.195
Positive Predictive Value
MaxExpect:
0.528
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Vsfold5
Matthews Correlation Coefficient
ProbKnot:
0.512
Vsfold5:
0.260
Sensitivity
ProbKnot:
0.508
Vsfold5:
0.195
Positive Predictive Value
ProbKnot:
0.517
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Vsfold5
Matthews Correlation Coefficient
MXScarna(20):
0.500
Vsfold5:
0.260
Sensitivity
MXScarna(20):
0.353
Vsfold5:
0.195
Positive Predictive Value
MXScarna(20):
0.710
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Vsfold5
Matthews Correlation Coefficient
UNAFold:
0.494
Vsfold5:
0.260
Sensitivity
UNAFold:
0.491
Vsfold5:
0.195
Positive Predictive Value
UNAFold:
0.498
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Vsfold5
Matthews Correlation Coefficient
Fold:
0.493
Vsfold5:
0.260
Sensitivity
Fold:
0.496
Vsfold5:
0.195
Positive Predictive Value
Fold:
0.490
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Vsfold5
Matthews Correlation Coefficient
RNAfold:
0.485
Vsfold5:
0.260
Sensitivity
RNAfold:
0.486
Vsfold5:
0.195
Positive Predictive Value
RNAfold:
0.485
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Vsfold5
Matthews Correlation Coefficient
PknotsRG:
0.474
Vsfold5:
0.260
Sensitivity
PknotsRG:
0.440
Vsfold5:
0.195
Positive Predictive Value
PknotsRG:
0.510
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Vsfold5
Matthews Correlation Coefficient
CRWrnafold:
0.430
Vsfold5:
0.260
Sensitivity
CRWrnafold:
0.379
Vsfold5:
0.195
Positive Predictive Value
CRWrnafold:
0.490
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Vsfold5
Matthews Correlation Coefficient
RNAsubopt:
0.422
Vsfold5:
0.260
Sensitivity
RNAsubopt:
0.344
Vsfold5:
0.195
Positive Predictive Value
RNAsubopt:
0.519
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Vsfold5
Matthews Correlation Coefficient
TurboFold(20):
0.419
Vsfold5:
0.260
Sensitivity
TurboFold(20):
0.210
Vsfold5:
0.195
Positive Predictive Value
TurboFold(20):
0.837
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Vsfold5
Matthews Correlation Coefficient
PPfold(20):
0.416
Vsfold5:
0.260
Sensitivity
PPfold(20):
0.197
Vsfold5:
0.195
Positive Predictive Value
PPfold(20):
0.879
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Vsfold5
Matthews Correlation Coefficient
Afold:
0.408
Vsfold5:
0.260
Sensitivity
Afold:
0.343
Vsfold5:
0.195
Positive Predictive Value
Afold:
0.487
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Vsfold5
Matthews Correlation Coefficient
Carnac(20):
0.401
Vsfold5:
0.260
Sensitivity
Carnac(20):
0.180
Vsfold5:
0.195
Positive Predictive Value
Carnac(20):
0.892
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Vsfold5
Matthews Correlation Coefficient
McQFold:
0.389
Vsfold5:
0.260
Sensitivity
McQFold:
0.308
Vsfold5:
0.195
Positive Predictive Value
McQFold:
0.492
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(20):
0.377
Vsfold5:
0.260
Sensitivity
RSpredict(20):
0.241
Vsfold5:
0.195
Positive Predictive Value
RSpredict(20):
0.590
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Vsfold5
Matthews Correlation Coefficient
RNAshapes:
0.367
Vsfold5:
0.260
Sensitivity
RNAshapes:
0.285
Vsfold5:
0.195
Positive Predictive Value
RNAshapes:
0.474
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Vsfold5
Matthews Correlation Coefficient
RNASLOpt:
0.364
Vsfold5:
0.260
Sensitivity
RNASLOpt:
0.261
Vsfold5:
0.195
Positive Predictive Value
RNASLOpt:
0.510
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Vsfold5
Matthews Correlation Coefficient
Multilign(20):
0.350
Vsfold5:
0.260
Sensitivity
Multilign(20):
0.161
Vsfold5:
0.195
Positive Predictive Value
Multilign(20):
0.762
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Vsfold5
Matthews Correlation Coefficient
Murlet(20):
0.321
Vsfold5:
0.260
Sensitivity
Murlet(20):
0.131
Vsfold5:
0.195
Positive Predictive Value
Murlet(20):
0.787
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Vsfold5
Matthews Correlation Coefficient
RNAwolf:
0.310
Vsfold5:
0.260
Sensitivity
RNAwolf:
0.261
Vsfold5:
0.195
Positive Predictive Value
RNAwolf:
0.369
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Vsfold5
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Vsfold5:
0.260
Sensitivity
RSpredict(seed):
0.146
Vsfold5:
0.195
Positive Predictive Value
RSpredict(seed):
0.597
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Vsfold5
Matthews Correlation Coefficient
CMfinder(20):
0.292
Vsfold5:
0.260
Sensitivity
CMfinder(20):
0.107
Vsfold5:
0.195
Positive Predictive Value
CMfinder(20):
0.794
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Vsfold5
Matthews Correlation Coefficient
Vsfold4:
0.270
Vsfold5:
0.260
Sensitivity
Vsfold4:
0.198
Vsfold5:
0.195
Positive Predictive Value
Vsfold4:
0.371
Vsfold5:
0.347
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Vsfold5 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold5:
0.260
RNASampler(20):
0.246
Sensitivity
Vsfold5:
0.195
RNASampler(20):
0.069
Positive Predictive Value
Vsfold5:
0.347
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs HotKnots
Matthews Correlation Coefficient
Vsfold5:
0.260
HotKnots:
0.212
Sensitivity
Vsfold5:
0.195
HotKnots:
0.069
Positive Predictive Value
Vsfold5:
0.347
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold5:
0.260
Mastr(20):
0.187
Sensitivity
Vsfold5:
0.195
Mastr(20):
0.044
Positive Predictive Value
Vsfold5:
0.347
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Cylofold
Matthews Correlation Coefficient
Vsfold5:
0.260
Cylofold:
0.178
Sensitivity
Vsfold5:
0.195
Cylofold:
0.053
Positive Predictive Value
Vsfold5:
0.347
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Pknots
Matthews Correlation Coefficient
Vsfold5:
0.260
Pknots:
0.165
Sensitivity
Vsfold5:
0.195
Pknots:
0.041
Positive Predictive Value
Vsfold5:
0.347
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Alterna
Matthews Correlation Coefficient
Vsfold5:
0.260
Alterna:
0.145
Sensitivity
Vsfold5:
0.195
Alterna:
0.036
Positive Predictive Value
Vsfold5:
0.347
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs MCFold
Matthews Correlation Coefficient
Vsfold5:
0.260
MCFold:
0.138
Sensitivity
Vsfold5:
0.195
MCFold:
0.044
Positive Predictive Value
Vsfold5:
0.347
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs RDfolder
Matthews Correlation Coefficient
Vsfold5:
0.260
RDfolder:
0.138
Sensitivity
Vsfold5:
0.195
RDfolder:
0.031
Positive Predictive Value
Vsfold5:
0.347
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Murlet(seed):
0.110
Sensitivity
Vsfold5:
0.195
Murlet(seed):
0.015
Positive Predictive Value
Vsfold5:
0.347
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
CMfinder(seed):
0.111
Sensitivity
Vsfold5:
0.195
CMfinder(seed):
0.016
Positive Predictive Value
Vsfold5:
0.347
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
TurboFold(seed):
0.106
Sensitivity
Vsfold5:
0.195
TurboFold(seed):
0.014
Positive Predictive Value
Vsfold5:
0.347
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
PPfold(seed):
0.097
Sensitivity
Vsfold5:
0.195
PPfold(seed):
0.012
Positive Predictive Value
Vsfold5:
0.347
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Carnac(seed):
0.093
Sensitivity
Vsfold5:
0.195
Carnac(seed):
0.010
Positive Predictive Value
Vsfold5:
0.347
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
RNASampler(seed):
0.088
Sensitivity
Vsfold5:
0.195
RNASampler(seed):
0.009
Positive Predictive Value
Vsfold5:
0.347
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Mastr(seed):
0.072
Sensitivity
Vsfold5:
0.195
Mastr(seed):
0.006
Positive Predictive Value
Vsfold5:
0.347
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs NanoFolder
Matthews Correlation Coefficient
Vsfold5:
0.260
NanoFolder:
0.068
Sensitivity
Vsfold5:
0.195
NanoFolder:
0.027
Positive Predictive Value
Vsfold5:
0.347
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Vsfold5 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Multilign(seed):
0.059
Sensitivity
Vsfold5:
0.195
Multilign(seed):
0.005
Positive Predictive Value
Vsfold5:
0.347
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNASampler(20) |
1242
ContextFold vs RNASampler(20)
Matthews Correlation Coefficient
ContextFold:
0.754
RNASampler(20):
0.246
Sensitivity
ContextFold:
0.717
RNASampler(20):
0.069
Positive Predictive Value
ContextFold:
0.794
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNASampler(20):
0.246
Sensitivity
PETfold_pre2.0(seed):
0.667
RNASampler(20):
0.069
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNASampler(20):
0.246
Sensitivity
CentroidAlifold(seed):
0.414
RNASampler(20):
0.069
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNASampler(20)
Matthews Correlation Coefficient
IPknot:
0.607
RNASampler(20):
0.246
Sensitivity
IPknot:
0.551
RNASampler(20):
0.069
Positive Predictive Value
IPknot:
0.670
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNASampler(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNASampler(20):
0.246
Sensitivity
PETfold_pre2.0(20):
0.410
RNASampler(20):
0.069
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNASampler(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNASampler(20):
0.246
Sensitivity
CentroidFold:
0.515
RNASampler(20):
0.069
Positive Predictive Value
CentroidFold:
0.633
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNASampler(20):
0.246
Sensitivity
CentroidAlifold(20):
0.369
RNASampler(20):
0.069
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNASampler(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNASampler(20):
0.246
Sensitivity
CentroidHomfold‑LAST:
0.359
RNASampler(20):
0.069
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNASampler(20)
Matthews Correlation Coefficient
Contrafold:
0.556
RNASampler(20):
0.246
Sensitivity
Contrafold:
0.542
RNASampler(20):
0.069
Positive Predictive Value
Contrafold:
0.571
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNASampler(20):
0.246
Sensitivity
MXScarna(seed):
0.382
RNASampler(20):
0.069
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNASampler(20):
0.246
Sensitivity
RNAalifold(20):
0.359
RNASampler(20):
0.069
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNASampler(20)
Matthews Correlation Coefficient
Sfold:
0.527
RNASampler(20):
0.246
Sensitivity
Sfold:
0.477
RNASampler(20):
0.069
Positive Predictive Value
Sfold:
0.582
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNASampler(20):
0.246
Sensitivity
RNAalifold(seed):
0.328
RNASampler(20):
0.069
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNASampler(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
RNASampler(20):
0.246
Sensitivity
MaxExpect:
0.505
RNASampler(20):
0.069
Positive Predictive Value
MaxExpect:
0.528
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNASampler(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
RNASampler(20):
0.246
Sensitivity
ProbKnot:
0.508
RNASampler(20):
0.069
Positive Predictive Value
ProbKnot:
0.517
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNASampler(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNASampler(20):
0.246
Sensitivity
MXScarna(20):
0.353
RNASampler(20):
0.069
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNASampler(20)
Matthews Correlation Coefficient
UNAFold:
0.494
RNASampler(20):
0.246
Sensitivity
UNAFold:
0.491
RNASampler(20):
0.069
Positive Predictive Value
UNAFold:
0.498
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNASampler(20)
Matthews Correlation Coefficient
Fold:
0.493
RNASampler(20):
0.246
Sensitivity
Fold:
0.496
RNASampler(20):
0.069
Positive Predictive Value
Fold:
0.490
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RNASampler(20)
Matthews Correlation Coefficient
RNAfold:
0.485
RNASampler(20):
0.246
Sensitivity
RNAfold:
0.486
RNASampler(20):
0.069
Positive Predictive Value
RNAfold:
0.485
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RNASampler(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
RNASampler(20):
0.246
Sensitivity
PknotsRG:
0.440
RNASampler(20):
0.069
Positive Predictive Value
PknotsRG:
0.510
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RNASampler(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNASampler(20):
0.246
Sensitivity
CRWrnafold:
0.379
RNASampler(20):
0.069
Positive Predictive Value
CRWrnafold:
0.490
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RNASampler(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNASampler(20):
0.246
Sensitivity
RNAsubopt:
0.344
RNASampler(20):
0.069
Positive Predictive Value
RNAsubopt:
0.519
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RNASampler(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNASampler(20):
0.246
Sensitivity
TurboFold(20):
0.210
RNASampler(20):
0.069
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RNASampler(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
RNASampler(20):
0.246
Sensitivity
PPfold(20):
0.197
RNASampler(20):
0.069
Positive Predictive Value
PPfold(20):
0.879
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RNASampler(20)
Matthews Correlation Coefficient
Afold:
0.408
RNASampler(20):
0.246
Sensitivity
Afold:
0.343
RNASampler(20):
0.069
Positive Predictive Value
Afold:
0.487
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RNASampler(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
RNASampler(20):
0.246
Sensitivity
Carnac(20):
0.180
RNASampler(20):
0.069
Positive Predictive Value
Carnac(20):
0.892
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RNASampler(20)
Matthews Correlation Coefficient
McQFold:
0.389
RNASampler(20):
0.246
Sensitivity
McQFold:
0.308
RNASampler(20):
0.069
Positive Predictive Value
McQFold:
0.492
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNASampler(20):
0.246
Sensitivity
RSpredict(20):
0.241
RNASampler(20):
0.069
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs RNASampler(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
RNASampler(20):
0.246
Sensitivity
RNAshapes:
0.285
RNASampler(20):
0.069
Positive Predictive Value
RNAshapes:
0.474
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs RNASampler(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
RNASampler(20):
0.246
Sensitivity
RNASLOpt:
0.261
RNASampler(20):
0.069
Positive Predictive Value
RNASLOpt:
0.510
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs RNASampler(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
RNASampler(20):
0.246
Sensitivity
Multilign(20):
0.161
RNASampler(20):
0.069
Positive Predictive Value
Multilign(20):
0.762
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs RNASampler(20)
Matthews Correlation Coefficient
Murlet(20):
0.321
RNASampler(20):
0.246
Sensitivity
Murlet(20):
0.131
RNASampler(20):
0.069
Positive Predictive Value
Murlet(20):
0.787
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs RNASampler(20)
Matthews Correlation Coefficient
RNAwolf:
0.310
RNASampler(20):
0.246
Sensitivity
RNAwolf:
0.261
RNASampler(20):
0.069
Positive Predictive Value
RNAwolf:
0.369
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs RNASampler(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
RNASampler(20):
0.246
Sensitivity
RSpredict(seed):
0.146
RNASampler(20):
0.069
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs RNASampler(20)
Matthews Correlation Coefficient
CMfinder(20):
0.292
RNASampler(20):
0.246
Sensitivity
CMfinder(20):
0.107
RNASampler(20):
0.069
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold4:
0.270
RNASampler(20):
0.246
Sensitivity
Vsfold4:
0.198
RNASampler(20):
0.069
Positive Predictive Value
Vsfold4:
0.371
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs RNASampler(20)
Matthews Correlation Coefficient
Vsfold5:
0.260
RNASampler(20):
0.246
Sensitivity
Vsfold5:
0.195
RNASampler(20):
0.069
Positive Predictive Value
Vsfold5:
0.347
RNASampler(20):
0.877
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
RNASampler(20) vs HotKnots
Matthews Correlation Coefficient
RNASampler(20):
0.246
HotKnots:
0.212
Sensitivity
RNASampler(20):
0.069
HotKnots:
0.069
Positive Predictive Value
RNASampler(20):
0.877
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Mastr(20)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Mastr(20):
0.187
Sensitivity
RNASampler(20):
0.069
Mastr(20):
0.044
Positive Predictive Value
RNASampler(20):
0.877
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Cylofold
Matthews Correlation Coefficient
RNASampler(20):
0.246
Cylofold:
0.178
Sensitivity
RNASampler(20):
0.069
Cylofold:
0.053
Positive Predictive Value
RNASampler(20):
0.877
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Pknots
Matthews Correlation Coefficient
RNASampler(20):
0.246
Pknots:
0.165
Sensitivity
RNASampler(20):
0.069
Pknots:
0.041
Positive Predictive Value
RNASampler(20):
0.877
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Alterna
Matthews Correlation Coefficient
RNASampler(20):
0.246
Alterna:
0.145
Sensitivity
RNASampler(20):
0.069
Alterna:
0.036
Positive Predictive Value
RNASampler(20):
0.877
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs MCFold
Matthews Correlation Coefficient
RNASampler(20):
0.246
MCFold:
0.138
Sensitivity
RNASampler(20):
0.069
MCFold:
0.044
Positive Predictive Value
RNASampler(20):
0.877
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs RDfolder
Matthews Correlation Coefficient
RNASampler(20):
0.246
RDfolder:
0.138
Sensitivity
RNASampler(20):
0.069
RDfolder:
0.031
Positive Predictive Value
RNASampler(20):
0.877
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Murlet(seed):
0.110
Sensitivity
RNASampler(20):
0.069
Murlet(seed):
0.015
Positive Predictive Value
RNASampler(20):
0.877
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
CMfinder(seed):
0.111
Sensitivity
RNASampler(20):
0.069
CMfinder(seed):
0.016
Positive Predictive Value
RNASampler(20):
0.877
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
TurboFold(seed):
0.106
Sensitivity
RNASampler(20):
0.069
TurboFold(seed):
0.014
Positive Predictive Value
RNASampler(20):
0.877
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
PPfold(seed):
0.097
Sensitivity
RNASampler(20):
0.069
PPfold(seed):
0.012
Positive Predictive Value
RNASampler(20):
0.877
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Carnac(seed):
0.093
Sensitivity
RNASampler(20):
0.069
Carnac(seed):
0.010
Positive Predictive Value
RNASampler(20):
0.877
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
RNASampler(seed):
0.088
Sensitivity
RNASampler(20):
0.069
RNASampler(seed):
0.009
Positive Predictive Value
RNASampler(20):
0.877
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Mastr(seed):
0.072
Sensitivity
RNASampler(20):
0.069
Mastr(seed):
0.006
Positive Predictive Value
RNASampler(20):
0.877
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(20):
0.246
NanoFolder:
0.068
Sensitivity
RNASampler(20):
0.069
NanoFolder:
0.027
Positive Predictive Value
RNASampler(20):
0.877
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Multilign(seed):
0.059
Sensitivity
RNASampler(20):
0.069
Multilign(seed):
0.005
Positive Predictive Value
RNASampler(20):
0.877
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| HotKnots |
1242
ContextFold vs HotKnots
Matthews Correlation Coefficient
ContextFold:
0.754
HotKnots:
0.212
Sensitivity
ContextFold:
0.717
HotKnots:
0.069
Positive Predictive Value
ContextFold:
0.794
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
HotKnots:
0.212
Sensitivity
PETfold_pre2.0(seed):
0.667
HotKnots:
0.069
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
HotKnots:
0.212
Sensitivity
CentroidAlifold(seed):
0.414
HotKnots:
0.069
Positive Predictive Value
CentroidAlifold(seed):
0.901
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs HotKnots
Matthews Correlation Coefficient
IPknot:
0.607
HotKnots:
0.212
Sensitivity
IPknot:
0.551
HotKnots:
0.069
Positive Predictive Value
IPknot:
0.670
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs HotKnots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
HotKnots:
0.212
Sensitivity
PETfold_pre2.0(20):
0.410
HotKnots:
0.069
Positive Predictive Value
PETfold_pre2.0(20):
0.807
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs HotKnots
Matthews Correlation Coefficient
CentroidFold:
0.571
HotKnots:
0.212
Sensitivity
CentroidFold:
0.515
HotKnots:
0.069
Positive Predictive Value
CentroidFold:
0.633
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs HotKnots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
HotKnots:
0.212
Sensitivity
CentroidAlifold(20):
0.369
HotKnots:
0.069
Positive Predictive Value
CentroidAlifold(20):
0.891
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs HotKnots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
HotKnots:
0.212
Sensitivity
CentroidHomfold‑LAST:
0.359
HotKnots:
0.069
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs HotKnots
Matthews Correlation Coefficient
Contrafold:
0.556
HotKnots:
0.212
Sensitivity
Contrafold:
0.542
HotKnots:
0.069
Positive Predictive Value
Contrafold:
0.571
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs HotKnots
Matthews Correlation Coefficient
MXScarna(seed):
0.536
HotKnots:
0.212
Sensitivity
MXScarna(seed):
0.382
HotKnots:
0.069
Positive Predictive Value
MXScarna(seed):
0.753
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(20):
0.532
HotKnots:
0.212
Sensitivity
RNAalifold(20):
0.359
HotKnots:
0.069
Positive Predictive Value
RNAalifold(20):
0.787
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs HotKnots
Matthews Correlation Coefficient
Sfold:
0.527
HotKnots:
0.212
Sensitivity
Sfold:
0.477
HotKnots:
0.069
Positive Predictive Value
Sfold:
0.582
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs HotKnots
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
HotKnots:
0.212
Sensitivity
RNAalifold(seed):
0.328
HotKnots:
0.069
Positive Predictive Value
RNAalifold(seed):
0.835
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs HotKnots
Matthews Correlation Coefficient
MaxExpect:
0.516
HotKnots:
0.212
Sensitivity
MaxExpect:
0.505
HotKnots:
0.069
Positive Predictive Value
MaxExpect:
0.528
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs HotKnots
Matthews Correlation Coefficient
ProbKnot:
0.512
HotKnots:
0.212
Sensitivity
ProbKnot:
0.508
HotKnots:
0.069
Positive Predictive Value
ProbKnot:
0.517
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs HotKnots
Matthews Correlation Coefficient
MXScarna(20):
0.500
HotKnots:
0.212
Sensitivity
MXScarna(20):
0.353
HotKnots:
0.069
Positive Predictive Value
MXScarna(20):
0.710
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs HotKnots
Matthews Correlation Coefficient
UNAFold:
0.494
HotKnots:
0.212
Sensitivity
UNAFold:
0.491
HotKnots:
0.069
Positive Predictive Value
UNAFold:
0.498
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs HotKnots
Matthews Correlation Coefficient
Fold:
0.493
HotKnots:
0.212
Sensitivity
Fold:
0.496
HotKnots:
0.069
Positive Predictive Value
Fold:
0.490
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs HotKnots
Matthews Correlation Coefficient
RNAfold:
0.485
HotKnots:
0.212
Sensitivity
RNAfold:
0.486
HotKnots:
0.069
Positive Predictive Value
RNAfold:
0.485
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs HotKnots
Matthews Correlation Coefficient
PknotsRG:
0.474
HotKnots:
0.212
Sensitivity
PknotsRG:
0.440
HotKnots:
0.069
Positive Predictive Value
PknotsRG:
0.510
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs HotKnots
Matthews Correlation Coefficient
CRWrnafold:
0.430
HotKnots:
0.212
Sensitivity
CRWrnafold:
0.379
HotKnots:
0.069
Positive Predictive Value
CRWrnafold:
0.490
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs HotKnots
Matthews Correlation Coefficient
RNAsubopt:
0.422
HotKnots:
0.212
Sensitivity
RNAsubopt:
0.344
HotKnots:
0.069
Positive Predictive Value
RNAsubopt:
0.519
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs HotKnots
Matthews Correlation Coefficient
TurboFold(20):
0.419
HotKnots:
0.212
Sensitivity
TurboFold(20):
0.210
HotKnots:
0.069
Positive Predictive Value
TurboFold(20):
0.837
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs HotKnots
Matthews Correlation Coefficient
PPfold(20):
0.416
HotKnots:
0.212
Sensitivity
PPfold(20):
0.197
HotKnots:
0.069
Positive Predictive Value
PPfold(20):
0.879
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs HotKnots
Matthews Correlation Coefficient
Afold:
0.408
HotKnots:
0.212
Sensitivity
Afold:
0.343
HotKnots:
0.069
Positive Predictive Value
Afold:
0.487
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs HotKnots
Matthews Correlation Coefficient
Carnac(20):
0.401
HotKnots:
0.212
Sensitivity
Carnac(20):
0.180
HotKnots:
0.069
Positive Predictive Value
Carnac(20):
0.892
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs HotKnots
Matthews Correlation Coefficient
McQFold:
0.389
HotKnots:
0.212
Sensitivity
McQFold:
0.308
HotKnots:
0.069
Positive Predictive Value
McQFold:
0.492
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs HotKnots
Matthews Correlation Coefficient
RSpredict(20):
0.377
HotKnots:
0.212
Sensitivity
RSpredict(20):
0.241
HotKnots:
0.069
Positive Predictive Value
RSpredict(20):
0.590
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs HotKnots
Matthews Correlation Coefficient
RNAshapes:
0.367
HotKnots:
0.212
Sensitivity
RNAshapes:
0.285
HotKnots:
0.069
Positive Predictive Value
RNAshapes:
0.474
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs HotKnots
Matthews Correlation Coefficient
RNASLOpt:
0.364
HotKnots:
0.212
Sensitivity
RNASLOpt:
0.261
HotKnots:
0.069
Positive Predictive Value
RNASLOpt:
0.510
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs HotKnots
Matthews Correlation Coefficient
Multilign(20):
0.350
HotKnots:
0.212
Sensitivity
Multilign(20):
0.161
HotKnots:
0.069
Positive Predictive Value
Multilign(20):
0.762
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs HotKnots
Matthews Correlation Coefficient
Murlet(20):
0.321
HotKnots:
0.212
Sensitivity
Murlet(20):
0.131
HotKnots:
0.069
Positive Predictive Value
Murlet(20):
0.787
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs HotKnots
Matthews Correlation Coefficient
RNAwolf:
0.310
HotKnots:
0.212
Sensitivity
RNAwolf:
0.261
HotKnots:
0.069
Positive Predictive Value
RNAwolf:
0.369
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs HotKnots
Matthews Correlation Coefficient
RSpredict(seed):
0.295
HotKnots:
0.212
Sensitivity
RSpredict(seed):
0.146
HotKnots:
0.069
Positive Predictive Value
RSpredict(seed):
0.597
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs HotKnots
Matthews Correlation Coefficient
CMfinder(20):
0.292
HotKnots:
0.212
Sensitivity
CMfinder(20):
0.107
HotKnots:
0.069
Positive Predictive Value
CMfinder(20):
0.794
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs HotKnots
Matthews Correlation Coefficient
Vsfold4:
0.270
HotKnots:
0.212
Sensitivity
Vsfold4:
0.198
HotKnots:
0.069
Positive Predictive Value
Vsfold4:
0.371
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs HotKnots
Matthews Correlation Coefficient
Vsfold5:
0.260
HotKnots:
0.212
Sensitivity
Vsfold5:
0.195
HotKnots:
0.069
Positive Predictive Value
Vsfold5:
0.347
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs HotKnots
Matthews Correlation Coefficient
RNASampler(20):
0.246
HotKnots:
0.212
Sensitivity
RNASampler(20):
0.069
HotKnots:
0.069
Positive Predictive Value
RNASampler(20):
0.877
HotKnots:
0.659
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
HotKnots vs Mastr(20)
Matthews Correlation Coefficient
HotKnots:
0.212
Mastr(20):
0.187
Sensitivity
HotKnots:
0.069
Mastr(20):
0.044
Positive Predictive Value
HotKnots:
0.659
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Cylofold
Matthews Correlation Coefficient
HotKnots:
0.212
Cylofold:
0.178
Sensitivity
HotKnots:
0.069
Cylofold:
0.053
Positive Predictive Value
HotKnots:
0.659
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Pknots
Matthews Correlation Coefficient
HotKnots:
0.212
Pknots:
0.165
Sensitivity
HotKnots:
0.069
Pknots:
0.041
Positive Predictive Value
HotKnots:
0.659
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Alterna
Matthews Correlation Coefficient
HotKnots:
0.212
Alterna:
0.145
Sensitivity
HotKnots:
0.069
Alterna:
0.036
Positive Predictive Value
HotKnots:
0.659
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs MCFold
Matthews Correlation Coefficient
HotKnots:
0.212
MCFold:
0.138
Sensitivity
HotKnots:
0.069
MCFold:
0.044
Positive Predictive Value
HotKnots:
0.659
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs RDfolder
Matthews Correlation Coefficient
HotKnots:
0.212
RDfolder:
0.138
Sensitivity
HotKnots:
0.069
RDfolder:
0.031
Positive Predictive Value
HotKnots:
0.659
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Murlet(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Murlet(seed):
0.110
Sensitivity
HotKnots:
0.069
Murlet(seed):
0.015
Positive Predictive Value
HotKnots:
0.659
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs CMfinder(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
CMfinder(seed):
0.111
Sensitivity
HotKnots:
0.069
CMfinder(seed):
0.016
Positive Predictive Value
HotKnots:
0.659
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs TurboFold(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
TurboFold(seed):
0.106
Sensitivity
HotKnots:
0.069
TurboFold(seed):
0.014
Positive Predictive Value
HotKnots:
0.659
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs PPfold(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
PPfold(seed):
0.097
Sensitivity
HotKnots:
0.069
PPfold(seed):
0.012
Positive Predictive Value
HotKnots:
0.659
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Carnac(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Carnac(seed):
0.093
Sensitivity
HotKnots:
0.069
Carnac(seed):
0.010
Positive Predictive Value
HotKnots:
0.659
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs RNASampler(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
RNASampler(seed):
0.088
Sensitivity
HotKnots:
0.069
RNASampler(seed):
0.009
Positive Predictive Value
HotKnots:
0.659
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Mastr(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Mastr(seed):
0.072
Sensitivity
HotKnots:
0.069
Mastr(seed):
0.006
Positive Predictive Value
HotKnots:
0.659
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs NanoFolder
Matthews Correlation Coefficient
HotKnots:
0.212
NanoFolder:
0.068
Sensitivity
HotKnots:
0.069
NanoFolder:
0.027
Positive Predictive Value
HotKnots:
0.659
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
HotKnots vs Multilign(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Multilign(seed):
0.059
Sensitivity
HotKnots:
0.069
Multilign(seed):
0.005
Positive Predictive Value
HotKnots:
0.659
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Mastr(20) |
1242
ContextFold vs Mastr(20)
Matthews Correlation Coefficient
ContextFold:
0.754
Mastr(20):
0.187
Sensitivity
ContextFold:
0.717
Mastr(20):
0.044
Positive Predictive Value
ContextFold:
0.794
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Mastr(20):
0.187
Sensitivity
PETfold_pre2.0(seed):
0.667
Mastr(20):
0.044
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Mastr(20):
0.187
Sensitivity
CentroidAlifold(seed):
0.414
Mastr(20):
0.044
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Mastr(20)
Matthews Correlation Coefficient
IPknot:
0.607
Mastr(20):
0.187
Sensitivity
IPknot:
0.551
Mastr(20):
0.044
Positive Predictive Value
IPknot:
0.670
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Mastr(20)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Mastr(20):
0.187
Sensitivity
PETfold_pre2.0(20):
0.410
Mastr(20):
0.044
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Mastr(20)
Matthews Correlation Coefficient
CentroidFold:
0.571
Mastr(20):
0.187
Sensitivity
CentroidFold:
0.515
Mastr(20):
0.044
Positive Predictive Value
CentroidFold:
0.633
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Mastr(20)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Mastr(20):
0.187
Sensitivity
CentroidAlifold(20):
0.369
Mastr(20):
0.044
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Mastr(20)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Mastr(20):
0.187
Sensitivity
CentroidHomfold‑LAST:
0.359
Mastr(20):
0.044
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Mastr(20)
Matthews Correlation Coefficient
Contrafold:
0.556
Mastr(20):
0.187
Sensitivity
Contrafold:
0.542
Mastr(20):
0.044
Positive Predictive Value
Contrafold:
0.571
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Mastr(20):
0.187
Sensitivity
MXScarna(seed):
0.382
Mastr(20):
0.044
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Mastr(20):
0.187
Sensitivity
RNAalifold(20):
0.359
Mastr(20):
0.044
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Mastr(20)
Matthews Correlation Coefficient
Sfold:
0.527
Mastr(20):
0.187
Sensitivity
Sfold:
0.477
Mastr(20):
0.044
Positive Predictive Value
Sfold:
0.582
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Mastr(20)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Mastr(20):
0.187
Sensitivity
RNAalifold(seed):
0.328
Mastr(20):
0.044
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Mastr(20)
Matthews Correlation Coefficient
MaxExpect:
0.516
Mastr(20):
0.187
Sensitivity
MaxExpect:
0.505
Mastr(20):
0.044
Positive Predictive Value
MaxExpect:
0.528
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Mastr(20)
Matthews Correlation Coefficient
ProbKnot:
0.512
Mastr(20):
0.187
Sensitivity
ProbKnot:
0.508
Mastr(20):
0.044
Positive Predictive Value
ProbKnot:
0.517
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Mastr(20)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Mastr(20):
0.187
Sensitivity
MXScarna(20):
0.353
Mastr(20):
0.044
Positive Predictive Value
MXScarna(20):
0.710
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Mastr(20)
Matthews Correlation Coefficient
UNAFold:
0.494
Mastr(20):
0.187
Sensitivity
UNAFold:
0.491
Mastr(20):
0.044
Positive Predictive Value
UNAFold:
0.498
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Mastr(20)
Matthews Correlation Coefficient
Fold:
0.493
Mastr(20):
0.187
Sensitivity
Fold:
0.496
Mastr(20):
0.044
Positive Predictive Value
Fold:
0.490
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Mastr(20)
Matthews Correlation Coefficient
RNAfold:
0.485
Mastr(20):
0.187
Sensitivity
RNAfold:
0.486
Mastr(20):
0.044
Positive Predictive Value
RNAfold:
0.485
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Mastr(20)
Matthews Correlation Coefficient
PknotsRG:
0.474
Mastr(20):
0.187
Sensitivity
PknotsRG:
0.440
Mastr(20):
0.044
Positive Predictive Value
PknotsRG:
0.510
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Mastr(20)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Mastr(20):
0.187
Sensitivity
CRWrnafold:
0.379
Mastr(20):
0.044
Positive Predictive Value
CRWrnafold:
0.490
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Mastr(20)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Mastr(20):
0.187
Sensitivity
RNAsubopt:
0.344
Mastr(20):
0.044
Positive Predictive Value
RNAsubopt:
0.519
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Mastr(20)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Mastr(20):
0.187
Sensitivity
TurboFold(20):
0.210
Mastr(20):
0.044
Positive Predictive Value
TurboFold(20):
0.837
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Mastr(20)
Matthews Correlation Coefficient
PPfold(20):
0.416
Mastr(20):
0.187
Sensitivity
PPfold(20):
0.197
Mastr(20):
0.044
Positive Predictive Value
PPfold(20):
0.879
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Mastr(20)
Matthews Correlation Coefficient
Afold:
0.408
Mastr(20):
0.187
Sensitivity
Afold:
0.343
Mastr(20):
0.044
Positive Predictive Value
Afold:
0.487
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Mastr(20)
Matthews Correlation Coefficient
Carnac(20):
0.401
Mastr(20):
0.187
Sensitivity
Carnac(20):
0.180
Mastr(20):
0.044
Positive Predictive Value
Carnac(20):
0.892
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Mastr(20)
Matthews Correlation Coefficient
McQFold:
0.389
Mastr(20):
0.187
Sensitivity
McQFold:
0.308
Mastr(20):
0.044
Positive Predictive Value
McQFold:
0.492
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Mastr(20):
0.187
Sensitivity
RSpredict(20):
0.241
Mastr(20):
0.044
Positive Predictive Value
RSpredict(20):
0.590
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Mastr(20)
Matthews Correlation Coefficient
RNAshapes:
0.367
Mastr(20):
0.187
Sensitivity
RNAshapes:
0.285
Mastr(20):
0.044
Positive Predictive Value
RNAshapes:
0.474
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Mastr(20)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Mastr(20):
0.187
Sensitivity
RNASLOpt:
0.261
Mastr(20):
0.044
Positive Predictive Value
RNASLOpt:
0.510
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Mastr(20)
Matthews Correlation Coefficient
Multilign(20):
0.350
Mastr(20):
0.187
Sensitivity
Multilign(20):
0.161
Mastr(20):
0.044
Positive Predictive Value
Multilign(20):
0.762
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Mastr(20)
Matthews Correlation Coefficient
Murlet(20):
0.321
Mastr(20):
0.187
Sensitivity
Murlet(20):
0.131
Mastr(20):
0.044
Positive Predictive Value
Murlet(20):
0.787
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Mastr(20)
Matthews Correlation Coefficient
RNAwolf:
0.310
Mastr(20):
0.187
Sensitivity
RNAwolf:
0.261
Mastr(20):
0.044
Positive Predictive Value
RNAwolf:
0.369
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Mastr(20)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Mastr(20):
0.187
Sensitivity
RSpredict(seed):
0.146
Mastr(20):
0.044
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Mastr(20)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Mastr(20):
0.187
Sensitivity
CMfinder(20):
0.107
Mastr(20):
0.044
Positive Predictive Value
CMfinder(20):
0.794
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold4:
0.270
Mastr(20):
0.187
Sensitivity
Vsfold4:
0.198
Mastr(20):
0.044
Positive Predictive Value
Vsfold4:
0.371
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Mastr(20)
Matthews Correlation Coefficient
Vsfold5:
0.260
Mastr(20):
0.187
Sensitivity
Vsfold5:
0.195
Mastr(20):
0.044
Positive Predictive Value
Vsfold5:
0.347
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Mastr(20)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Mastr(20):
0.187
Sensitivity
RNASampler(20):
0.069
Mastr(20):
0.044
Positive Predictive Value
RNASampler(20):
0.877
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Mastr(20)
Matthews Correlation Coefficient
HotKnots:
0.212
Mastr(20):
0.187
Sensitivity
HotKnots:
0.069
Mastr(20):
0.044
Positive Predictive Value
HotKnots:
0.659
Mastr(20):
0.790
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Mastr(20) vs Cylofold
Matthews Correlation Coefficient
Mastr(20):
0.187
Cylofold:
0.178
Sensitivity
Mastr(20):
0.044
Cylofold:
0.053
Positive Predictive Value
Mastr(20):
0.790
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Pknots
Matthews Correlation Coefficient
Mastr(20):
0.187
Pknots:
0.165
Sensitivity
Mastr(20):
0.044
Pknots:
0.041
Positive Predictive Value
Mastr(20):
0.790
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Alterna
Matthews Correlation Coefficient
Mastr(20):
0.187
Alterna:
0.145
Sensitivity
Mastr(20):
0.044
Alterna:
0.036
Positive Predictive Value
Mastr(20):
0.790
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs MCFold
Matthews Correlation Coefficient
Mastr(20):
0.187
MCFold:
0.138
Sensitivity
Mastr(20):
0.044
MCFold:
0.044
Positive Predictive Value
Mastr(20):
0.790
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs RDfolder
Matthews Correlation Coefficient
Mastr(20):
0.187
RDfolder:
0.138
Sensitivity
Mastr(20):
0.044
RDfolder:
0.031
Positive Predictive Value
Mastr(20):
0.790
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Murlet(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Murlet(seed):
0.110
Sensitivity
Mastr(20):
0.044
Murlet(seed):
0.015
Positive Predictive Value
Mastr(20):
0.790
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
CMfinder(seed):
0.111
Sensitivity
Mastr(20):
0.044
CMfinder(seed):
0.016
Positive Predictive Value
Mastr(20):
0.790
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
TurboFold(seed):
0.106
Sensitivity
Mastr(20):
0.044
TurboFold(seed):
0.014
Positive Predictive Value
Mastr(20):
0.790
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs PPfold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
PPfold(seed):
0.097
Sensitivity
Mastr(20):
0.044
PPfold(seed):
0.012
Positive Predictive Value
Mastr(20):
0.790
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Carnac(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Carnac(seed):
0.093
Sensitivity
Mastr(20):
0.044
Carnac(seed):
0.010
Positive Predictive Value
Mastr(20):
0.790
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
RNASampler(seed):
0.088
Sensitivity
Mastr(20):
0.044
RNASampler(seed):
0.009
Positive Predictive Value
Mastr(20):
0.790
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Mastr(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Mastr(seed):
0.072
Sensitivity
Mastr(20):
0.044
Mastr(seed):
0.006
Positive Predictive Value
Mastr(20):
0.790
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs NanoFolder
Matthews Correlation Coefficient
Mastr(20):
0.187
NanoFolder:
0.068
Sensitivity
Mastr(20):
0.044
NanoFolder:
0.027
Positive Predictive Value
Mastr(20):
0.790
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Mastr(20) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Multilign(seed):
0.059
Sensitivity
Mastr(20):
0.044
Multilign(seed):
0.005
Positive Predictive Value
Mastr(20):
0.790
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Cylofold |
1242
ContextFold vs Cylofold
Matthews Correlation Coefficient
ContextFold:
0.754
Cylofold:
0.178
Sensitivity
ContextFold:
0.717
Cylofold:
0.053
Positive Predictive Value
ContextFold:
0.794
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Cylofold:
0.178
Sensitivity
PETfold_pre2.0(seed):
0.667
Cylofold:
0.053
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Cylofold:
0.178
Sensitivity
CentroidAlifold(seed):
0.414
Cylofold:
0.053
Positive Predictive Value
CentroidAlifold(seed):
0.901
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Cylofold
Matthews Correlation Coefficient
IPknot:
0.607
Cylofold:
0.178
Sensitivity
IPknot:
0.551
Cylofold:
0.053
Positive Predictive Value
IPknot:
0.670
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Cylofold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Cylofold:
0.178
Sensitivity
PETfold_pre2.0(20):
0.410
Cylofold:
0.053
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Cylofold
Matthews Correlation Coefficient
CentroidFold:
0.571
Cylofold:
0.178
Sensitivity
CentroidFold:
0.515
Cylofold:
0.053
Positive Predictive Value
CentroidFold:
0.633
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Cylofold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Cylofold:
0.178
Sensitivity
CentroidAlifold(20):
0.369
Cylofold:
0.053
Positive Predictive Value
CentroidAlifold(20):
0.891
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Cylofold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Cylofold:
0.178
Sensitivity
CentroidHomfold‑LAST:
0.359
Cylofold:
0.053
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Cylofold
Matthews Correlation Coefficient
Contrafold:
0.556
Cylofold:
0.178
Sensitivity
Contrafold:
0.542
Cylofold:
0.053
Positive Predictive Value
Contrafold:
0.571
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Cylofold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Cylofold:
0.178
Sensitivity
MXScarna(seed):
0.382
Cylofold:
0.053
Positive Predictive Value
MXScarna(seed):
0.753
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Cylofold:
0.178
Sensitivity
RNAalifold(20):
0.359
Cylofold:
0.053
Positive Predictive Value
RNAalifold(20):
0.787
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Cylofold
Matthews Correlation Coefficient
Sfold:
0.527
Cylofold:
0.178
Sensitivity
Sfold:
0.477
Cylofold:
0.053
Positive Predictive Value
Sfold:
0.582
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Cylofold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Cylofold:
0.178
Sensitivity
RNAalifold(seed):
0.328
Cylofold:
0.053
Positive Predictive Value
RNAalifold(seed):
0.835
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Cylofold
Matthews Correlation Coefficient
MaxExpect:
0.516
Cylofold:
0.178
Sensitivity
MaxExpect:
0.505
Cylofold:
0.053
Positive Predictive Value
MaxExpect:
0.528
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Cylofold
Matthews Correlation Coefficient
ProbKnot:
0.512
Cylofold:
0.178
Sensitivity
ProbKnot:
0.508
Cylofold:
0.053
Positive Predictive Value
ProbKnot:
0.517
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Cylofold
Matthews Correlation Coefficient
MXScarna(20):
0.500
Cylofold:
0.178
Sensitivity
MXScarna(20):
0.353
Cylofold:
0.053
Positive Predictive Value
MXScarna(20):
0.710
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Cylofold
Matthews Correlation Coefficient
UNAFold:
0.494
Cylofold:
0.178
Sensitivity
UNAFold:
0.491
Cylofold:
0.053
Positive Predictive Value
UNAFold:
0.498
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Cylofold
Matthews Correlation Coefficient
Fold:
0.493
Cylofold:
0.178
Sensitivity
Fold:
0.496
Cylofold:
0.053
Positive Predictive Value
Fold:
0.490
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Cylofold
Matthews Correlation Coefficient
RNAfold:
0.485
Cylofold:
0.178
Sensitivity
RNAfold:
0.486
Cylofold:
0.053
Positive Predictive Value
RNAfold:
0.485
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Cylofold
Matthews Correlation Coefficient
PknotsRG:
0.474
Cylofold:
0.178
Sensitivity
PknotsRG:
0.440
Cylofold:
0.053
Positive Predictive Value
PknotsRG:
0.510
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Cylofold
Matthews Correlation Coefficient
CRWrnafold:
0.430
Cylofold:
0.178
Sensitivity
CRWrnafold:
0.379
Cylofold:
0.053
Positive Predictive Value
CRWrnafold:
0.490
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Cylofold
Matthews Correlation Coefficient
RNAsubopt:
0.422
Cylofold:
0.178
Sensitivity
RNAsubopt:
0.344
Cylofold:
0.053
Positive Predictive Value
RNAsubopt:
0.519
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Cylofold
Matthews Correlation Coefficient
TurboFold(20):
0.419
Cylofold:
0.178
Sensitivity
TurboFold(20):
0.210
Cylofold:
0.053
Positive Predictive Value
TurboFold(20):
0.837
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Cylofold
Matthews Correlation Coefficient
PPfold(20):
0.416
Cylofold:
0.178
Sensitivity
PPfold(20):
0.197
Cylofold:
0.053
Positive Predictive Value
PPfold(20):
0.879
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Cylofold
Matthews Correlation Coefficient
Afold:
0.408
Cylofold:
0.178
Sensitivity
Afold:
0.343
Cylofold:
0.053
Positive Predictive Value
Afold:
0.487
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Cylofold
Matthews Correlation Coefficient
Carnac(20):
0.401
Cylofold:
0.178
Sensitivity
Carnac(20):
0.180
Cylofold:
0.053
Positive Predictive Value
Carnac(20):
0.892
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Cylofold
Matthews Correlation Coefficient
McQFold:
0.389
Cylofold:
0.178
Sensitivity
McQFold:
0.308
Cylofold:
0.053
Positive Predictive Value
McQFold:
0.492
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Cylofold
Matthews Correlation Coefficient
RSpredict(20):
0.377
Cylofold:
0.178
Sensitivity
RSpredict(20):
0.241
Cylofold:
0.053
Positive Predictive Value
RSpredict(20):
0.590
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Cylofold
Matthews Correlation Coefficient
RNAshapes:
0.367
Cylofold:
0.178
Sensitivity
RNAshapes:
0.285
Cylofold:
0.053
Positive Predictive Value
RNAshapes:
0.474
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Cylofold
Matthews Correlation Coefficient
RNASLOpt:
0.364
Cylofold:
0.178
Sensitivity
RNASLOpt:
0.261
Cylofold:
0.053
Positive Predictive Value
RNASLOpt:
0.510
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Cylofold
Matthews Correlation Coefficient
Multilign(20):
0.350
Cylofold:
0.178
Sensitivity
Multilign(20):
0.161
Cylofold:
0.053
Positive Predictive Value
Multilign(20):
0.762
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Cylofold
Matthews Correlation Coefficient
Murlet(20):
0.321
Cylofold:
0.178
Sensitivity
Murlet(20):
0.131
Cylofold:
0.053
Positive Predictive Value
Murlet(20):
0.787
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Cylofold
Matthews Correlation Coefficient
RNAwolf:
0.310
Cylofold:
0.178
Sensitivity
RNAwolf:
0.261
Cylofold:
0.053
Positive Predictive Value
RNAwolf:
0.369
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Cylofold
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Cylofold:
0.178
Sensitivity
RSpredict(seed):
0.146
Cylofold:
0.053
Positive Predictive Value
RSpredict(seed):
0.597
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Cylofold
Matthews Correlation Coefficient
CMfinder(20):
0.292
Cylofold:
0.178
Sensitivity
CMfinder(20):
0.107
Cylofold:
0.053
Positive Predictive Value
CMfinder(20):
0.794
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Cylofold
Matthews Correlation Coefficient
Vsfold4:
0.270
Cylofold:
0.178
Sensitivity
Vsfold4:
0.198
Cylofold:
0.053
Positive Predictive Value
Vsfold4:
0.371
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Cylofold
Matthews Correlation Coefficient
Vsfold5:
0.260
Cylofold:
0.178
Sensitivity
Vsfold5:
0.195
Cylofold:
0.053
Positive Predictive Value
Vsfold5:
0.347
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Cylofold
Matthews Correlation Coefficient
RNASampler(20):
0.246
Cylofold:
0.178
Sensitivity
RNASampler(20):
0.069
Cylofold:
0.053
Positive Predictive Value
RNASampler(20):
0.877
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Cylofold
Matthews Correlation Coefficient
HotKnots:
0.212
Cylofold:
0.178
Sensitivity
HotKnots:
0.069
Cylofold:
0.053
Positive Predictive Value
HotKnots:
0.659
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Cylofold
Matthews Correlation Coefficient
Mastr(20):
0.187
Cylofold:
0.178
Sensitivity
Mastr(20):
0.044
Cylofold:
0.053
Positive Predictive Value
Mastr(20):
0.790
Cylofold:
0.597
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Cylofold vs Pknots
Matthews Correlation Coefficient
Cylofold:
0.178
Pknots:
0.165
Sensitivity
Cylofold:
0.053
Pknots:
0.041
Positive Predictive Value
Cylofold:
0.597
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Alterna
Matthews Correlation Coefficient
Cylofold:
0.178
Alterna:
0.145
Sensitivity
Cylofold:
0.053
Alterna:
0.036
Positive Predictive Value
Cylofold:
0.597
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs MCFold
Matthews Correlation Coefficient
Cylofold:
0.178
MCFold:
0.138
Sensitivity
Cylofold:
0.053
MCFold:
0.044
Positive Predictive Value
Cylofold:
0.597
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs RDfolder
Matthews Correlation Coefficient
Cylofold:
0.178
RDfolder:
0.138
Sensitivity
Cylofold:
0.053
RDfolder:
0.031
Positive Predictive Value
Cylofold:
0.597
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Murlet(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Murlet(seed):
0.110
Sensitivity
Cylofold:
0.053
Murlet(seed):
0.015
Positive Predictive Value
Cylofold:
0.597
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs CMfinder(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
CMfinder(seed):
0.111
Sensitivity
Cylofold:
0.053
CMfinder(seed):
0.016
Positive Predictive Value
Cylofold:
0.597
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs TurboFold(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
TurboFold(seed):
0.106
Sensitivity
Cylofold:
0.053
TurboFold(seed):
0.014
Positive Predictive Value
Cylofold:
0.597
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs PPfold(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
PPfold(seed):
0.097
Sensitivity
Cylofold:
0.053
PPfold(seed):
0.012
Positive Predictive Value
Cylofold:
0.597
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Carnac(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Carnac(seed):
0.093
Sensitivity
Cylofold:
0.053
Carnac(seed):
0.010
Positive Predictive Value
Cylofold:
0.597
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs RNASampler(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
RNASampler(seed):
0.088
Sensitivity
Cylofold:
0.053
RNASampler(seed):
0.009
Positive Predictive Value
Cylofold:
0.597
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Mastr(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Mastr(seed):
0.072
Sensitivity
Cylofold:
0.053
Mastr(seed):
0.006
Positive Predictive Value
Cylofold:
0.597
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs NanoFolder
Matthews Correlation Coefficient
Cylofold:
0.178
NanoFolder:
0.068
Sensitivity
Cylofold:
0.053
NanoFolder:
0.027
Positive Predictive Value
Cylofold:
0.597
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Cylofold vs Multilign(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Multilign(seed):
0.059
Sensitivity
Cylofold:
0.053
Multilign(seed):
0.005
Positive Predictive Value
Cylofold:
0.597
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Pknots |
1242
ContextFold vs Pknots
Matthews Correlation Coefficient
ContextFold:
0.754
Pknots:
0.165
Sensitivity
ContextFold:
0.717
Pknots:
0.041
Positive Predictive Value
ContextFold:
0.794
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Pknots:
0.165
Sensitivity
PETfold_pre2.0(seed):
0.667
Pknots:
0.041
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Pknots:
0.165
Sensitivity
CentroidAlifold(seed):
0.414
Pknots:
0.041
Positive Predictive Value
CentroidAlifold(seed):
0.901
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Pknots
Matthews Correlation Coefficient
IPknot:
0.607
Pknots:
0.165
Sensitivity
IPknot:
0.551
Pknots:
0.041
Positive Predictive Value
IPknot:
0.670
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Pknots
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Pknots:
0.165
Sensitivity
PETfold_pre2.0(20):
0.410
Pknots:
0.041
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Pknots
Matthews Correlation Coefficient
CentroidFold:
0.571
Pknots:
0.165
Sensitivity
CentroidFold:
0.515
Pknots:
0.041
Positive Predictive Value
CentroidFold:
0.633
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Pknots
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Pknots:
0.165
Sensitivity
CentroidAlifold(20):
0.369
Pknots:
0.041
Positive Predictive Value
CentroidAlifold(20):
0.891
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Pknots
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Pknots:
0.165
Sensitivity
CentroidHomfold‑LAST:
0.359
Pknots:
0.041
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Pknots
Matthews Correlation Coefficient
Contrafold:
0.556
Pknots:
0.165
Sensitivity
Contrafold:
0.542
Pknots:
0.041
Positive Predictive Value
Contrafold:
0.571
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Pknots
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Pknots:
0.165
Sensitivity
MXScarna(seed):
0.382
Pknots:
0.041
Positive Predictive Value
MXScarna(seed):
0.753
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Pknots
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Pknots:
0.165
Sensitivity
RNAalifold(20):
0.359
Pknots:
0.041
Positive Predictive Value
RNAalifold(20):
0.787
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Pknots
Matthews Correlation Coefficient
Sfold:
0.527
Pknots:
0.165
Sensitivity
Sfold:
0.477
Pknots:
0.041
Positive Predictive Value
Sfold:
0.582
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Pknots
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Pknots:
0.165
Sensitivity
RNAalifold(seed):
0.328
Pknots:
0.041
Positive Predictive Value
RNAalifold(seed):
0.835
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Pknots
Matthews Correlation Coefficient
MaxExpect:
0.516
Pknots:
0.165
Sensitivity
MaxExpect:
0.505
Pknots:
0.041
Positive Predictive Value
MaxExpect:
0.528
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Pknots
Matthews Correlation Coefficient
ProbKnot:
0.512
Pknots:
0.165
Sensitivity
ProbKnot:
0.508
Pknots:
0.041
Positive Predictive Value
ProbKnot:
0.517
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Pknots
Matthews Correlation Coefficient
MXScarna(20):
0.500
Pknots:
0.165
Sensitivity
MXScarna(20):
0.353
Pknots:
0.041
Positive Predictive Value
MXScarna(20):
0.710
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Pknots
Matthews Correlation Coefficient
UNAFold:
0.494
Pknots:
0.165
Sensitivity
UNAFold:
0.491
Pknots:
0.041
Positive Predictive Value
UNAFold:
0.498
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Pknots
Matthews Correlation Coefficient
Fold:
0.493
Pknots:
0.165
Sensitivity
Fold:
0.496
Pknots:
0.041
Positive Predictive Value
Fold:
0.490
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Pknots
Matthews Correlation Coefficient
RNAfold:
0.485
Pknots:
0.165
Sensitivity
RNAfold:
0.486
Pknots:
0.041
Positive Predictive Value
RNAfold:
0.485
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Pknots
Matthews Correlation Coefficient
PknotsRG:
0.474
Pknots:
0.165
Sensitivity
PknotsRG:
0.440
Pknots:
0.041
Positive Predictive Value
PknotsRG:
0.510
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Pknots
Matthews Correlation Coefficient
CRWrnafold:
0.430
Pknots:
0.165
Sensitivity
CRWrnafold:
0.379
Pknots:
0.041
Positive Predictive Value
CRWrnafold:
0.490
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Pknots
Matthews Correlation Coefficient
RNAsubopt:
0.422
Pknots:
0.165
Sensitivity
RNAsubopt:
0.344
Pknots:
0.041
Positive Predictive Value
RNAsubopt:
0.519
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Pknots
Matthews Correlation Coefficient
TurboFold(20):
0.419
Pknots:
0.165
Sensitivity
TurboFold(20):
0.210
Pknots:
0.041
Positive Predictive Value
TurboFold(20):
0.837
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Pknots
Matthews Correlation Coefficient
PPfold(20):
0.416
Pknots:
0.165
Sensitivity
PPfold(20):
0.197
Pknots:
0.041
Positive Predictive Value
PPfold(20):
0.879
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Pknots
Matthews Correlation Coefficient
Afold:
0.408
Pknots:
0.165
Sensitivity
Afold:
0.343
Pknots:
0.041
Positive Predictive Value
Afold:
0.487
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Pknots
Matthews Correlation Coefficient
Carnac(20):
0.401
Pknots:
0.165
Sensitivity
Carnac(20):
0.180
Pknots:
0.041
Positive Predictive Value
Carnac(20):
0.892
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Pknots
Matthews Correlation Coefficient
McQFold:
0.389
Pknots:
0.165
Sensitivity
McQFold:
0.308
Pknots:
0.041
Positive Predictive Value
McQFold:
0.492
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Pknots
Matthews Correlation Coefficient
RSpredict(20):
0.377
Pknots:
0.165
Sensitivity
RSpredict(20):
0.241
Pknots:
0.041
Positive Predictive Value
RSpredict(20):
0.590
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Pknots
Matthews Correlation Coefficient
RNAshapes:
0.367
Pknots:
0.165
Sensitivity
RNAshapes:
0.285
Pknots:
0.041
Positive Predictive Value
RNAshapes:
0.474
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Pknots
Matthews Correlation Coefficient
RNASLOpt:
0.364
Pknots:
0.165
Sensitivity
RNASLOpt:
0.261
Pknots:
0.041
Positive Predictive Value
RNASLOpt:
0.510
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Pknots
Matthews Correlation Coefficient
Multilign(20):
0.350
Pknots:
0.165
Sensitivity
Multilign(20):
0.161
Pknots:
0.041
Positive Predictive Value
Multilign(20):
0.762
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Pknots
Matthews Correlation Coefficient
Murlet(20):
0.321
Pknots:
0.165
Sensitivity
Murlet(20):
0.131
Pknots:
0.041
Positive Predictive Value
Murlet(20):
0.787
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Pknots
Matthews Correlation Coefficient
RNAwolf:
0.310
Pknots:
0.165
Sensitivity
RNAwolf:
0.261
Pknots:
0.041
Positive Predictive Value
RNAwolf:
0.369
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Pknots
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Pknots:
0.165
Sensitivity
RSpredict(seed):
0.146
Pknots:
0.041
Positive Predictive Value
RSpredict(seed):
0.597
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Pknots
Matthews Correlation Coefficient
CMfinder(20):
0.292
Pknots:
0.165
Sensitivity
CMfinder(20):
0.107
Pknots:
0.041
Positive Predictive Value
CMfinder(20):
0.794
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Pknots
Matthews Correlation Coefficient
Vsfold4:
0.270
Pknots:
0.165
Sensitivity
Vsfold4:
0.198
Pknots:
0.041
Positive Predictive Value
Vsfold4:
0.371
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Pknots
Matthews Correlation Coefficient
Vsfold5:
0.260
Pknots:
0.165
Sensitivity
Vsfold5:
0.195
Pknots:
0.041
Positive Predictive Value
Vsfold5:
0.347
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Pknots
Matthews Correlation Coefficient
RNASampler(20):
0.246
Pknots:
0.165
Sensitivity
RNASampler(20):
0.069
Pknots:
0.041
Positive Predictive Value
RNASampler(20):
0.877
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Pknots
Matthews Correlation Coefficient
HotKnots:
0.212
Pknots:
0.165
Sensitivity
HotKnots:
0.069
Pknots:
0.041
Positive Predictive Value
HotKnots:
0.659
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Pknots
Matthews Correlation Coefficient
Mastr(20):
0.187
Pknots:
0.165
Sensitivity
Mastr(20):
0.044
Pknots:
0.041
Positive Predictive Value
Mastr(20):
0.790
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs Pknots
Matthews Correlation Coefficient
Cylofold:
0.178
Pknots:
0.165
Sensitivity
Cylofold:
0.053
Pknots:
0.041
Positive Predictive Value
Cylofold:
0.597
Pknots:
0.662
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Pknots vs Alterna
Matthews Correlation Coefficient
Pknots:
0.165
Alterna:
0.145
Sensitivity
Pknots:
0.041
Alterna:
0.036
Positive Predictive Value
Pknots:
0.662
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs MCFold
Matthews Correlation Coefficient
Pknots:
0.165
MCFold:
0.138
Sensitivity
Pknots:
0.041
MCFold:
0.044
Positive Predictive Value
Pknots:
0.662
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs RDfolder
Matthews Correlation Coefficient
Pknots:
0.165
RDfolder:
0.138
Sensitivity
Pknots:
0.041
RDfolder:
0.031
Positive Predictive Value
Pknots:
0.662
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Murlet(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Murlet(seed):
0.110
Sensitivity
Pknots:
0.041
Murlet(seed):
0.015
Positive Predictive Value
Pknots:
0.662
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs CMfinder(seed)
Matthews Correlation Coefficient
Pknots:
0.165
CMfinder(seed):
0.111
Sensitivity
Pknots:
0.041
CMfinder(seed):
0.016
Positive Predictive Value
Pknots:
0.662
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs TurboFold(seed)
Matthews Correlation Coefficient
Pknots:
0.165
TurboFold(seed):
0.106
Sensitivity
Pknots:
0.041
TurboFold(seed):
0.014
Positive Predictive Value
Pknots:
0.662
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs PPfold(seed)
Matthews Correlation Coefficient
Pknots:
0.165
PPfold(seed):
0.097
Sensitivity
Pknots:
0.041
PPfold(seed):
0.012
Positive Predictive Value
Pknots:
0.662
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Carnac(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Carnac(seed):
0.093
Sensitivity
Pknots:
0.041
Carnac(seed):
0.010
Positive Predictive Value
Pknots:
0.662
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs RNASampler(seed)
Matthews Correlation Coefficient
Pknots:
0.165
RNASampler(seed):
0.088
Sensitivity
Pknots:
0.041
RNASampler(seed):
0.009
Positive Predictive Value
Pknots:
0.662
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Mastr(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Mastr(seed):
0.072
Sensitivity
Pknots:
0.041
Mastr(seed):
0.006
Positive Predictive Value
Pknots:
0.662
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs NanoFolder
Matthews Correlation Coefficient
Pknots:
0.165
NanoFolder:
0.068
Sensitivity
Pknots:
0.041
NanoFolder:
0.027
Positive Predictive Value
Pknots:
0.662
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Pknots vs Multilign(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Multilign(seed):
0.059
Sensitivity
Pknots:
0.041
Multilign(seed):
0.005
Positive Predictive Value
Pknots:
0.662
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Alterna |
1242
ContextFold vs Alterna
Matthews Correlation Coefficient
ContextFold:
0.754
Alterna:
0.145
Sensitivity
ContextFold:
0.717
Alterna:
0.036
Positive Predictive Value
ContextFold:
0.794
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Alterna:
0.145
Sensitivity
PETfold_pre2.0(seed):
0.667
Alterna:
0.036
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Alterna:
0.145
Sensitivity
CentroidAlifold(seed):
0.414
Alterna:
0.036
Positive Predictive Value
CentroidAlifold(seed):
0.901
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Alterna
Matthews Correlation Coefficient
IPknot:
0.607
Alterna:
0.145
Sensitivity
IPknot:
0.551
Alterna:
0.036
Positive Predictive Value
IPknot:
0.670
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Alterna
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Alterna:
0.145
Sensitivity
PETfold_pre2.0(20):
0.410
Alterna:
0.036
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Alterna
Matthews Correlation Coefficient
CentroidFold:
0.571
Alterna:
0.145
Sensitivity
CentroidFold:
0.515
Alterna:
0.036
Positive Predictive Value
CentroidFold:
0.633
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Alterna
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Alterna:
0.145
Sensitivity
CentroidAlifold(20):
0.369
Alterna:
0.036
Positive Predictive Value
CentroidAlifold(20):
0.891
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Alterna
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Alterna:
0.145
Sensitivity
CentroidHomfold‑LAST:
0.359
Alterna:
0.036
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Alterna
Matthews Correlation Coefficient
Contrafold:
0.556
Alterna:
0.145
Sensitivity
Contrafold:
0.542
Alterna:
0.036
Positive Predictive Value
Contrafold:
0.571
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Alterna
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Alterna:
0.145
Sensitivity
MXScarna(seed):
0.382
Alterna:
0.036
Positive Predictive Value
MXScarna(seed):
0.753
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Alterna
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Alterna:
0.145
Sensitivity
RNAalifold(20):
0.359
Alterna:
0.036
Positive Predictive Value
RNAalifold(20):
0.787
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Alterna
Matthews Correlation Coefficient
Sfold:
0.527
Alterna:
0.145
Sensitivity
Sfold:
0.477
Alterna:
0.036
Positive Predictive Value
Sfold:
0.582
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Alterna
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Alterna:
0.145
Sensitivity
RNAalifold(seed):
0.328
Alterna:
0.036
Positive Predictive Value
RNAalifold(seed):
0.835
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Alterna
Matthews Correlation Coefficient
MaxExpect:
0.516
Alterna:
0.145
Sensitivity
MaxExpect:
0.505
Alterna:
0.036
Positive Predictive Value
MaxExpect:
0.528
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Alterna
Matthews Correlation Coefficient
ProbKnot:
0.512
Alterna:
0.145
Sensitivity
ProbKnot:
0.508
Alterna:
0.036
Positive Predictive Value
ProbKnot:
0.517
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Alterna
Matthews Correlation Coefficient
MXScarna(20):
0.500
Alterna:
0.145
Sensitivity
MXScarna(20):
0.353
Alterna:
0.036
Positive Predictive Value
MXScarna(20):
0.710
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Alterna
Matthews Correlation Coefficient
UNAFold:
0.494
Alterna:
0.145
Sensitivity
UNAFold:
0.491
Alterna:
0.036
Positive Predictive Value
UNAFold:
0.498
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Alterna
Matthews Correlation Coefficient
Fold:
0.493
Alterna:
0.145
Sensitivity
Fold:
0.496
Alterna:
0.036
Positive Predictive Value
Fold:
0.490
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Alterna
Matthews Correlation Coefficient
RNAfold:
0.485
Alterna:
0.145
Sensitivity
RNAfold:
0.486
Alterna:
0.036
Positive Predictive Value
RNAfold:
0.485
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Alterna
Matthews Correlation Coefficient
PknotsRG:
0.474
Alterna:
0.145
Sensitivity
PknotsRG:
0.440
Alterna:
0.036
Positive Predictive Value
PknotsRG:
0.510
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Alterna
Matthews Correlation Coefficient
CRWrnafold:
0.430
Alterna:
0.145
Sensitivity
CRWrnafold:
0.379
Alterna:
0.036
Positive Predictive Value
CRWrnafold:
0.490
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Alterna
Matthews Correlation Coefficient
RNAsubopt:
0.422
Alterna:
0.145
Sensitivity
RNAsubopt:
0.344
Alterna:
0.036
Positive Predictive Value
RNAsubopt:
0.519
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Alterna
Matthews Correlation Coefficient
TurboFold(20):
0.419
Alterna:
0.145
Sensitivity
TurboFold(20):
0.210
Alterna:
0.036
Positive Predictive Value
TurboFold(20):
0.837
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Alterna
Matthews Correlation Coefficient
PPfold(20):
0.416
Alterna:
0.145
Sensitivity
PPfold(20):
0.197
Alterna:
0.036
Positive Predictive Value
PPfold(20):
0.879
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Alterna
Matthews Correlation Coefficient
Afold:
0.408
Alterna:
0.145
Sensitivity
Afold:
0.343
Alterna:
0.036
Positive Predictive Value
Afold:
0.487
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Alterna
Matthews Correlation Coefficient
Carnac(20):
0.401
Alterna:
0.145
Sensitivity
Carnac(20):
0.180
Alterna:
0.036
Positive Predictive Value
Carnac(20):
0.892
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Alterna
Matthews Correlation Coefficient
McQFold:
0.389
Alterna:
0.145
Sensitivity
McQFold:
0.308
Alterna:
0.036
Positive Predictive Value
McQFold:
0.492
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Alterna
Matthews Correlation Coefficient
RSpredict(20):
0.377
Alterna:
0.145
Sensitivity
RSpredict(20):
0.241
Alterna:
0.036
Positive Predictive Value
RSpredict(20):
0.590
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Alterna
Matthews Correlation Coefficient
RNAshapes:
0.367
Alterna:
0.145
Sensitivity
RNAshapes:
0.285
Alterna:
0.036
Positive Predictive Value
RNAshapes:
0.474
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Alterna
Matthews Correlation Coefficient
RNASLOpt:
0.364
Alterna:
0.145
Sensitivity
RNASLOpt:
0.261
Alterna:
0.036
Positive Predictive Value
RNASLOpt:
0.510
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Alterna
Matthews Correlation Coefficient
Multilign(20):
0.350
Alterna:
0.145
Sensitivity
Multilign(20):
0.161
Alterna:
0.036
Positive Predictive Value
Multilign(20):
0.762
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Alterna
Matthews Correlation Coefficient
Murlet(20):
0.321
Alterna:
0.145
Sensitivity
Murlet(20):
0.131
Alterna:
0.036
Positive Predictive Value
Murlet(20):
0.787
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Alterna
Matthews Correlation Coefficient
RNAwolf:
0.310
Alterna:
0.145
Sensitivity
RNAwolf:
0.261
Alterna:
0.036
Positive Predictive Value
RNAwolf:
0.369
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Alterna
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Alterna:
0.145
Sensitivity
RSpredict(seed):
0.146
Alterna:
0.036
Positive Predictive Value
RSpredict(seed):
0.597
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Alterna
Matthews Correlation Coefficient
CMfinder(20):
0.292
Alterna:
0.145
Sensitivity
CMfinder(20):
0.107
Alterna:
0.036
Positive Predictive Value
CMfinder(20):
0.794
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Alterna
Matthews Correlation Coefficient
Vsfold4:
0.270
Alterna:
0.145
Sensitivity
Vsfold4:
0.198
Alterna:
0.036
Positive Predictive Value
Vsfold4:
0.371
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Alterna
Matthews Correlation Coefficient
Vsfold5:
0.260
Alterna:
0.145
Sensitivity
Vsfold5:
0.195
Alterna:
0.036
Positive Predictive Value
Vsfold5:
0.347
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Alterna
Matthews Correlation Coefficient
RNASampler(20):
0.246
Alterna:
0.145
Sensitivity
RNASampler(20):
0.069
Alterna:
0.036
Positive Predictive Value
RNASampler(20):
0.877
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Alterna
Matthews Correlation Coefficient
HotKnots:
0.212
Alterna:
0.145
Sensitivity
HotKnots:
0.069
Alterna:
0.036
Positive Predictive Value
HotKnots:
0.659
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Alterna
Matthews Correlation Coefficient
Mastr(20):
0.187
Alterna:
0.145
Sensitivity
Mastr(20):
0.044
Alterna:
0.036
Positive Predictive Value
Mastr(20):
0.790
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs Alterna
Matthews Correlation Coefficient
Cylofold:
0.178
Alterna:
0.145
Sensitivity
Cylofold:
0.053
Alterna:
0.036
Positive Predictive Value
Cylofold:
0.597
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs Alterna
Matthews Correlation Coefficient
Pknots:
0.165
Alterna:
0.145
Sensitivity
Pknots:
0.041
Alterna:
0.036
Positive Predictive Value
Pknots:
0.662
Alterna:
0.582
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Alterna vs MCFold
Matthews Correlation Coefficient
Alterna:
0.145
MCFold:
0.138
Sensitivity
Alterna:
0.036
MCFold:
0.044
Positive Predictive Value
Alterna:
0.582
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs RDfolder
Matthews Correlation Coefficient
Alterna:
0.145
RDfolder:
0.138
Sensitivity
Alterna:
0.036
RDfolder:
0.031
Positive Predictive Value
Alterna:
0.582
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Murlet(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Murlet(seed):
0.110
Sensitivity
Alterna:
0.036
Murlet(seed):
0.015
Positive Predictive Value
Alterna:
0.582
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs CMfinder(seed)
Matthews Correlation Coefficient
Alterna:
0.145
CMfinder(seed):
0.111
Sensitivity
Alterna:
0.036
CMfinder(seed):
0.016
Positive Predictive Value
Alterna:
0.582
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs TurboFold(seed)
Matthews Correlation Coefficient
Alterna:
0.145
TurboFold(seed):
0.106
Sensitivity
Alterna:
0.036
TurboFold(seed):
0.014
Positive Predictive Value
Alterna:
0.582
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs PPfold(seed)
Matthews Correlation Coefficient
Alterna:
0.145
PPfold(seed):
0.097
Sensitivity
Alterna:
0.036
PPfold(seed):
0.012
Positive Predictive Value
Alterna:
0.582
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Carnac(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Carnac(seed):
0.093
Sensitivity
Alterna:
0.036
Carnac(seed):
0.010
Positive Predictive Value
Alterna:
0.582
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs RNASampler(seed)
Matthews Correlation Coefficient
Alterna:
0.145
RNASampler(seed):
0.088
Sensitivity
Alterna:
0.036
RNASampler(seed):
0.009
Positive Predictive Value
Alterna:
0.582
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Mastr(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Mastr(seed):
0.072
Sensitivity
Alterna:
0.036
Mastr(seed):
0.006
Positive Predictive Value
Alterna:
0.582
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs NanoFolder
Matthews Correlation Coefficient
Alterna:
0.145
NanoFolder:
0.068
Sensitivity
Alterna:
0.036
NanoFolder:
0.027
Positive Predictive Value
Alterna:
0.582
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Alterna vs Multilign(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Multilign(seed):
0.059
Sensitivity
Alterna:
0.036
Multilign(seed):
0.005
Positive Predictive Value
Alterna:
0.582
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| MCFold |
1242
ContextFold vs MCFold
Matthews Correlation Coefficient
ContextFold:
0.754
MCFold:
0.138
Sensitivity
ContextFold:
0.717
MCFold:
0.044
Positive Predictive Value
ContextFold:
0.794
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
MCFold:
0.138
Sensitivity
PETfold_pre2.0(seed):
0.667
MCFold:
0.044
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
MCFold:
0.138
Sensitivity
CentroidAlifold(seed):
0.414
MCFold:
0.044
Positive Predictive Value
CentroidAlifold(seed):
0.901
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs MCFold
Matthews Correlation Coefficient
IPknot:
0.607
MCFold:
0.138
Sensitivity
IPknot:
0.551
MCFold:
0.044
Positive Predictive Value
IPknot:
0.670
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs MCFold
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
MCFold:
0.138
Sensitivity
PETfold_pre2.0(20):
0.410
MCFold:
0.044
Positive Predictive Value
PETfold_pre2.0(20):
0.807
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs MCFold
Matthews Correlation Coefficient
CentroidFold:
0.571
MCFold:
0.138
Sensitivity
CentroidFold:
0.515
MCFold:
0.044
Positive Predictive Value
CentroidFold:
0.633
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs MCFold
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
MCFold:
0.138
Sensitivity
CentroidAlifold(20):
0.369
MCFold:
0.044
Positive Predictive Value
CentroidAlifold(20):
0.891
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs MCFold
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
MCFold:
0.138
Sensitivity
CentroidHomfold‑LAST:
0.359
MCFold:
0.044
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs MCFold
Matthews Correlation Coefficient
Contrafold:
0.556
MCFold:
0.138
Sensitivity
Contrafold:
0.542
MCFold:
0.044
Positive Predictive Value
Contrafold:
0.571
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs MCFold
Matthews Correlation Coefficient
MXScarna(seed):
0.536
MCFold:
0.138
Sensitivity
MXScarna(seed):
0.382
MCFold:
0.044
Positive Predictive Value
MXScarna(seed):
0.753
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs MCFold
Matthews Correlation Coefficient
RNAalifold(20):
0.532
MCFold:
0.138
Sensitivity
RNAalifold(20):
0.359
MCFold:
0.044
Positive Predictive Value
RNAalifold(20):
0.787
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs MCFold
Matthews Correlation Coefficient
Sfold:
0.527
MCFold:
0.138
Sensitivity
Sfold:
0.477
MCFold:
0.044
Positive Predictive Value
Sfold:
0.582
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs MCFold
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
MCFold:
0.138
Sensitivity
RNAalifold(seed):
0.328
MCFold:
0.044
Positive Predictive Value
RNAalifold(seed):
0.835
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs MCFold
Matthews Correlation Coefficient
MaxExpect:
0.516
MCFold:
0.138
Sensitivity
MaxExpect:
0.505
MCFold:
0.044
Positive Predictive Value
MaxExpect:
0.528
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs MCFold
Matthews Correlation Coefficient
ProbKnot:
0.512
MCFold:
0.138
Sensitivity
ProbKnot:
0.508
MCFold:
0.044
Positive Predictive Value
ProbKnot:
0.517
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs MCFold
Matthews Correlation Coefficient
MXScarna(20):
0.500
MCFold:
0.138
Sensitivity
MXScarna(20):
0.353
MCFold:
0.044
Positive Predictive Value
MXScarna(20):
0.710
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs MCFold
Matthews Correlation Coefficient
UNAFold:
0.494
MCFold:
0.138
Sensitivity
UNAFold:
0.491
MCFold:
0.044
Positive Predictive Value
UNAFold:
0.498
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs MCFold
Matthews Correlation Coefficient
Fold:
0.493
MCFold:
0.138
Sensitivity
Fold:
0.496
MCFold:
0.044
Positive Predictive Value
Fold:
0.490
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs MCFold
Matthews Correlation Coefficient
RNAfold:
0.485
MCFold:
0.138
Sensitivity
RNAfold:
0.486
MCFold:
0.044
Positive Predictive Value
RNAfold:
0.485
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs MCFold
Matthews Correlation Coefficient
PknotsRG:
0.474
MCFold:
0.138
Sensitivity
PknotsRG:
0.440
MCFold:
0.044
Positive Predictive Value
PknotsRG:
0.510
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs MCFold
Matthews Correlation Coefficient
CRWrnafold:
0.430
MCFold:
0.138
Sensitivity
CRWrnafold:
0.379
MCFold:
0.044
Positive Predictive Value
CRWrnafold:
0.490
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs MCFold
Matthews Correlation Coefficient
RNAsubopt:
0.422
MCFold:
0.138
Sensitivity
RNAsubopt:
0.344
MCFold:
0.044
Positive Predictive Value
RNAsubopt:
0.519
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs MCFold
Matthews Correlation Coefficient
TurboFold(20):
0.419
MCFold:
0.138
Sensitivity
TurboFold(20):
0.210
MCFold:
0.044
Positive Predictive Value
TurboFold(20):
0.837
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs MCFold
Matthews Correlation Coefficient
PPfold(20):
0.416
MCFold:
0.138
Sensitivity
PPfold(20):
0.197
MCFold:
0.044
Positive Predictive Value
PPfold(20):
0.879
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs MCFold
Matthews Correlation Coefficient
Afold:
0.408
MCFold:
0.138
Sensitivity
Afold:
0.343
MCFold:
0.044
Positive Predictive Value
Afold:
0.487
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs MCFold
Matthews Correlation Coefficient
Carnac(20):
0.401
MCFold:
0.138
Sensitivity
Carnac(20):
0.180
MCFold:
0.044
Positive Predictive Value
Carnac(20):
0.892
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs MCFold
Matthews Correlation Coefficient
McQFold:
0.389
MCFold:
0.138
Sensitivity
McQFold:
0.308
MCFold:
0.044
Positive Predictive Value
McQFold:
0.492
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs MCFold
Matthews Correlation Coefficient
RSpredict(20):
0.377
MCFold:
0.138
Sensitivity
RSpredict(20):
0.241
MCFold:
0.044
Positive Predictive Value
RSpredict(20):
0.590
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs MCFold
Matthews Correlation Coefficient
RNAshapes:
0.367
MCFold:
0.138
Sensitivity
RNAshapes:
0.285
MCFold:
0.044
Positive Predictive Value
RNAshapes:
0.474
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs MCFold
Matthews Correlation Coefficient
RNASLOpt:
0.364
MCFold:
0.138
Sensitivity
RNASLOpt:
0.261
MCFold:
0.044
Positive Predictive Value
RNASLOpt:
0.510
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs MCFold
Matthews Correlation Coefficient
Multilign(20):
0.350
MCFold:
0.138
Sensitivity
Multilign(20):
0.161
MCFold:
0.044
Positive Predictive Value
Multilign(20):
0.762
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs MCFold
Matthews Correlation Coefficient
Murlet(20):
0.321
MCFold:
0.138
Sensitivity
Murlet(20):
0.131
MCFold:
0.044
Positive Predictive Value
Murlet(20):
0.787
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs MCFold
Matthews Correlation Coefficient
RNAwolf:
0.310
MCFold:
0.138
Sensitivity
RNAwolf:
0.261
MCFold:
0.044
Positive Predictive Value
RNAwolf:
0.369
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs MCFold
Matthews Correlation Coefficient
RSpredict(seed):
0.295
MCFold:
0.138
Sensitivity
RSpredict(seed):
0.146
MCFold:
0.044
Positive Predictive Value
RSpredict(seed):
0.597
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs MCFold
Matthews Correlation Coefficient
CMfinder(20):
0.292
MCFold:
0.138
Sensitivity
CMfinder(20):
0.107
MCFold:
0.044
Positive Predictive Value
CMfinder(20):
0.794
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs MCFold
Matthews Correlation Coefficient
Vsfold4:
0.270
MCFold:
0.138
Sensitivity
Vsfold4:
0.198
MCFold:
0.044
Positive Predictive Value
Vsfold4:
0.371
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs MCFold
Matthews Correlation Coefficient
Vsfold5:
0.260
MCFold:
0.138
Sensitivity
Vsfold5:
0.195
MCFold:
0.044
Positive Predictive Value
Vsfold5:
0.347
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs MCFold
Matthews Correlation Coefficient
RNASampler(20):
0.246
MCFold:
0.138
Sensitivity
RNASampler(20):
0.069
MCFold:
0.044
Positive Predictive Value
RNASampler(20):
0.877
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs MCFold
Matthews Correlation Coefficient
HotKnots:
0.212
MCFold:
0.138
Sensitivity
HotKnots:
0.069
MCFold:
0.044
Positive Predictive Value
HotKnots:
0.659
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs MCFold
Matthews Correlation Coefficient
Mastr(20):
0.187
MCFold:
0.138
Sensitivity
Mastr(20):
0.044
MCFold:
0.044
Positive Predictive Value
Mastr(20):
0.790
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs MCFold
Matthews Correlation Coefficient
Cylofold:
0.178
MCFold:
0.138
Sensitivity
Cylofold:
0.053
MCFold:
0.044
Positive Predictive Value
Cylofold:
0.597
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs MCFold
Matthews Correlation Coefficient
Pknots:
0.165
MCFold:
0.138
Sensitivity
Pknots:
0.041
MCFold:
0.044
Positive Predictive Value
Pknots:
0.662
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs MCFold
Matthews Correlation Coefficient
Alterna:
0.145
MCFold:
0.138
Sensitivity
Alterna:
0.036
MCFold:
0.044
Positive Predictive Value
Alterna:
0.582
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
RDfolder vs MCFold
Matthews Correlation Coefficient
RDfolder:
0.138
MCFold:
0.138
Sensitivity
RDfolder:
0.031
MCFold:
0.044
Positive Predictive Value
RDfolder:
0.615
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.0186613811547
|
+
MCFold vs Murlet(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Murlet(seed):
0.110
Sensitivity
MCFold:
0.044
Murlet(seed):
0.015
Positive Predictive Value
MCFold:
0.432
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs CMfinder(seed)
Matthews Correlation Coefficient
MCFold:
0.138
CMfinder(seed):
0.111
Sensitivity
MCFold:
0.044
CMfinder(seed):
0.016
Positive Predictive Value
MCFold:
0.432
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs TurboFold(seed)
Matthews Correlation Coefficient
MCFold:
0.138
TurboFold(seed):
0.106
Sensitivity
MCFold:
0.044
TurboFold(seed):
0.014
Positive Predictive Value
MCFold:
0.432
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs PPfold(seed)
Matthews Correlation Coefficient
MCFold:
0.138
PPfold(seed):
0.097
Sensitivity
MCFold:
0.044
PPfold(seed):
0.012
Positive Predictive Value
MCFold:
0.432
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Carnac(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Carnac(seed):
0.093
Sensitivity
MCFold:
0.044
Carnac(seed):
0.010
Positive Predictive Value
MCFold:
0.432
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs RNASampler(seed)
Matthews Correlation Coefficient
MCFold:
0.138
RNASampler(seed):
0.088
Sensitivity
MCFold:
0.044
RNASampler(seed):
0.009
Positive Predictive Value
MCFold:
0.432
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Mastr(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Mastr(seed):
0.072
Sensitivity
MCFold:
0.044
Mastr(seed):
0.006
Positive Predictive Value
MCFold:
0.432
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs NanoFolder
Matthews Correlation Coefficient
MCFold:
0.138
NanoFolder:
0.068
Sensitivity
MCFold:
0.044
NanoFolder:
0.027
Positive Predictive Value
MCFold:
0.432
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
MCFold vs Multilign(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Multilign(seed):
0.059
Sensitivity
MCFold:
0.044
Multilign(seed):
0.005
Positive Predictive Value
MCFold:
0.432
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RDfolder |
1242
ContextFold vs RDfolder
Matthews Correlation Coefficient
ContextFold:
0.754
RDfolder:
0.138
Sensitivity
ContextFold:
0.717
RDfolder:
0.031
Positive Predictive Value
ContextFold:
0.794
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RDfolder:
0.138
Sensitivity
PETfold_pre2.0(seed):
0.667
RDfolder:
0.031
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RDfolder:
0.138
Sensitivity
CentroidAlifold(seed):
0.414
RDfolder:
0.031
Positive Predictive Value
CentroidAlifold(seed):
0.901
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RDfolder
Matthews Correlation Coefficient
IPknot:
0.607
RDfolder:
0.138
Sensitivity
IPknot:
0.551
RDfolder:
0.031
Positive Predictive Value
IPknot:
0.670
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RDfolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RDfolder:
0.138
Sensitivity
PETfold_pre2.0(20):
0.410
RDfolder:
0.031
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RDfolder
Matthews Correlation Coefficient
CentroidFold:
0.571
RDfolder:
0.138
Sensitivity
CentroidFold:
0.515
RDfolder:
0.031
Positive Predictive Value
CentroidFold:
0.633
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RDfolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RDfolder:
0.138
Sensitivity
CentroidAlifold(20):
0.369
RDfolder:
0.031
Positive Predictive Value
CentroidAlifold(20):
0.891
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RDfolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RDfolder:
0.138
Sensitivity
CentroidHomfold‑LAST:
0.359
RDfolder:
0.031
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RDfolder
Matthews Correlation Coefficient
Contrafold:
0.556
RDfolder:
0.138
Sensitivity
Contrafold:
0.542
RDfolder:
0.031
Positive Predictive Value
Contrafold:
0.571
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RDfolder
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RDfolder:
0.138
Sensitivity
MXScarna(seed):
0.382
RDfolder:
0.031
Positive Predictive Value
MXScarna(seed):
0.753
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RDfolder:
0.138
Sensitivity
RNAalifold(20):
0.359
RDfolder:
0.031
Positive Predictive Value
RNAalifold(20):
0.787
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RDfolder
Matthews Correlation Coefficient
Sfold:
0.527
RDfolder:
0.138
Sensitivity
Sfold:
0.477
RDfolder:
0.031
Positive Predictive Value
Sfold:
0.582
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RDfolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RDfolder:
0.138
Sensitivity
RNAalifold(seed):
0.328
RDfolder:
0.031
Positive Predictive Value
RNAalifold(seed):
0.835
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RDfolder
Matthews Correlation Coefficient
MaxExpect:
0.516
RDfolder:
0.138
Sensitivity
MaxExpect:
0.505
RDfolder:
0.031
Positive Predictive Value
MaxExpect:
0.528
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RDfolder
Matthews Correlation Coefficient
ProbKnot:
0.512
RDfolder:
0.138
Sensitivity
ProbKnot:
0.508
RDfolder:
0.031
Positive Predictive Value
ProbKnot:
0.517
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RDfolder
Matthews Correlation Coefficient
MXScarna(20):
0.500
RDfolder:
0.138
Sensitivity
MXScarna(20):
0.353
RDfolder:
0.031
Positive Predictive Value
MXScarna(20):
0.710
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RDfolder
Matthews Correlation Coefficient
UNAFold:
0.494
RDfolder:
0.138
Sensitivity
UNAFold:
0.491
RDfolder:
0.031
Positive Predictive Value
UNAFold:
0.498
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RDfolder
Matthews Correlation Coefficient
Fold:
0.493
RDfolder:
0.138
Sensitivity
Fold:
0.496
RDfolder:
0.031
Positive Predictive Value
Fold:
0.490
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RDfolder
Matthews Correlation Coefficient
RNAfold:
0.485
RDfolder:
0.138
Sensitivity
RNAfold:
0.486
RDfolder:
0.031
Positive Predictive Value
RNAfold:
0.485
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RDfolder
Matthews Correlation Coefficient
PknotsRG:
0.474
RDfolder:
0.138
Sensitivity
PknotsRG:
0.440
RDfolder:
0.031
Positive Predictive Value
PknotsRG:
0.510
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RDfolder
Matthews Correlation Coefficient
CRWrnafold:
0.430
RDfolder:
0.138
Sensitivity
CRWrnafold:
0.379
RDfolder:
0.031
Positive Predictive Value
CRWrnafold:
0.490
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RDfolder
Matthews Correlation Coefficient
RNAsubopt:
0.422
RDfolder:
0.138
Sensitivity
RNAsubopt:
0.344
RDfolder:
0.031
Positive Predictive Value
RNAsubopt:
0.519
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RDfolder
Matthews Correlation Coefficient
TurboFold(20):
0.419
RDfolder:
0.138
Sensitivity
TurboFold(20):
0.210
RDfolder:
0.031
Positive Predictive Value
TurboFold(20):
0.837
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RDfolder
Matthews Correlation Coefficient
PPfold(20):
0.416
RDfolder:
0.138
Sensitivity
PPfold(20):
0.197
RDfolder:
0.031
Positive Predictive Value
PPfold(20):
0.879
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RDfolder
Matthews Correlation Coefficient
Afold:
0.408
RDfolder:
0.138
Sensitivity
Afold:
0.343
RDfolder:
0.031
Positive Predictive Value
Afold:
0.487
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RDfolder
Matthews Correlation Coefficient
Carnac(20):
0.401
RDfolder:
0.138
Sensitivity
Carnac(20):
0.180
RDfolder:
0.031
Positive Predictive Value
Carnac(20):
0.892
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RDfolder
Matthews Correlation Coefficient
McQFold:
0.389
RDfolder:
0.138
Sensitivity
McQFold:
0.308
RDfolder:
0.031
Positive Predictive Value
McQFold:
0.492
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RDfolder
Matthews Correlation Coefficient
RSpredict(20):
0.377
RDfolder:
0.138
Sensitivity
RSpredict(20):
0.241
RDfolder:
0.031
Positive Predictive Value
RSpredict(20):
0.590
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs RDfolder
Matthews Correlation Coefficient
RNAshapes:
0.367
RDfolder:
0.138
Sensitivity
RNAshapes:
0.285
RDfolder:
0.031
Positive Predictive Value
RNAshapes:
0.474
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs RDfolder
Matthews Correlation Coefficient
RNASLOpt:
0.364
RDfolder:
0.138
Sensitivity
RNASLOpt:
0.261
RDfolder:
0.031
Positive Predictive Value
RNASLOpt:
0.510
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs RDfolder
Matthews Correlation Coefficient
Multilign(20):
0.350
RDfolder:
0.138
Sensitivity
Multilign(20):
0.161
RDfolder:
0.031
Positive Predictive Value
Multilign(20):
0.762
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs RDfolder
Matthews Correlation Coefficient
Murlet(20):
0.321
RDfolder:
0.138
Sensitivity
Murlet(20):
0.131
RDfolder:
0.031
Positive Predictive Value
Murlet(20):
0.787
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs RDfolder
Matthews Correlation Coefficient
RNAwolf:
0.310
RDfolder:
0.138
Sensitivity
RNAwolf:
0.261
RDfolder:
0.031
Positive Predictive Value
RNAwolf:
0.369
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs RDfolder
Matthews Correlation Coefficient
RSpredict(seed):
0.295
RDfolder:
0.138
Sensitivity
RSpredict(seed):
0.146
RDfolder:
0.031
Positive Predictive Value
RSpredict(seed):
0.597
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs RDfolder
Matthews Correlation Coefficient
CMfinder(20):
0.292
RDfolder:
0.138
Sensitivity
CMfinder(20):
0.107
RDfolder:
0.031
Positive Predictive Value
CMfinder(20):
0.794
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs RDfolder
Matthews Correlation Coefficient
Vsfold4:
0.270
RDfolder:
0.138
Sensitivity
Vsfold4:
0.198
RDfolder:
0.031
Positive Predictive Value
Vsfold4:
0.371
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs RDfolder
Matthews Correlation Coefficient
Vsfold5:
0.260
RDfolder:
0.138
Sensitivity
Vsfold5:
0.195
RDfolder:
0.031
Positive Predictive Value
Vsfold5:
0.347
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs RDfolder
Matthews Correlation Coefficient
RNASampler(20):
0.246
RDfolder:
0.138
Sensitivity
RNASampler(20):
0.069
RDfolder:
0.031
Positive Predictive Value
RNASampler(20):
0.877
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs RDfolder
Matthews Correlation Coefficient
HotKnots:
0.212
RDfolder:
0.138
Sensitivity
HotKnots:
0.069
RDfolder:
0.031
Positive Predictive Value
HotKnots:
0.659
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs RDfolder
Matthews Correlation Coefficient
Mastr(20):
0.187
RDfolder:
0.138
Sensitivity
Mastr(20):
0.044
RDfolder:
0.031
Positive Predictive Value
Mastr(20):
0.790
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs RDfolder
Matthews Correlation Coefficient
Cylofold:
0.178
RDfolder:
0.138
Sensitivity
Cylofold:
0.053
RDfolder:
0.031
Positive Predictive Value
Cylofold:
0.597
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs RDfolder
Matthews Correlation Coefficient
Pknots:
0.165
RDfolder:
0.138
Sensitivity
Pknots:
0.041
RDfolder:
0.031
Positive Predictive Value
Pknots:
0.662
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs RDfolder
Matthews Correlation Coefficient
Alterna:
0.145
RDfolder:
0.138
Sensitivity
Alterna:
0.036
RDfolder:
0.031
Positive Predictive Value
Alterna:
0.582
RDfolder:
0.615
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs MCFold
Matthews Correlation Coefficient
RDfolder:
0.138
MCFold:
0.138
Sensitivity
RDfolder:
0.031
MCFold:
0.044
Positive Predictive Value
RDfolder:
0.615
MCFold:
0.432
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.0186613811547
|
|
+
RDfolder vs Murlet(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Murlet(seed):
0.110
Sensitivity
RDfolder:
0.031
Murlet(seed):
0.015
Positive Predictive Value
RDfolder:
0.615
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs CMfinder(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
CMfinder(seed):
0.111
Sensitivity
RDfolder:
0.031
CMfinder(seed):
0.016
Positive Predictive Value
RDfolder:
0.615
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs TurboFold(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
TurboFold(seed):
0.106
Sensitivity
RDfolder:
0.031
TurboFold(seed):
0.014
Positive Predictive Value
RDfolder:
0.615
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs PPfold(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
PPfold(seed):
0.097
Sensitivity
RDfolder:
0.031
PPfold(seed):
0.012
Positive Predictive Value
RDfolder:
0.615
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs Carnac(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Carnac(seed):
0.093
Sensitivity
RDfolder:
0.031
Carnac(seed):
0.010
Positive Predictive Value
RDfolder:
0.615
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs RNASampler(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
RNASampler(seed):
0.088
Sensitivity
RDfolder:
0.031
RNASampler(seed):
0.009
Positive Predictive Value
RDfolder:
0.615
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs Mastr(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Mastr(seed):
0.072
Sensitivity
RDfolder:
0.031
Mastr(seed):
0.006
Positive Predictive Value
RDfolder:
0.615
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs NanoFolder
Matthews Correlation Coefficient
RDfolder:
0.138
NanoFolder:
0.068
Sensitivity
RDfolder:
0.031
NanoFolder:
0.027
Positive Predictive Value
RDfolder:
0.615
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RDfolder vs Multilign(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Multilign(seed):
0.059
Sensitivity
RDfolder:
0.031
Multilign(seed):
0.005
Positive Predictive Value
RDfolder:
0.615
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Murlet(seed) |
1242
ContextFold vs Murlet(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Murlet(seed):
0.110
Sensitivity
ContextFold:
0.717
Murlet(seed):
0.015
Positive Predictive Value
ContextFold:
0.794
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Murlet(seed):
0.110
Sensitivity
PETfold_pre2.0(seed):
0.667
Murlet(seed):
0.015
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Murlet(seed):
0.110
Sensitivity
CentroidAlifold(seed):
0.414
Murlet(seed):
0.015
Positive Predictive Value
CentroidAlifold(seed):
0.901
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Murlet(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Murlet(seed):
0.110
Sensitivity
IPknot:
0.551
Murlet(seed):
0.015
Positive Predictive Value
IPknot:
0.670
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Murlet(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Murlet(seed):
0.110
Sensitivity
PETfold_pre2.0(20):
0.410
Murlet(seed):
0.015
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Murlet(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Murlet(seed):
0.110
Sensitivity
CentroidFold:
0.515
Murlet(seed):
0.015
Positive Predictive Value
CentroidFold:
0.633
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Murlet(seed):
0.110
Sensitivity
CentroidAlifold(20):
0.369
Murlet(seed):
0.015
Positive Predictive Value
CentroidAlifold(20):
0.891
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Murlet(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Murlet(seed):
0.110
Sensitivity
CentroidHomfold‑LAST:
0.359
Murlet(seed):
0.015
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Murlet(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Murlet(seed):
0.110
Sensitivity
Contrafold:
0.542
Murlet(seed):
0.015
Positive Predictive Value
Contrafold:
0.571
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Murlet(seed):
0.110
Sensitivity
MXScarna(seed):
0.382
Murlet(seed):
0.015
Positive Predictive Value
MXScarna(seed):
0.753
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Murlet(seed):
0.110
Sensitivity
RNAalifold(20):
0.359
Murlet(seed):
0.015
Positive Predictive Value
RNAalifold(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Murlet(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Murlet(seed):
0.110
Sensitivity
Sfold:
0.477
Murlet(seed):
0.015
Positive Predictive Value
Sfold:
0.582
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Murlet(seed):
0.110
Sensitivity
RNAalifold(seed):
0.328
Murlet(seed):
0.015
Positive Predictive Value
RNAalifold(seed):
0.835
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Murlet(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Murlet(seed):
0.110
Sensitivity
MaxExpect:
0.505
Murlet(seed):
0.015
Positive Predictive Value
MaxExpect:
0.528
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Murlet(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Murlet(seed):
0.110
Sensitivity
ProbKnot:
0.508
Murlet(seed):
0.015
Positive Predictive Value
ProbKnot:
0.517
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Murlet(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Murlet(seed):
0.110
Sensitivity
MXScarna(20):
0.353
Murlet(seed):
0.015
Positive Predictive Value
MXScarna(20):
0.710
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Murlet(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Murlet(seed):
0.110
Sensitivity
UNAFold:
0.491
Murlet(seed):
0.015
Positive Predictive Value
UNAFold:
0.498
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Murlet(seed)
Matthews Correlation Coefficient
Fold:
0.493
Murlet(seed):
0.110
Sensitivity
Fold:
0.496
Murlet(seed):
0.015
Positive Predictive Value
Fold:
0.490
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Murlet(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Murlet(seed):
0.110
Sensitivity
RNAfold:
0.486
Murlet(seed):
0.015
Positive Predictive Value
RNAfold:
0.485
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Murlet(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Murlet(seed):
0.110
Sensitivity
PknotsRG:
0.440
Murlet(seed):
0.015
Positive Predictive Value
PknotsRG:
0.510
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Murlet(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Murlet(seed):
0.110
Sensitivity
CRWrnafold:
0.379
Murlet(seed):
0.015
Positive Predictive Value
CRWrnafold:
0.490
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Murlet(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Murlet(seed):
0.110
Sensitivity
RNAsubopt:
0.344
Murlet(seed):
0.015
Positive Predictive Value
RNAsubopt:
0.519
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Murlet(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Murlet(seed):
0.110
Sensitivity
TurboFold(20):
0.210
Murlet(seed):
0.015
Positive Predictive Value
TurboFold(20):
0.837
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Murlet(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Murlet(seed):
0.110
Sensitivity
PPfold(20):
0.197
Murlet(seed):
0.015
Positive Predictive Value
PPfold(20):
0.879
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Murlet(seed)
Matthews Correlation Coefficient
Afold:
0.408
Murlet(seed):
0.110
Sensitivity
Afold:
0.343
Murlet(seed):
0.015
Positive Predictive Value
Afold:
0.487
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Murlet(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Murlet(seed):
0.110
Sensitivity
Carnac(20):
0.180
Murlet(seed):
0.015
Positive Predictive Value
Carnac(20):
0.892
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Murlet(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Murlet(seed):
0.110
Sensitivity
McQFold:
0.308
Murlet(seed):
0.015
Positive Predictive Value
McQFold:
0.492
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Murlet(seed):
0.110
Sensitivity
RSpredict(20):
0.241
Murlet(seed):
0.015
Positive Predictive Value
RSpredict(20):
0.590
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Murlet(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Murlet(seed):
0.110
Sensitivity
RNAshapes:
0.285
Murlet(seed):
0.015
Positive Predictive Value
RNAshapes:
0.474
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Murlet(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Murlet(seed):
0.110
Sensitivity
RNASLOpt:
0.261
Murlet(seed):
0.015
Positive Predictive Value
RNASLOpt:
0.510
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Murlet(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Murlet(seed):
0.110
Sensitivity
Multilign(20):
0.161
Murlet(seed):
0.015
Positive Predictive Value
Multilign(20):
0.762
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Murlet(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Murlet(seed):
0.110
Sensitivity
Murlet(20):
0.131
Murlet(seed):
0.015
Positive Predictive Value
Murlet(20):
0.787
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Murlet(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Murlet(seed):
0.110
Sensitivity
RNAwolf:
0.261
Murlet(seed):
0.015
Positive Predictive Value
RNAwolf:
0.369
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Murlet(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Murlet(seed):
0.110
Sensitivity
RSpredict(seed):
0.146
Murlet(seed):
0.015
Positive Predictive Value
RSpredict(seed):
0.597
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Murlet(seed):
0.110
Sensitivity
CMfinder(20):
0.107
Murlet(seed):
0.015
Positive Predictive Value
CMfinder(20):
0.794
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Murlet(seed):
0.110
Sensitivity
Vsfold4:
0.198
Murlet(seed):
0.015
Positive Predictive Value
Vsfold4:
0.371
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Murlet(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Murlet(seed):
0.110
Sensitivity
Vsfold5:
0.195
Murlet(seed):
0.015
Positive Predictive Value
Vsfold5:
0.347
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Murlet(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Murlet(seed):
0.110
Sensitivity
RNASampler(20):
0.069
Murlet(seed):
0.015
Positive Predictive Value
RNASampler(20):
0.877
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Murlet(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Murlet(seed):
0.110
Sensitivity
HotKnots:
0.069
Murlet(seed):
0.015
Positive Predictive Value
HotKnots:
0.659
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Murlet(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Murlet(seed):
0.110
Sensitivity
Mastr(20):
0.044
Murlet(seed):
0.015
Positive Predictive Value
Mastr(20):
0.790
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs Murlet(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Murlet(seed):
0.110
Sensitivity
Cylofold:
0.053
Murlet(seed):
0.015
Positive Predictive Value
Cylofold:
0.597
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs Murlet(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Murlet(seed):
0.110
Sensitivity
Pknots:
0.041
Murlet(seed):
0.015
Positive Predictive Value
Pknots:
0.662
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs Murlet(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Murlet(seed):
0.110
Sensitivity
Alterna:
0.036
Murlet(seed):
0.015
Positive Predictive Value
Alterna:
0.582
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs Murlet(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Murlet(seed):
0.110
Sensitivity
MCFold:
0.044
Murlet(seed):
0.015
Positive Predictive Value
MCFold:
0.432
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs Murlet(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Murlet(seed):
0.110
Sensitivity
RDfolder:
0.031
Murlet(seed):
0.015
Positive Predictive Value
RDfolder:
0.615
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
=
CMfinder(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Murlet(seed):
0.110
Sensitivity
CMfinder(seed):
0.016
Murlet(seed):
0.015
Positive Predictive Value
CMfinder(seed):
0.757
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.253241979086
|
+
Murlet(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
TurboFold(seed):
0.106
Sensitivity
Murlet(seed):
0.015
TurboFold(seed):
0.014
Positive Predictive Value
Murlet(seed):
0.811
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.50402730567e-05
|
+
Murlet(seed) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
PPfold(seed):
0.097
Sensitivity
Murlet(seed):
0.015
PPfold(seed):
0.012
Positive Predictive Value
Murlet(seed):
0.811
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
Carnac(seed):
0.093
Sensitivity
Murlet(seed):
0.015
Carnac(seed):
0.010
Positive Predictive Value
Murlet(seed):
0.811
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
RNASampler(seed):
0.088
Sensitivity
Murlet(seed):
0.015
RNASampler(seed):
0.009
Positive Predictive Value
Murlet(seed):
0.811
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
Mastr(seed):
0.072
Sensitivity
Murlet(seed):
0.015
Mastr(seed):
0.006
Positive Predictive Value
Murlet(seed):
0.811
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs NanoFolder
Matthews Correlation Coefficient
Murlet(seed):
0.110
NanoFolder:
0.068
Sensitivity
Murlet(seed):
0.015
NanoFolder:
0.027
Positive Predictive Value
Murlet(seed):
0.811
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Murlet(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
Multilign(seed):
0.059
Sensitivity
Murlet(seed):
0.015
Multilign(seed):
0.005
Positive Predictive Value
Murlet(seed):
0.811
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| CMfinder(seed) |
1242
ContextFold vs CMfinder(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
CMfinder(seed):
0.111
Sensitivity
ContextFold:
0.717
CMfinder(seed):
0.016
Positive Predictive Value
ContextFold:
0.794
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
CMfinder(seed):
0.111
Sensitivity
PETfold_pre2.0(seed):
0.667
CMfinder(seed):
0.016
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
CMfinder(seed):
0.111
Sensitivity
CentroidAlifold(seed):
0.414
CMfinder(seed):
0.016
Positive Predictive Value
CentroidAlifold(seed):
0.901
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs CMfinder(seed)
Matthews Correlation Coefficient
IPknot:
0.607
CMfinder(seed):
0.111
Sensitivity
IPknot:
0.551
CMfinder(seed):
0.016
Positive Predictive Value
IPknot:
0.670
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
CMfinder(seed):
0.111
Sensitivity
PETfold_pre2.0(20):
0.410
CMfinder(seed):
0.016
Positive Predictive Value
PETfold_pre2.0(20):
0.807
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
CMfinder(seed):
0.111
Sensitivity
CentroidFold:
0.515
CMfinder(seed):
0.016
Positive Predictive Value
CentroidFold:
0.633
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
CMfinder(seed):
0.111
Sensitivity
CentroidAlifold(20):
0.369
CMfinder(seed):
0.016
Positive Predictive Value
CentroidAlifold(20):
0.891
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs CMfinder(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
CMfinder(seed):
0.111
Sensitivity
CentroidHomfold‑LAST:
0.359
CMfinder(seed):
0.016
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs CMfinder(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
CMfinder(seed):
0.111
Sensitivity
Contrafold:
0.542
CMfinder(seed):
0.016
Positive Predictive Value
Contrafold:
0.571
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
CMfinder(seed):
0.111
Sensitivity
MXScarna(seed):
0.382
CMfinder(seed):
0.016
Positive Predictive Value
MXScarna(seed):
0.753
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
CMfinder(seed):
0.111
Sensitivity
RNAalifold(20):
0.359
CMfinder(seed):
0.016
Positive Predictive Value
RNAalifold(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs CMfinder(seed)
Matthews Correlation Coefficient
Sfold:
0.527
CMfinder(seed):
0.111
Sensitivity
Sfold:
0.477
CMfinder(seed):
0.016
Positive Predictive Value
Sfold:
0.582
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
CMfinder(seed):
0.111
Sensitivity
RNAalifold(seed):
0.328
CMfinder(seed):
0.016
Positive Predictive Value
RNAalifold(seed):
0.835
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs CMfinder(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
CMfinder(seed):
0.111
Sensitivity
MaxExpect:
0.505
CMfinder(seed):
0.016
Positive Predictive Value
MaxExpect:
0.528
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs CMfinder(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
CMfinder(seed):
0.111
Sensitivity
ProbKnot:
0.508
CMfinder(seed):
0.016
Positive Predictive Value
ProbKnot:
0.517
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs CMfinder(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
CMfinder(seed):
0.111
Sensitivity
MXScarna(20):
0.353
CMfinder(seed):
0.016
Positive Predictive Value
MXScarna(20):
0.710
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs CMfinder(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
CMfinder(seed):
0.111
Sensitivity
UNAFold:
0.491
CMfinder(seed):
0.016
Positive Predictive Value
UNAFold:
0.498
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs CMfinder(seed)
Matthews Correlation Coefficient
Fold:
0.493
CMfinder(seed):
0.111
Sensitivity
Fold:
0.496
CMfinder(seed):
0.016
Positive Predictive Value
Fold:
0.490
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs CMfinder(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
CMfinder(seed):
0.111
Sensitivity
RNAfold:
0.486
CMfinder(seed):
0.016
Positive Predictive Value
RNAfold:
0.485
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs CMfinder(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
CMfinder(seed):
0.111
Sensitivity
PknotsRG:
0.440
CMfinder(seed):
0.016
Positive Predictive Value
PknotsRG:
0.510
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs CMfinder(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
CMfinder(seed):
0.111
Sensitivity
CRWrnafold:
0.379
CMfinder(seed):
0.016
Positive Predictive Value
CRWrnafold:
0.490
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs CMfinder(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
CMfinder(seed):
0.111
Sensitivity
RNAsubopt:
0.344
CMfinder(seed):
0.016
Positive Predictive Value
RNAsubopt:
0.519
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
CMfinder(seed):
0.111
Sensitivity
TurboFold(20):
0.210
CMfinder(seed):
0.016
Positive Predictive Value
TurboFold(20):
0.837
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs CMfinder(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
CMfinder(seed):
0.111
Sensitivity
PPfold(20):
0.197
CMfinder(seed):
0.016
Positive Predictive Value
PPfold(20):
0.879
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs CMfinder(seed)
Matthews Correlation Coefficient
Afold:
0.408
CMfinder(seed):
0.111
Sensitivity
Afold:
0.343
CMfinder(seed):
0.016
Positive Predictive Value
Afold:
0.487
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
CMfinder(seed):
0.111
Sensitivity
Carnac(20):
0.180
CMfinder(seed):
0.016
Positive Predictive Value
Carnac(20):
0.892
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs CMfinder(seed)
Matthews Correlation Coefficient
McQFold:
0.389
CMfinder(seed):
0.111
Sensitivity
McQFold:
0.308
CMfinder(seed):
0.016
Positive Predictive Value
McQFold:
0.492
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
CMfinder(seed):
0.111
Sensitivity
RSpredict(20):
0.241
CMfinder(seed):
0.016
Positive Predictive Value
RSpredict(20):
0.590
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs CMfinder(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
CMfinder(seed):
0.111
Sensitivity
RNAshapes:
0.285
CMfinder(seed):
0.016
Positive Predictive Value
RNAshapes:
0.474
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs CMfinder(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
CMfinder(seed):
0.111
Sensitivity
RNASLOpt:
0.261
CMfinder(seed):
0.016
Positive Predictive Value
RNASLOpt:
0.510
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
CMfinder(seed):
0.111
Sensitivity
Multilign(20):
0.161
CMfinder(seed):
0.016
Positive Predictive Value
Multilign(20):
0.762
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
CMfinder(seed):
0.111
Sensitivity
Murlet(20):
0.131
CMfinder(seed):
0.016
Positive Predictive Value
Murlet(20):
0.787
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs CMfinder(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
CMfinder(seed):
0.111
Sensitivity
RNAwolf:
0.261
CMfinder(seed):
0.016
Positive Predictive Value
RNAwolf:
0.369
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs CMfinder(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
CMfinder(seed):
0.111
Sensitivity
RSpredict(seed):
0.146
CMfinder(seed):
0.016
Positive Predictive Value
RSpredict(seed):
0.597
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs CMfinder(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
CMfinder(seed):
0.111
Sensitivity
CMfinder(20):
0.107
CMfinder(seed):
0.016
Positive Predictive Value
CMfinder(20):
0.794
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
CMfinder(seed):
0.111
Sensitivity
Vsfold4:
0.198
CMfinder(seed):
0.016
Positive Predictive Value
Vsfold4:
0.371
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs CMfinder(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
CMfinder(seed):
0.111
Sensitivity
Vsfold5:
0.195
CMfinder(seed):
0.016
Positive Predictive Value
Vsfold5:
0.347
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs CMfinder(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
CMfinder(seed):
0.111
Sensitivity
RNASampler(20):
0.069
CMfinder(seed):
0.016
Positive Predictive Value
RNASampler(20):
0.877
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs CMfinder(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
CMfinder(seed):
0.111
Sensitivity
HotKnots:
0.069
CMfinder(seed):
0.016
Positive Predictive Value
HotKnots:
0.659
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs CMfinder(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
CMfinder(seed):
0.111
Sensitivity
Mastr(20):
0.044
CMfinder(seed):
0.016
Positive Predictive Value
Mastr(20):
0.790
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs CMfinder(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
CMfinder(seed):
0.111
Sensitivity
Cylofold:
0.053
CMfinder(seed):
0.016
Positive Predictive Value
Cylofold:
0.597
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs CMfinder(seed)
Matthews Correlation Coefficient
Pknots:
0.165
CMfinder(seed):
0.111
Sensitivity
Pknots:
0.041
CMfinder(seed):
0.016
Positive Predictive Value
Pknots:
0.662
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs CMfinder(seed)
Matthews Correlation Coefficient
Alterna:
0.145
CMfinder(seed):
0.111
Sensitivity
Alterna:
0.036
CMfinder(seed):
0.016
Positive Predictive Value
Alterna:
0.582
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs CMfinder(seed)
Matthews Correlation Coefficient
MCFold:
0.138
CMfinder(seed):
0.111
Sensitivity
MCFold:
0.044
CMfinder(seed):
0.016
Positive Predictive Value
MCFold:
0.432
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs CMfinder(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
CMfinder(seed):
0.111
Sensitivity
RDfolder:
0.031
CMfinder(seed):
0.016
Positive Predictive Value
RDfolder:
0.615
CMfinder(seed):
0.757
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs Murlet(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Murlet(seed):
0.110
Sensitivity
CMfinder(seed):
0.016
Murlet(seed):
0.015
Positive Predictive Value
CMfinder(seed):
0.757
Murlet(seed):
0.811
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 0.253241979086
|
|
+
CMfinder(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
TurboFold(seed):
0.106
Sensitivity
CMfinder(seed):
0.016
TurboFold(seed):
0.014
Positive Predictive Value
CMfinder(seed):
0.757
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 8.81311508309e-08
|
+
CMfinder(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
PPfold(seed):
0.097
Sensitivity
CMfinder(seed):
0.016
PPfold(seed):
0.012
Positive Predictive Value
CMfinder(seed):
0.757
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Carnac(seed):
0.093
Sensitivity
CMfinder(seed):
0.016
Carnac(seed):
0.010
Positive Predictive Value
CMfinder(seed):
0.757
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
RNASampler(seed):
0.088
Sensitivity
CMfinder(seed):
0.016
RNASampler(seed):
0.009
Positive Predictive Value
CMfinder(seed):
0.757
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Mastr(seed):
0.072
Sensitivity
CMfinder(seed):
0.016
Mastr(seed):
0.006
Positive Predictive Value
CMfinder(seed):
0.757
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(seed):
0.111
NanoFolder:
0.068
Sensitivity
CMfinder(seed):
0.016
NanoFolder:
0.027
Positive Predictive Value
CMfinder(seed):
0.757
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
CMfinder(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Multilign(seed):
0.059
Sensitivity
CMfinder(seed):
0.016
Multilign(seed):
0.005
Positive Predictive Value
CMfinder(seed):
0.757
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| TurboFold(seed) |
1242
ContextFold vs TurboFold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
TurboFold(seed):
0.106
Sensitivity
ContextFold:
0.717
TurboFold(seed):
0.014
Positive Predictive Value
ContextFold:
0.794
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
TurboFold(seed):
0.106
Sensitivity
PETfold_pre2.0(seed):
0.667
TurboFold(seed):
0.014
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
TurboFold(seed):
0.106
Sensitivity
CentroidAlifold(seed):
0.414
TurboFold(seed):
0.014
Positive Predictive Value
CentroidAlifold(seed):
0.901
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs TurboFold(seed)
Matthews Correlation Coefficient
IPknot:
0.607
TurboFold(seed):
0.106
Sensitivity
IPknot:
0.551
TurboFold(seed):
0.014
Positive Predictive Value
IPknot:
0.670
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
TurboFold(seed):
0.106
Sensitivity
PETfold_pre2.0(20):
0.410
TurboFold(seed):
0.014
Positive Predictive Value
PETfold_pre2.0(20):
0.807
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
TurboFold(seed):
0.106
Sensitivity
CentroidFold:
0.515
TurboFold(seed):
0.014
Positive Predictive Value
CentroidFold:
0.633
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
TurboFold(seed):
0.106
Sensitivity
CentroidAlifold(20):
0.369
TurboFold(seed):
0.014
Positive Predictive Value
CentroidAlifold(20):
0.891
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs TurboFold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
TurboFold(seed):
0.106
Sensitivity
CentroidHomfold‑LAST:
0.359
TurboFold(seed):
0.014
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs TurboFold(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
TurboFold(seed):
0.106
Sensitivity
Contrafold:
0.542
TurboFold(seed):
0.014
Positive Predictive Value
Contrafold:
0.571
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
TurboFold(seed):
0.106
Sensitivity
MXScarna(seed):
0.382
TurboFold(seed):
0.014
Positive Predictive Value
MXScarna(seed):
0.753
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
TurboFold(seed):
0.106
Sensitivity
RNAalifold(20):
0.359
TurboFold(seed):
0.014
Positive Predictive Value
RNAalifold(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs TurboFold(seed)
Matthews Correlation Coefficient
Sfold:
0.527
TurboFold(seed):
0.106
Sensitivity
Sfold:
0.477
TurboFold(seed):
0.014
Positive Predictive Value
Sfold:
0.582
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
TurboFold(seed):
0.106
Sensitivity
RNAalifold(seed):
0.328
TurboFold(seed):
0.014
Positive Predictive Value
RNAalifold(seed):
0.835
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs TurboFold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
TurboFold(seed):
0.106
Sensitivity
MaxExpect:
0.505
TurboFold(seed):
0.014
Positive Predictive Value
MaxExpect:
0.528
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs TurboFold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
TurboFold(seed):
0.106
Sensitivity
ProbKnot:
0.508
TurboFold(seed):
0.014
Positive Predictive Value
ProbKnot:
0.517
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs TurboFold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
TurboFold(seed):
0.106
Sensitivity
MXScarna(20):
0.353
TurboFold(seed):
0.014
Positive Predictive Value
MXScarna(20):
0.710
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs TurboFold(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
TurboFold(seed):
0.106
Sensitivity
UNAFold:
0.491
TurboFold(seed):
0.014
Positive Predictive Value
UNAFold:
0.498
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs TurboFold(seed)
Matthews Correlation Coefficient
Fold:
0.493
TurboFold(seed):
0.106
Sensitivity
Fold:
0.496
TurboFold(seed):
0.014
Positive Predictive Value
Fold:
0.490
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs TurboFold(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
TurboFold(seed):
0.106
Sensitivity
RNAfold:
0.486
TurboFold(seed):
0.014
Positive Predictive Value
RNAfold:
0.485
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs TurboFold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
TurboFold(seed):
0.106
Sensitivity
PknotsRG:
0.440
TurboFold(seed):
0.014
Positive Predictive Value
PknotsRG:
0.510
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs TurboFold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
TurboFold(seed):
0.106
Sensitivity
CRWrnafold:
0.379
TurboFold(seed):
0.014
Positive Predictive Value
CRWrnafold:
0.490
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs TurboFold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
TurboFold(seed):
0.106
Sensitivity
RNAsubopt:
0.344
TurboFold(seed):
0.014
Positive Predictive Value
RNAsubopt:
0.519
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
TurboFold(seed):
0.106
Sensitivity
TurboFold(20):
0.210
TurboFold(seed):
0.014
Positive Predictive Value
TurboFold(20):
0.837
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs TurboFold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
TurboFold(seed):
0.106
Sensitivity
PPfold(20):
0.197
TurboFold(seed):
0.014
Positive Predictive Value
PPfold(20):
0.879
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs TurboFold(seed)
Matthews Correlation Coefficient
Afold:
0.408
TurboFold(seed):
0.106
Sensitivity
Afold:
0.343
TurboFold(seed):
0.014
Positive Predictive Value
Afold:
0.487
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
TurboFold(seed):
0.106
Sensitivity
Carnac(20):
0.180
TurboFold(seed):
0.014
Positive Predictive Value
Carnac(20):
0.892
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs TurboFold(seed)
Matthews Correlation Coefficient
McQFold:
0.389
TurboFold(seed):
0.106
Sensitivity
McQFold:
0.308
TurboFold(seed):
0.014
Positive Predictive Value
McQFold:
0.492
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
TurboFold(seed):
0.106
Sensitivity
RSpredict(20):
0.241
TurboFold(seed):
0.014
Positive Predictive Value
RSpredict(20):
0.590
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs TurboFold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
TurboFold(seed):
0.106
Sensitivity
RNAshapes:
0.285
TurboFold(seed):
0.014
Positive Predictive Value
RNAshapes:
0.474
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs TurboFold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
TurboFold(seed):
0.106
Sensitivity
RNASLOpt:
0.261
TurboFold(seed):
0.014
Positive Predictive Value
RNASLOpt:
0.510
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
TurboFold(seed):
0.106
Sensitivity
Multilign(20):
0.161
TurboFold(seed):
0.014
Positive Predictive Value
Multilign(20):
0.762
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
TurboFold(seed):
0.106
Sensitivity
Murlet(20):
0.131
TurboFold(seed):
0.014
Positive Predictive Value
Murlet(20):
0.787
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs TurboFold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
TurboFold(seed):
0.106
Sensitivity
RNAwolf:
0.261
TurboFold(seed):
0.014
Positive Predictive Value
RNAwolf:
0.369
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
TurboFold(seed):
0.106
Sensitivity
RSpredict(seed):
0.146
TurboFold(seed):
0.014
Positive Predictive Value
RSpredict(seed):
0.597
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
TurboFold(seed):
0.106
Sensitivity
CMfinder(20):
0.107
TurboFold(seed):
0.014
Positive Predictive Value
CMfinder(20):
0.794
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
TurboFold(seed):
0.106
Sensitivity
Vsfold4:
0.198
TurboFold(seed):
0.014
Positive Predictive Value
Vsfold4:
0.371
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs TurboFold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
TurboFold(seed):
0.106
Sensitivity
Vsfold5:
0.195
TurboFold(seed):
0.014
Positive Predictive Value
Vsfold5:
0.347
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs TurboFold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
TurboFold(seed):
0.106
Sensitivity
RNASampler(20):
0.069
TurboFold(seed):
0.014
Positive Predictive Value
RNASampler(20):
0.877
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs TurboFold(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
TurboFold(seed):
0.106
Sensitivity
HotKnots:
0.069
TurboFold(seed):
0.014
Positive Predictive Value
HotKnots:
0.659
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs TurboFold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
TurboFold(seed):
0.106
Sensitivity
Mastr(20):
0.044
TurboFold(seed):
0.014
Positive Predictive Value
Mastr(20):
0.790
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs TurboFold(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
TurboFold(seed):
0.106
Sensitivity
Cylofold:
0.053
TurboFold(seed):
0.014
Positive Predictive Value
Cylofold:
0.597
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs TurboFold(seed)
Matthews Correlation Coefficient
Pknots:
0.165
TurboFold(seed):
0.106
Sensitivity
Pknots:
0.041
TurboFold(seed):
0.014
Positive Predictive Value
Pknots:
0.662
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs TurboFold(seed)
Matthews Correlation Coefficient
Alterna:
0.145
TurboFold(seed):
0.106
Sensitivity
Alterna:
0.036
TurboFold(seed):
0.014
Positive Predictive Value
Alterna:
0.582
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs TurboFold(seed)
Matthews Correlation Coefficient
MCFold:
0.138
TurboFold(seed):
0.106
Sensitivity
MCFold:
0.044
TurboFold(seed):
0.014
Positive Predictive Value
MCFold:
0.432
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs TurboFold(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
TurboFold(seed):
0.106
Sensitivity
RDfolder:
0.031
TurboFold(seed):
0.014
Positive Predictive Value
RDfolder:
0.615
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
TurboFold(seed):
0.106
Sensitivity
Murlet(seed):
0.015
TurboFold(seed):
0.014
Positive Predictive Value
Murlet(seed):
0.811
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.50402730567e-05
|
1242
CMfinder(seed) vs TurboFold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
TurboFold(seed):
0.106
Sensitivity
CMfinder(seed):
0.016
TurboFold(seed):
0.014
Positive Predictive Value
CMfinder(seed):
0.757
TurboFold(seed):
0.822
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 8.81311508309e-08
|
|
+
TurboFold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
PPfold(seed):
0.097
Sensitivity
TurboFold(seed):
0.014
PPfold(seed):
0.012
Positive Predictive Value
TurboFold(seed):
0.822
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
Carnac(seed):
0.093
Sensitivity
TurboFold(seed):
0.014
Carnac(seed):
0.010
Positive Predictive Value
TurboFold(seed):
0.822
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
RNASampler(seed):
0.088
Sensitivity
TurboFold(seed):
0.014
RNASampler(seed):
0.009
Positive Predictive Value
TurboFold(seed):
0.822
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
Mastr(seed):
0.072
Sensitivity
TurboFold(seed):
0.014
Mastr(seed):
0.006
Positive Predictive Value
TurboFold(seed):
0.822
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(seed):
0.106
NanoFolder:
0.068
Sensitivity
TurboFold(seed):
0.014
NanoFolder:
0.027
Positive Predictive Value
TurboFold(seed):
0.822
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
TurboFold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
Multilign(seed):
0.059
Sensitivity
TurboFold(seed):
0.014
Multilign(seed):
0.005
Positive Predictive Value
TurboFold(seed):
0.822
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| PPfold(seed) |
1242
ContextFold vs PPfold(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
PPfold(seed):
0.097
Sensitivity
ContextFold:
0.717
PPfold(seed):
0.012
Positive Predictive Value
ContextFold:
0.794
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
PPfold(seed):
0.097
Sensitivity
PETfold_pre2.0(seed):
0.667
PPfold(seed):
0.012
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
PPfold(seed):
0.097
Sensitivity
CentroidAlifold(seed):
0.414
PPfold(seed):
0.012
Positive Predictive Value
CentroidAlifold(seed):
0.901
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs PPfold(seed)
Matthews Correlation Coefficient
IPknot:
0.607
PPfold(seed):
0.097
Sensitivity
IPknot:
0.551
PPfold(seed):
0.012
Positive Predictive Value
IPknot:
0.670
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs PPfold(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
PPfold(seed):
0.097
Sensitivity
PETfold_pre2.0(20):
0.410
PPfold(seed):
0.012
Positive Predictive Value
PETfold_pre2.0(20):
0.807
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs PPfold(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
PPfold(seed):
0.097
Sensitivity
CentroidFold:
0.515
PPfold(seed):
0.012
Positive Predictive Value
CentroidFold:
0.633
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
PPfold(seed):
0.097
Sensitivity
CentroidAlifold(20):
0.369
PPfold(seed):
0.012
Positive Predictive Value
CentroidAlifold(20):
0.891
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs PPfold(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
PPfold(seed):
0.097
Sensitivity
CentroidHomfold‑LAST:
0.359
PPfold(seed):
0.012
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs PPfold(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
PPfold(seed):
0.097
Sensitivity
Contrafold:
0.542
PPfold(seed):
0.012
Positive Predictive Value
Contrafold:
0.571
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
PPfold(seed):
0.097
Sensitivity
MXScarna(seed):
0.382
PPfold(seed):
0.012
Positive Predictive Value
MXScarna(seed):
0.753
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
PPfold(seed):
0.097
Sensitivity
RNAalifold(20):
0.359
PPfold(seed):
0.012
Positive Predictive Value
RNAalifold(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs PPfold(seed)
Matthews Correlation Coefficient
Sfold:
0.527
PPfold(seed):
0.097
Sensitivity
Sfold:
0.477
PPfold(seed):
0.012
Positive Predictive Value
Sfold:
0.582
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
PPfold(seed):
0.097
Sensitivity
RNAalifold(seed):
0.328
PPfold(seed):
0.012
Positive Predictive Value
RNAalifold(seed):
0.835
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs PPfold(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
PPfold(seed):
0.097
Sensitivity
MaxExpect:
0.505
PPfold(seed):
0.012
Positive Predictive Value
MaxExpect:
0.528
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs PPfold(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
PPfold(seed):
0.097
Sensitivity
ProbKnot:
0.508
PPfold(seed):
0.012
Positive Predictive Value
ProbKnot:
0.517
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs PPfold(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
PPfold(seed):
0.097
Sensitivity
MXScarna(20):
0.353
PPfold(seed):
0.012
Positive Predictive Value
MXScarna(20):
0.710
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs PPfold(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
PPfold(seed):
0.097
Sensitivity
UNAFold:
0.491
PPfold(seed):
0.012
Positive Predictive Value
UNAFold:
0.498
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs PPfold(seed)
Matthews Correlation Coefficient
Fold:
0.493
PPfold(seed):
0.097
Sensitivity
Fold:
0.496
PPfold(seed):
0.012
Positive Predictive Value
Fold:
0.490
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs PPfold(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
PPfold(seed):
0.097
Sensitivity
RNAfold:
0.486
PPfold(seed):
0.012
Positive Predictive Value
RNAfold:
0.485
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs PPfold(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
PPfold(seed):
0.097
Sensitivity
PknotsRG:
0.440
PPfold(seed):
0.012
Positive Predictive Value
PknotsRG:
0.510
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs PPfold(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
PPfold(seed):
0.097
Sensitivity
CRWrnafold:
0.379
PPfold(seed):
0.012
Positive Predictive Value
CRWrnafold:
0.490
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs PPfold(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
PPfold(seed):
0.097
Sensitivity
RNAsubopt:
0.344
PPfold(seed):
0.012
Positive Predictive Value
RNAsubopt:
0.519
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
PPfold(seed):
0.097
Sensitivity
TurboFold(20):
0.210
PPfold(seed):
0.012
Positive Predictive Value
TurboFold(20):
0.837
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs PPfold(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
PPfold(seed):
0.097
Sensitivity
PPfold(20):
0.197
PPfold(seed):
0.012
Positive Predictive Value
PPfold(20):
0.879
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs PPfold(seed)
Matthews Correlation Coefficient
Afold:
0.408
PPfold(seed):
0.097
Sensitivity
Afold:
0.343
PPfold(seed):
0.012
Positive Predictive Value
Afold:
0.487
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs PPfold(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
PPfold(seed):
0.097
Sensitivity
Carnac(20):
0.180
PPfold(seed):
0.012
Positive Predictive Value
Carnac(20):
0.892
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs PPfold(seed)
Matthews Correlation Coefficient
McQFold:
0.389
PPfold(seed):
0.097
Sensitivity
McQFold:
0.308
PPfold(seed):
0.012
Positive Predictive Value
McQFold:
0.492
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
PPfold(seed):
0.097
Sensitivity
RSpredict(20):
0.241
PPfold(seed):
0.012
Positive Predictive Value
RSpredict(20):
0.590
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs PPfold(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
PPfold(seed):
0.097
Sensitivity
RNAshapes:
0.285
PPfold(seed):
0.012
Positive Predictive Value
RNAshapes:
0.474
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs PPfold(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
PPfold(seed):
0.097
Sensitivity
RNASLOpt:
0.261
PPfold(seed):
0.012
Positive Predictive Value
RNASLOpt:
0.510
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs PPfold(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
PPfold(seed):
0.097
Sensitivity
Multilign(20):
0.161
PPfold(seed):
0.012
Positive Predictive Value
Multilign(20):
0.762
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
PPfold(seed):
0.097
Sensitivity
Murlet(20):
0.131
PPfold(seed):
0.012
Positive Predictive Value
Murlet(20):
0.787
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs PPfold(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
PPfold(seed):
0.097
Sensitivity
RNAwolf:
0.261
PPfold(seed):
0.012
Positive Predictive Value
RNAwolf:
0.369
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs PPfold(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
PPfold(seed):
0.097
Sensitivity
RSpredict(seed):
0.146
PPfold(seed):
0.012
Positive Predictive Value
RSpredict(seed):
0.597
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
PPfold(seed):
0.097
Sensitivity
CMfinder(20):
0.107
PPfold(seed):
0.012
Positive Predictive Value
CMfinder(20):
0.794
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
PPfold(seed):
0.097
Sensitivity
Vsfold4:
0.198
PPfold(seed):
0.012
Positive Predictive Value
Vsfold4:
0.371
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs PPfold(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
PPfold(seed):
0.097
Sensitivity
Vsfold5:
0.195
PPfold(seed):
0.012
Positive Predictive Value
Vsfold5:
0.347
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs PPfold(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
PPfold(seed):
0.097
Sensitivity
RNASampler(20):
0.069
PPfold(seed):
0.012
Positive Predictive Value
RNASampler(20):
0.877
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs PPfold(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
PPfold(seed):
0.097
Sensitivity
HotKnots:
0.069
PPfold(seed):
0.012
Positive Predictive Value
HotKnots:
0.659
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs PPfold(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
PPfold(seed):
0.097
Sensitivity
Mastr(20):
0.044
PPfold(seed):
0.012
Positive Predictive Value
Mastr(20):
0.790
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs PPfold(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
PPfold(seed):
0.097
Sensitivity
Cylofold:
0.053
PPfold(seed):
0.012
Positive Predictive Value
Cylofold:
0.597
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs PPfold(seed)
Matthews Correlation Coefficient
Pknots:
0.165
PPfold(seed):
0.097
Sensitivity
Pknots:
0.041
PPfold(seed):
0.012
Positive Predictive Value
Pknots:
0.662
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs PPfold(seed)
Matthews Correlation Coefficient
Alterna:
0.145
PPfold(seed):
0.097
Sensitivity
Alterna:
0.036
PPfold(seed):
0.012
Positive Predictive Value
Alterna:
0.582
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs PPfold(seed)
Matthews Correlation Coefficient
MCFold:
0.138
PPfold(seed):
0.097
Sensitivity
MCFold:
0.044
PPfold(seed):
0.012
Positive Predictive Value
MCFold:
0.432
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs PPfold(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
PPfold(seed):
0.097
Sensitivity
RDfolder:
0.031
PPfold(seed):
0.012
Positive Predictive Value
RDfolder:
0.615
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs PPfold(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
PPfold(seed):
0.097
Sensitivity
Murlet(seed):
0.015
PPfold(seed):
0.012
Positive Predictive Value
Murlet(seed):
0.811
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs PPfold(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
PPfold(seed):
0.097
Sensitivity
CMfinder(seed):
0.016
PPfold(seed):
0.012
Positive Predictive Value
CMfinder(seed):
0.757
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(seed) vs PPfold(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
PPfold(seed):
0.097
Sensitivity
TurboFold(seed):
0.014
PPfold(seed):
0.012
Positive Predictive Value
TurboFold(seed):
0.822
PPfold(seed):
0.794
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
PPfold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
Carnac(seed):
0.093
Sensitivity
PPfold(seed):
0.012
Carnac(seed):
0.010
Positive Predictive Value
PPfold(seed):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 6.18847062639e-05
|
+
PPfold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
RNASampler(seed):
0.088
Sensitivity
PPfold(seed):
0.012
RNASampler(seed):
0.009
Positive Predictive Value
PPfold(seed):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
Mastr(seed):
0.072
Sensitivity
PPfold(seed):
0.012
Mastr(seed):
0.006
Positive Predictive Value
PPfold(seed):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(seed) vs NanoFolder
Matthews Correlation Coefficient
PPfold(seed):
0.097
NanoFolder:
0.068
Sensitivity
PPfold(seed):
0.012
NanoFolder:
0.027
Positive Predictive Value
PPfold(seed):
0.794
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
PPfold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
Multilign(seed):
0.059
Sensitivity
PPfold(seed):
0.012
Multilign(seed):
0.005
Positive Predictive Value
PPfold(seed):
0.794
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Carnac(seed) |
1242
ContextFold vs Carnac(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Carnac(seed):
0.093
Sensitivity
ContextFold:
0.717
Carnac(seed):
0.010
Positive Predictive Value
ContextFold:
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Carnac(seed):
0.093
Sensitivity
PETfold_pre2.0(seed):
0.667
Carnac(seed):
0.010
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Carnac(seed):
0.093
Sensitivity
CentroidAlifold(seed):
0.414
Carnac(seed):
0.010
Positive Predictive Value
CentroidAlifold(seed):
0.901
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Carnac(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Carnac(seed):
0.093
Sensitivity
IPknot:
0.551
Carnac(seed):
0.010
Positive Predictive Value
IPknot:
0.670
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Carnac(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Carnac(seed):
0.093
Sensitivity
PETfold_pre2.0(20):
0.410
Carnac(seed):
0.010
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Carnac(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Carnac(seed):
0.093
Sensitivity
CentroidFold:
0.515
Carnac(seed):
0.010
Positive Predictive Value
CentroidFold:
0.633
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Carnac(seed):
0.093
Sensitivity
CentroidAlifold(20):
0.369
Carnac(seed):
0.010
Positive Predictive Value
CentroidAlifold(20):
0.891
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Carnac(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Carnac(seed):
0.093
Sensitivity
CentroidHomfold‑LAST:
0.359
Carnac(seed):
0.010
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Carnac(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Carnac(seed):
0.093
Sensitivity
Contrafold:
0.542
Carnac(seed):
0.010
Positive Predictive Value
Contrafold:
0.571
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Carnac(seed):
0.093
Sensitivity
MXScarna(seed):
0.382
Carnac(seed):
0.010
Positive Predictive Value
MXScarna(seed):
0.753
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Carnac(seed):
0.093
Sensitivity
RNAalifold(20):
0.359
Carnac(seed):
0.010
Positive Predictive Value
RNAalifold(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Carnac(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Carnac(seed):
0.093
Sensitivity
Sfold:
0.477
Carnac(seed):
0.010
Positive Predictive Value
Sfold:
0.582
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Carnac(seed):
0.093
Sensitivity
RNAalifold(seed):
0.328
Carnac(seed):
0.010
Positive Predictive Value
RNAalifold(seed):
0.835
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Carnac(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Carnac(seed):
0.093
Sensitivity
MaxExpect:
0.505
Carnac(seed):
0.010
Positive Predictive Value
MaxExpect:
0.528
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Carnac(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Carnac(seed):
0.093
Sensitivity
ProbKnot:
0.508
Carnac(seed):
0.010
Positive Predictive Value
ProbKnot:
0.517
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Carnac(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Carnac(seed):
0.093
Sensitivity
MXScarna(20):
0.353
Carnac(seed):
0.010
Positive Predictive Value
MXScarna(20):
0.710
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Carnac(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Carnac(seed):
0.093
Sensitivity
UNAFold:
0.491
Carnac(seed):
0.010
Positive Predictive Value
UNAFold:
0.498
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Carnac(seed)
Matthews Correlation Coefficient
Fold:
0.493
Carnac(seed):
0.093
Sensitivity
Fold:
0.496
Carnac(seed):
0.010
Positive Predictive Value
Fold:
0.490
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Carnac(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Carnac(seed):
0.093
Sensitivity
RNAfold:
0.486
Carnac(seed):
0.010
Positive Predictive Value
RNAfold:
0.485
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Carnac(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Carnac(seed):
0.093
Sensitivity
PknotsRG:
0.440
Carnac(seed):
0.010
Positive Predictive Value
PknotsRG:
0.510
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Carnac(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Carnac(seed):
0.093
Sensitivity
CRWrnafold:
0.379
Carnac(seed):
0.010
Positive Predictive Value
CRWrnafold:
0.490
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Carnac(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Carnac(seed):
0.093
Sensitivity
RNAsubopt:
0.344
Carnac(seed):
0.010
Positive Predictive Value
RNAsubopt:
0.519
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Carnac(seed):
0.093
Sensitivity
TurboFold(20):
0.210
Carnac(seed):
0.010
Positive Predictive Value
TurboFold(20):
0.837
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Carnac(seed):
0.093
Sensitivity
PPfold(20):
0.197
Carnac(seed):
0.010
Positive Predictive Value
PPfold(20):
0.879
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Carnac(seed)
Matthews Correlation Coefficient
Afold:
0.408
Carnac(seed):
0.093
Sensitivity
Afold:
0.343
Carnac(seed):
0.010
Positive Predictive Value
Afold:
0.487
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Carnac(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Carnac(seed):
0.093
Sensitivity
Carnac(20):
0.180
Carnac(seed):
0.010
Positive Predictive Value
Carnac(20):
0.892
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Carnac(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Carnac(seed):
0.093
Sensitivity
McQFold:
0.308
Carnac(seed):
0.010
Positive Predictive Value
McQFold:
0.492
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Carnac(seed):
0.093
Sensitivity
RSpredict(20):
0.241
Carnac(seed):
0.010
Positive Predictive Value
RSpredict(20):
0.590
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Carnac(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Carnac(seed):
0.093
Sensitivity
RNAshapes:
0.285
Carnac(seed):
0.010
Positive Predictive Value
RNAshapes:
0.474
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Carnac(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Carnac(seed):
0.093
Sensitivity
RNASLOpt:
0.261
Carnac(seed):
0.010
Positive Predictive Value
RNASLOpt:
0.510
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Carnac(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Carnac(seed):
0.093
Sensitivity
Multilign(20):
0.161
Carnac(seed):
0.010
Positive Predictive Value
Multilign(20):
0.762
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Carnac(seed):
0.093
Sensitivity
Murlet(20):
0.131
Carnac(seed):
0.010
Positive Predictive Value
Murlet(20):
0.787
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Carnac(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Carnac(seed):
0.093
Sensitivity
RNAwolf:
0.261
Carnac(seed):
0.010
Positive Predictive Value
RNAwolf:
0.369
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Carnac(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Carnac(seed):
0.093
Sensitivity
RSpredict(seed):
0.146
Carnac(seed):
0.010
Positive Predictive Value
RSpredict(seed):
0.597
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Carnac(seed):
0.093
Sensitivity
CMfinder(20):
0.107
Carnac(seed):
0.010
Positive Predictive Value
CMfinder(20):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Carnac(seed):
0.093
Sensitivity
Vsfold4:
0.198
Carnac(seed):
0.010
Positive Predictive Value
Vsfold4:
0.371
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Carnac(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Carnac(seed):
0.093
Sensitivity
Vsfold5:
0.195
Carnac(seed):
0.010
Positive Predictive Value
Vsfold5:
0.347
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Carnac(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Carnac(seed):
0.093
Sensitivity
RNASampler(20):
0.069
Carnac(seed):
0.010
Positive Predictive Value
RNASampler(20):
0.877
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Carnac(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Carnac(seed):
0.093
Sensitivity
HotKnots:
0.069
Carnac(seed):
0.010
Positive Predictive Value
HotKnots:
0.659
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Carnac(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Carnac(seed):
0.093
Sensitivity
Mastr(20):
0.044
Carnac(seed):
0.010
Positive Predictive Value
Mastr(20):
0.790
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs Carnac(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Carnac(seed):
0.093
Sensitivity
Cylofold:
0.053
Carnac(seed):
0.010
Positive Predictive Value
Cylofold:
0.597
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs Carnac(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Carnac(seed):
0.093
Sensitivity
Pknots:
0.041
Carnac(seed):
0.010
Positive Predictive Value
Pknots:
0.662
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs Carnac(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Carnac(seed):
0.093
Sensitivity
Alterna:
0.036
Carnac(seed):
0.010
Positive Predictive Value
Alterna:
0.582
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs Carnac(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Carnac(seed):
0.093
Sensitivity
MCFold:
0.044
Carnac(seed):
0.010
Positive Predictive Value
MCFold:
0.432
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs Carnac(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Carnac(seed):
0.093
Sensitivity
RDfolder:
0.031
Carnac(seed):
0.010
Positive Predictive Value
RDfolder:
0.615
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs Carnac(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
Carnac(seed):
0.093
Sensitivity
Murlet(seed):
0.015
Carnac(seed):
0.010
Positive Predictive Value
Murlet(seed):
0.811
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs Carnac(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Carnac(seed):
0.093
Sensitivity
CMfinder(seed):
0.016
Carnac(seed):
0.010
Positive Predictive Value
CMfinder(seed):
0.757
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
Carnac(seed):
0.093
Sensitivity
TurboFold(seed):
0.014
Carnac(seed):
0.010
Positive Predictive Value
TurboFold(seed):
0.822
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(seed) vs Carnac(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
Carnac(seed):
0.093
Sensitivity
PPfold(seed):
0.012
Carnac(seed):
0.010
Positive Predictive Value
PPfold(seed):
0.794
Carnac(seed):
0.867
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 6.18847062639e-05
|
|
+
Carnac(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.093
RNASampler(seed):
0.088
Sensitivity
Carnac(seed):
0.010
RNASampler(seed):
0.009
Positive Predictive Value
Carnac(seed):
0.867
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 2.45033651981e-07
|
+
Carnac(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.093
Mastr(seed):
0.072
Sensitivity
Carnac(seed):
0.010
Mastr(seed):
0.006
Positive Predictive Value
Carnac(seed):
0.867
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(seed) vs NanoFolder
Matthews Correlation Coefficient
Carnac(seed):
0.093
NanoFolder:
0.068
Sensitivity
Carnac(seed):
0.010
NanoFolder:
0.027
Positive Predictive Value
Carnac(seed):
0.867
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
Carnac(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.093
Multilign(seed):
0.059
Sensitivity
Carnac(seed):
0.010
Multilign(seed):
0.005
Positive Predictive Value
Carnac(seed):
0.867
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| RNASampler(seed) |
1242
ContextFold vs RNASampler(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
RNASampler(seed):
0.088
Sensitivity
ContextFold:
0.717
RNASampler(seed):
0.009
Positive Predictive Value
ContextFold:
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
RNASampler(seed):
0.088
Sensitivity
PETfold_pre2.0(seed):
0.667
RNASampler(seed):
0.009
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
RNASampler(seed):
0.088
Sensitivity
CentroidAlifold(seed):
0.414
RNASampler(seed):
0.009
Positive Predictive Value
CentroidAlifold(seed):
0.901
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs RNASampler(seed)
Matthews Correlation Coefficient
IPknot:
0.607
RNASampler(seed):
0.088
Sensitivity
IPknot:
0.551
RNASampler(seed):
0.009
Positive Predictive Value
IPknot:
0.670
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
RNASampler(seed):
0.088
Sensitivity
PETfold_pre2.0(20):
0.410
RNASampler(seed):
0.009
Positive Predictive Value
PETfold_pre2.0(20):
0.807
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
RNASampler(seed):
0.088
Sensitivity
CentroidFold:
0.515
RNASampler(seed):
0.009
Positive Predictive Value
CentroidFold:
0.633
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
RNASampler(seed):
0.088
Sensitivity
CentroidAlifold(20):
0.369
RNASampler(seed):
0.009
Positive Predictive Value
CentroidAlifold(20):
0.891
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs RNASampler(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
RNASampler(seed):
0.088
Sensitivity
CentroidHomfold‑LAST:
0.359
RNASampler(seed):
0.009
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs RNASampler(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
RNASampler(seed):
0.088
Sensitivity
Contrafold:
0.542
RNASampler(seed):
0.009
Positive Predictive Value
Contrafold:
0.571
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
RNASampler(seed):
0.088
Sensitivity
MXScarna(seed):
0.382
RNASampler(seed):
0.009
Positive Predictive Value
MXScarna(seed):
0.753
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
RNASampler(seed):
0.088
Sensitivity
RNAalifold(20):
0.359
RNASampler(seed):
0.009
Positive Predictive Value
RNAalifold(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs RNASampler(seed)
Matthews Correlation Coefficient
Sfold:
0.527
RNASampler(seed):
0.088
Sensitivity
Sfold:
0.477
RNASampler(seed):
0.009
Positive Predictive Value
Sfold:
0.582
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
RNASampler(seed):
0.088
Sensitivity
RNAalifold(seed):
0.328
RNASampler(seed):
0.009
Positive Predictive Value
RNAalifold(seed):
0.835
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs RNASampler(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
RNASampler(seed):
0.088
Sensitivity
MaxExpect:
0.505
RNASampler(seed):
0.009
Positive Predictive Value
MaxExpect:
0.528
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs RNASampler(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
RNASampler(seed):
0.088
Sensitivity
ProbKnot:
0.508
RNASampler(seed):
0.009
Positive Predictive Value
ProbKnot:
0.517
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs RNASampler(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
RNASampler(seed):
0.088
Sensitivity
MXScarna(20):
0.353
RNASampler(seed):
0.009
Positive Predictive Value
MXScarna(20):
0.710
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs RNASampler(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
RNASampler(seed):
0.088
Sensitivity
UNAFold:
0.491
RNASampler(seed):
0.009
Positive Predictive Value
UNAFold:
0.498
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs RNASampler(seed)
Matthews Correlation Coefficient
Fold:
0.493
RNASampler(seed):
0.088
Sensitivity
Fold:
0.496
RNASampler(seed):
0.009
Positive Predictive Value
Fold:
0.490
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs RNASampler(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
RNASampler(seed):
0.088
Sensitivity
RNAfold:
0.486
RNASampler(seed):
0.009
Positive Predictive Value
RNAfold:
0.485
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs RNASampler(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
RNASampler(seed):
0.088
Sensitivity
PknotsRG:
0.440
RNASampler(seed):
0.009
Positive Predictive Value
PknotsRG:
0.510
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs RNASampler(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
RNASampler(seed):
0.088
Sensitivity
CRWrnafold:
0.379
RNASampler(seed):
0.009
Positive Predictive Value
CRWrnafold:
0.490
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs RNASampler(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
RNASampler(seed):
0.088
Sensitivity
RNAsubopt:
0.344
RNASampler(seed):
0.009
Positive Predictive Value
RNAsubopt:
0.519
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
RNASampler(seed):
0.088
Sensitivity
TurboFold(20):
0.210
RNASampler(seed):
0.009
Positive Predictive Value
TurboFold(20):
0.837
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
RNASampler(seed):
0.088
Sensitivity
PPfold(20):
0.197
RNASampler(seed):
0.009
Positive Predictive Value
PPfold(20):
0.879
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs RNASampler(seed)
Matthews Correlation Coefficient
Afold:
0.408
RNASampler(seed):
0.088
Sensitivity
Afold:
0.343
RNASampler(seed):
0.009
Positive Predictive Value
Afold:
0.487
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
RNASampler(seed):
0.088
Sensitivity
Carnac(20):
0.180
RNASampler(seed):
0.009
Positive Predictive Value
Carnac(20):
0.892
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs RNASampler(seed)
Matthews Correlation Coefficient
McQFold:
0.389
RNASampler(seed):
0.088
Sensitivity
McQFold:
0.308
RNASampler(seed):
0.009
Positive Predictive Value
McQFold:
0.492
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
RNASampler(seed):
0.088
Sensitivity
RSpredict(20):
0.241
RNASampler(seed):
0.009
Positive Predictive Value
RSpredict(20):
0.590
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs RNASampler(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
RNASampler(seed):
0.088
Sensitivity
RNAshapes:
0.285
RNASampler(seed):
0.009
Positive Predictive Value
RNAshapes:
0.474
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs RNASampler(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
RNASampler(seed):
0.088
Sensitivity
RNASLOpt:
0.261
RNASampler(seed):
0.009
Positive Predictive Value
RNASLOpt:
0.510
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
RNASampler(seed):
0.088
Sensitivity
Multilign(20):
0.161
RNASampler(seed):
0.009
Positive Predictive Value
Multilign(20):
0.762
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
RNASampler(seed):
0.088
Sensitivity
Murlet(20):
0.131
RNASampler(seed):
0.009
Positive Predictive Value
Murlet(20):
0.787
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs RNASampler(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
RNASampler(seed):
0.088
Sensitivity
RNAwolf:
0.261
RNASampler(seed):
0.009
Positive Predictive Value
RNAwolf:
0.369
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
RNASampler(seed):
0.088
Sensitivity
RSpredict(seed):
0.146
RNASampler(seed):
0.009
Positive Predictive Value
RSpredict(seed):
0.597
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
RNASampler(seed):
0.088
Sensitivity
CMfinder(20):
0.107
RNASampler(seed):
0.009
Positive Predictive Value
CMfinder(20):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
RNASampler(seed):
0.088
Sensitivity
Vsfold4:
0.198
RNASampler(seed):
0.009
Positive Predictive Value
Vsfold4:
0.371
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs RNASampler(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
RNASampler(seed):
0.088
Sensitivity
Vsfold5:
0.195
RNASampler(seed):
0.009
Positive Predictive Value
Vsfold5:
0.347
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs RNASampler(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
RNASampler(seed):
0.088
Sensitivity
RNASampler(20):
0.069
RNASampler(seed):
0.009
Positive Predictive Value
RNASampler(20):
0.877
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs RNASampler(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
RNASampler(seed):
0.088
Sensitivity
HotKnots:
0.069
RNASampler(seed):
0.009
Positive Predictive Value
HotKnots:
0.659
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs RNASampler(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
RNASampler(seed):
0.088
Sensitivity
Mastr(20):
0.044
RNASampler(seed):
0.009
Positive Predictive Value
Mastr(20):
0.790
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs RNASampler(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
RNASampler(seed):
0.088
Sensitivity
Cylofold:
0.053
RNASampler(seed):
0.009
Positive Predictive Value
Cylofold:
0.597
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs RNASampler(seed)
Matthews Correlation Coefficient
Pknots:
0.165
RNASampler(seed):
0.088
Sensitivity
Pknots:
0.041
RNASampler(seed):
0.009
Positive Predictive Value
Pknots:
0.662
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs RNASampler(seed)
Matthews Correlation Coefficient
Alterna:
0.145
RNASampler(seed):
0.088
Sensitivity
Alterna:
0.036
RNASampler(seed):
0.009
Positive Predictive Value
Alterna:
0.582
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs RNASampler(seed)
Matthews Correlation Coefficient
MCFold:
0.138
RNASampler(seed):
0.088
Sensitivity
MCFold:
0.044
RNASampler(seed):
0.009
Positive Predictive Value
MCFold:
0.432
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs RNASampler(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
RNASampler(seed):
0.088
Sensitivity
RDfolder:
0.031
RNASampler(seed):
0.009
Positive Predictive Value
RDfolder:
0.615
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
RNASampler(seed):
0.088
Sensitivity
Murlet(seed):
0.015
RNASampler(seed):
0.009
Positive Predictive Value
Murlet(seed):
0.811
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
RNASampler(seed):
0.088
Sensitivity
CMfinder(seed):
0.016
RNASampler(seed):
0.009
Positive Predictive Value
CMfinder(seed):
0.757
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
RNASampler(seed):
0.088
Sensitivity
TurboFold(seed):
0.014
RNASampler(seed):
0.009
Positive Predictive Value
TurboFold(seed):
0.822
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
RNASampler(seed):
0.088
Sensitivity
PPfold(seed):
0.012
RNASampler(seed):
0.009
Positive Predictive Value
PPfold(seed):
0.794
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(seed) vs RNASampler(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.093
RNASampler(seed):
0.088
Sensitivity
Carnac(seed):
0.010
RNASampler(seed):
0.009
Positive Predictive Value
Carnac(seed):
0.867
RNASampler(seed):
0.847
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 2.45033651981e-07
|
|
+
RNASampler(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.088
Mastr(seed):
0.072
Sensitivity
RNASampler(seed):
0.009
Mastr(seed):
0.006
Positive Predictive Value
RNASampler(seed):
0.847
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(seed) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(seed):
0.088
NanoFolder:
0.068
Sensitivity
RNASampler(seed):
0.009
NanoFolder:
0.027
Positive Predictive Value
RNASampler(seed):
0.847
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
+
RNASampler(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.088
Multilign(seed):
0.059
Sensitivity
RNASampler(seed):
0.009
Multilign(seed):
0.005
Positive Predictive Value
RNASampler(seed):
0.847
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Mastr(seed) |
1242
ContextFold vs Mastr(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Mastr(seed):
0.072
Sensitivity
ContextFold:
0.717
Mastr(seed):
0.006
Positive Predictive Value
ContextFold:
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Mastr(seed):
0.072
Sensitivity
PETfold_pre2.0(seed):
0.667
Mastr(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Mastr(seed):
0.072
Sensitivity
CentroidAlifold(seed):
0.414
Mastr(seed):
0.006
Positive Predictive Value
CentroidAlifold(seed):
0.901
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Mastr(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Mastr(seed):
0.072
Sensitivity
IPknot:
0.551
Mastr(seed):
0.006
Positive Predictive Value
IPknot:
0.670
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Mastr(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Mastr(seed):
0.072
Sensitivity
PETfold_pre2.0(20):
0.410
Mastr(seed):
0.006
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Mastr(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Mastr(seed):
0.072
Sensitivity
CentroidFold:
0.515
Mastr(seed):
0.006
Positive Predictive Value
CentroidFold:
0.633
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Mastr(seed):
0.072
Sensitivity
CentroidAlifold(20):
0.369
Mastr(seed):
0.006
Positive Predictive Value
CentroidAlifold(20):
0.891
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Mastr(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Mastr(seed):
0.072
Sensitivity
CentroidHomfold‑LAST:
0.359
Mastr(seed):
0.006
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Mastr(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Mastr(seed):
0.072
Sensitivity
Contrafold:
0.542
Mastr(seed):
0.006
Positive Predictive Value
Contrafold:
0.571
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Mastr(seed):
0.072
Sensitivity
MXScarna(seed):
0.382
Mastr(seed):
0.006
Positive Predictive Value
MXScarna(seed):
0.753
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Mastr(seed):
0.072
Sensitivity
RNAalifold(20):
0.359
Mastr(seed):
0.006
Positive Predictive Value
RNAalifold(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Mastr(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Mastr(seed):
0.072
Sensitivity
Sfold:
0.477
Mastr(seed):
0.006
Positive Predictive Value
Sfold:
0.582
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Mastr(seed):
0.072
Sensitivity
RNAalifold(seed):
0.328
Mastr(seed):
0.006
Positive Predictive Value
RNAalifold(seed):
0.835
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Mastr(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Mastr(seed):
0.072
Sensitivity
MaxExpect:
0.505
Mastr(seed):
0.006
Positive Predictive Value
MaxExpect:
0.528
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Mastr(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Mastr(seed):
0.072
Sensitivity
ProbKnot:
0.508
Mastr(seed):
0.006
Positive Predictive Value
ProbKnot:
0.517
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Mastr(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Mastr(seed):
0.072
Sensitivity
MXScarna(20):
0.353
Mastr(seed):
0.006
Positive Predictive Value
MXScarna(20):
0.710
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Mastr(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Mastr(seed):
0.072
Sensitivity
UNAFold:
0.491
Mastr(seed):
0.006
Positive Predictive Value
UNAFold:
0.498
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Mastr(seed)
Matthews Correlation Coefficient
Fold:
0.493
Mastr(seed):
0.072
Sensitivity
Fold:
0.496
Mastr(seed):
0.006
Positive Predictive Value
Fold:
0.490
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Mastr(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Mastr(seed):
0.072
Sensitivity
RNAfold:
0.486
Mastr(seed):
0.006
Positive Predictive Value
RNAfold:
0.485
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Mastr(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Mastr(seed):
0.072
Sensitivity
PknotsRG:
0.440
Mastr(seed):
0.006
Positive Predictive Value
PknotsRG:
0.510
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Mastr(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Mastr(seed):
0.072
Sensitivity
CRWrnafold:
0.379
Mastr(seed):
0.006
Positive Predictive Value
CRWrnafold:
0.490
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Mastr(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Mastr(seed):
0.072
Sensitivity
RNAsubopt:
0.344
Mastr(seed):
0.006
Positive Predictive Value
RNAsubopt:
0.519
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Mastr(seed):
0.072
Sensitivity
TurboFold(20):
0.210
Mastr(seed):
0.006
Positive Predictive Value
TurboFold(20):
0.837
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Mastr(seed):
0.072
Sensitivity
PPfold(20):
0.197
Mastr(seed):
0.006
Positive Predictive Value
PPfold(20):
0.879
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Mastr(seed)
Matthews Correlation Coefficient
Afold:
0.408
Mastr(seed):
0.072
Sensitivity
Afold:
0.343
Mastr(seed):
0.006
Positive Predictive Value
Afold:
0.487
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Mastr(seed):
0.072
Sensitivity
Carnac(20):
0.180
Mastr(seed):
0.006
Positive Predictive Value
Carnac(20):
0.892
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Mastr(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Mastr(seed):
0.072
Sensitivity
McQFold:
0.308
Mastr(seed):
0.006
Positive Predictive Value
McQFold:
0.492
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Mastr(seed):
0.072
Sensitivity
RSpredict(20):
0.241
Mastr(seed):
0.006
Positive Predictive Value
RSpredict(20):
0.590
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Mastr(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Mastr(seed):
0.072
Sensitivity
RNAshapes:
0.285
Mastr(seed):
0.006
Positive Predictive Value
RNAshapes:
0.474
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Mastr(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Mastr(seed):
0.072
Sensitivity
RNASLOpt:
0.261
Mastr(seed):
0.006
Positive Predictive Value
RNASLOpt:
0.510
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Mastr(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Mastr(seed):
0.072
Sensitivity
Multilign(20):
0.161
Mastr(seed):
0.006
Positive Predictive Value
Multilign(20):
0.762
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Mastr(seed):
0.072
Sensitivity
Murlet(20):
0.131
Mastr(seed):
0.006
Positive Predictive Value
Murlet(20):
0.787
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Mastr(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Mastr(seed):
0.072
Sensitivity
RNAwolf:
0.261
Mastr(seed):
0.006
Positive Predictive Value
RNAwolf:
0.369
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Mastr(seed):
0.072
Sensitivity
RSpredict(seed):
0.146
Mastr(seed):
0.006
Positive Predictive Value
RSpredict(seed):
0.597
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Mastr(seed):
0.072
Sensitivity
CMfinder(20):
0.107
Mastr(seed):
0.006
Positive Predictive Value
CMfinder(20):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Mastr(seed):
0.072
Sensitivity
Vsfold4:
0.198
Mastr(seed):
0.006
Positive Predictive Value
Vsfold4:
0.371
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Mastr(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Mastr(seed):
0.072
Sensitivity
Vsfold5:
0.195
Mastr(seed):
0.006
Positive Predictive Value
Vsfold5:
0.347
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Mastr(seed):
0.072
Sensitivity
RNASampler(20):
0.069
Mastr(seed):
0.006
Positive Predictive Value
RNASampler(20):
0.877
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Mastr(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Mastr(seed):
0.072
Sensitivity
HotKnots:
0.069
Mastr(seed):
0.006
Positive Predictive Value
HotKnots:
0.659
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Mastr(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Mastr(seed):
0.072
Sensitivity
Mastr(20):
0.044
Mastr(seed):
0.006
Positive Predictive Value
Mastr(20):
0.790
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs Mastr(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Mastr(seed):
0.072
Sensitivity
Cylofold:
0.053
Mastr(seed):
0.006
Positive Predictive Value
Cylofold:
0.597
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs Mastr(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Mastr(seed):
0.072
Sensitivity
Pknots:
0.041
Mastr(seed):
0.006
Positive Predictive Value
Pknots:
0.662
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs Mastr(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Mastr(seed):
0.072
Sensitivity
Alterna:
0.036
Mastr(seed):
0.006
Positive Predictive Value
Alterna:
0.582
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs Mastr(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Mastr(seed):
0.072
Sensitivity
MCFold:
0.044
Mastr(seed):
0.006
Positive Predictive Value
MCFold:
0.432
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs Mastr(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Mastr(seed):
0.072
Sensitivity
RDfolder:
0.031
Mastr(seed):
0.006
Positive Predictive Value
RDfolder:
0.615
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
Mastr(seed):
0.072
Sensitivity
Murlet(seed):
0.015
Mastr(seed):
0.006
Positive Predictive Value
Murlet(seed):
0.811
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs Mastr(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Mastr(seed):
0.072
Sensitivity
CMfinder(seed):
0.016
Mastr(seed):
0.006
Positive Predictive Value
CMfinder(seed):
0.757
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
Mastr(seed):
0.072
Sensitivity
TurboFold(seed):
0.014
Mastr(seed):
0.006
Positive Predictive Value
TurboFold(seed):
0.822
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(seed) vs Mastr(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
Mastr(seed):
0.072
Sensitivity
PPfold(seed):
0.012
Mastr(seed):
0.006
Positive Predictive Value
PPfold(seed):
0.794
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(seed) vs Mastr(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.093
Mastr(seed):
0.072
Sensitivity
Carnac(seed):
0.010
Mastr(seed):
0.006
Positive Predictive Value
Carnac(seed):
0.867
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(seed) vs Mastr(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.088
Mastr(seed):
0.072
Sensitivity
RNASampler(seed):
0.009
Mastr(seed):
0.006
Positive Predictive Value
RNASampler(seed):
0.847
Mastr(seed):
0.820
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|
+
Mastr(seed) vs NanoFolder
Matthews Correlation Coefficient
Mastr(seed):
0.072
NanoFolder:
0.068
Sensitivity
Mastr(seed):
0.006
NanoFolder:
0.027
Positive Predictive Value
Mastr(seed):
0.820
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.14023792096e-06
|
+
Mastr(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(seed):
0.072
Multilign(seed):
0.059
Sensitivity
Mastr(seed):
0.006
Multilign(seed):
0.005
Positive Predictive Value
Mastr(seed):
0.820
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| NanoFolder |
1242
ContextFold vs NanoFolder
Matthews Correlation Coefficient
ContextFold:
0.754
NanoFolder:
0.068
Sensitivity
ContextFold:
0.717
NanoFolder:
0.027
Positive Predictive Value
ContextFold:
0.794
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
NanoFolder:
0.068
Sensitivity
PETfold_pre2.0(seed):
0.667
NanoFolder:
0.027
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
NanoFolder:
0.068
Sensitivity
CentroidAlifold(seed):
0.414
NanoFolder:
0.027
Positive Predictive Value
CentroidAlifold(seed):
0.901
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs NanoFolder
Matthews Correlation Coefficient
IPknot:
0.607
NanoFolder:
0.068
Sensitivity
IPknot:
0.551
NanoFolder:
0.027
Positive Predictive Value
IPknot:
0.670
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs NanoFolder
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
NanoFolder:
0.068
Sensitivity
PETfold_pre2.0(20):
0.410
NanoFolder:
0.027
Positive Predictive Value
PETfold_pre2.0(20):
0.807
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs NanoFolder
Matthews Correlation Coefficient
CentroidFold:
0.571
NanoFolder:
0.068
Sensitivity
CentroidFold:
0.515
NanoFolder:
0.027
Positive Predictive Value
CentroidFold:
0.633
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs NanoFolder
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
NanoFolder:
0.068
Sensitivity
CentroidAlifold(20):
0.369
NanoFolder:
0.027
Positive Predictive Value
CentroidAlifold(20):
0.891
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs NanoFolder
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
NanoFolder:
0.068
Sensitivity
CentroidHomfold‑LAST:
0.359
NanoFolder:
0.027
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs NanoFolder
Matthews Correlation Coefficient
Contrafold:
0.556
NanoFolder:
0.068
Sensitivity
Contrafold:
0.542
NanoFolder:
0.027
Positive Predictive Value
Contrafold:
0.571
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(seed):
0.536
NanoFolder:
0.068
Sensitivity
MXScarna(seed):
0.382
NanoFolder:
0.027
Positive Predictive Value
MXScarna(seed):
0.753
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(20):
0.532
NanoFolder:
0.068
Sensitivity
RNAalifold(20):
0.359
NanoFolder:
0.027
Positive Predictive Value
RNAalifold(20):
0.787
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs NanoFolder
Matthews Correlation Coefficient
Sfold:
0.527
NanoFolder:
0.068
Sensitivity
Sfold:
0.477
NanoFolder:
0.027
Positive Predictive Value
Sfold:
0.582
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs NanoFolder
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
NanoFolder:
0.068
Sensitivity
RNAalifold(seed):
0.328
NanoFolder:
0.027
Positive Predictive Value
RNAalifold(seed):
0.835
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs NanoFolder
Matthews Correlation Coefficient
MaxExpect:
0.516
NanoFolder:
0.068
Sensitivity
MaxExpect:
0.505
NanoFolder:
0.027
Positive Predictive Value
MaxExpect:
0.528
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs NanoFolder
Matthews Correlation Coefficient
ProbKnot:
0.512
NanoFolder:
0.068
Sensitivity
ProbKnot:
0.508
NanoFolder:
0.027
Positive Predictive Value
ProbKnot:
0.517
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs NanoFolder
Matthews Correlation Coefficient
MXScarna(20):
0.500
NanoFolder:
0.068
Sensitivity
MXScarna(20):
0.353
NanoFolder:
0.027
Positive Predictive Value
MXScarna(20):
0.710
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs NanoFolder
Matthews Correlation Coefficient
UNAFold:
0.494
NanoFolder:
0.068
Sensitivity
UNAFold:
0.491
NanoFolder:
0.027
Positive Predictive Value
UNAFold:
0.498
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs NanoFolder
Matthews Correlation Coefficient
Fold:
0.493
NanoFolder:
0.068
Sensitivity
Fold:
0.496
NanoFolder:
0.027
Positive Predictive Value
Fold:
0.490
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs NanoFolder
Matthews Correlation Coefficient
RNAfold:
0.485
NanoFolder:
0.068
Sensitivity
RNAfold:
0.486
NanoFolder:
0.027
Positive Predictive Value
RNAfold:
0.485
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs NanoFolder
Matthews Correlation Coefficient
PknotsRG:
0.474
NanoFolder:
0.068
Sensitivity
PknotsRG:
0.440
NanoFolder:
0.027
Positive Predictive Value
PknotsRG:
0.510
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs NanoFolder
Matthews Correlation Coefficient
CRWrnafold:
0.430
NanoFolder:
0.068
Sensitivity
CRWrnafold:
0.379
NanoFolder:
0.027
Positive Predictive Value
CRWrnafold:
0.490
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs NanoFolder
Matthews Correlation Coefficient
RNAsubopt:
0.422
NanoFolder:
0.068
Sensitivity
RNAsubopt:
0.344
NanoFolder:
0.027
Positive Predictive Value
RNAsubopt:
0.519
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(20):
0.419
NanoFolder:
0.068
Sensitivity
TurboFold(20):
0.210
NanoFolder:
0.027
Positive Predictive Value
TurboFold(20):
0.837
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs NanoFolder
Matthews Correlation Coefficient
PPfold(20):
0.416
NanoFolder:
0.068
Sensitivity
PPfold(20):
0.197
NanoFolder:
0.027
Positive Predictive Value
PPfold(20):
0.879
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs NanoFolder
Matthews Correlation Coefficient
Afold:
0.408
NanoFolder:
0.068
Sensitivity
Afold:
0.343
NanoFolder:
0.027
Positive Predictive Value
Afold:
0.487
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs NanoFolder
Matthews Correlation Coefficient
Carnac(20):
0.401
NanoFolder:
0.068
Sensitivity
Carnac(20):
0.180
NanoFolder:
0.027
Positive Predictive Value
Carnac(20):
0.892
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs NanoFolder
Matthews Correlation Coefficient
McQFold:
0.389
NanoFolder:
0.068
Sensitivity
McQFold:
0.308
NanoFolder:
0.027
Positive Predictive Value
McQFold:
0.492
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(20):
0.377
NanoFolder:
0.068
Sensitivity
RSpredict(20):
0.241
NanoFolder:
0.027
Positive Predictive Value
RSpredict(20):
0.590
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs NanoFolder
Matthews Correlation Coefficient
RNAshapes:
0.367
NanoFolder:
0.068
Sensitivity
RNAshapes:
0.285
NanoFolder:
0.027
Positive Predictive Value
RNAshapes:
0.474
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs NanoFolder
Matthews Correlation Coefficient
RNASLOpt:
0.364
NanoFolder:
0.068
Sensitivity
RNASLOpt:
0.261
NanoFolder:
0.027
Positive Predictive Value
RNASLOpt:
0.510
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs NanoFolder
Matthews Correlation Coefficient
Multilign(20):
0.350
NanoFolder:
0.068
Sensitivity
Multilign(20):
0.161
NanoFolder:
0.027
Positive Predictive Value
Multilign(20):
0.762
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs NanoFolder
Matthews Correlation Coefficient
Murlet(20):
0.321
NanoFolder:
0.068
Sensitivity
Murlet(20):
0.131
NanoFolder:
0.027
Positive Predictive Value
Murlet(20):
0.787
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs NanoFolder
Matthews Correlation Coefficient
RNAwolf:
0.310
NanoFolder:
0.068
Sensitivity
RNAwolf:
0.261
NanoFolder:
0.027
Positive Predictive Value
RNAwolf:
0.369
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs NanoFolder
Matthews Correlation Coefficient
RSpredict(seed):
0.295
NanoFolder:
0.068
Sensitivity
RSpredict(seed):
0.146
NanoFolder:
0.027
Positive Predictive Value
RSpredict(seed):
0.597
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(20):
0.292
NanoFolder:
0.068
Sensitivity
CMfinder(20):
0.107
NanoFolder:
0.027
Positive Predictive Value
CMfinder(20):
0.794
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs NanoFolder
Matthews Correlation Coefficient
Vsfold4:
0.270
NanoFolder:
0.068
Sensitivity
Vsfold4:
0.198
NanoFolder:
0.027
Positive Predictive Value
Vsfold4:
0.371
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs NanoFolder
Matthews Correlation Coefficient
Vsfold5:
0.260
NanoFolder:
0.068
Sensitivity
Vsfold5:
0.195
NanoFolder:
0.027
Positive Predictive Value
Vsfold5:
0.347
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(20):
0.246
NanoFolder:
0.068
Sensitivity
RNASampler(20):
0.069
NanoFolder:
0.027
Positive Predictive Value
RNASampler(20):
0.877
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs NanoFolder
Matthews Correlation Coefficient
HotKnots:
0.212
NanoFolder:
0.068
Sensitivity
HotKnots:
0.069
NanoFolder:
0.027
Positive Predictive Value
HotKnots:
0.659
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs NanoFolder
Matthews Correlation Coefficient
Mastr(20):
0.187
NanoFolder:
0.068
Sensitivity
Mastr(20):
0.044
NanoFolder:
0.027
Positive Predictive Value
Mastr(20):
0.790
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs NanoFolder
Matthews Correlation Coefficient
Cylofold:
0.178
NanoFolder:
0.068
Sensitivity
Cylofold:
0.053
NanoFolder:
0.027
Positive Predictive Value
Cylofold:
0.597
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs NanoFolder
Matthews Correlation Coefficient
Pknots:
0.165
NanoFolder:
0.068
Sensitivity
Pknots:
0.041
NanoFolder:
0.027
Positive Predictive Value
Pknots:
0.662
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs NanoFolder
Matthews Correlation Coefficient
Alterna:
0.145
NanoFolder:
0.068
Sensitivity
Alterna:
0.036
NanoFolder:
0.027
Positive Predictive Value
Alterna:
0.582
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs NanoFolder
Matthews Correlation Coefficient
MCFold:
0.138
NanoFolder:
0.068
Sensitivity
MCFold:
0.044
NanoFolder:
0.027
Positive Predictive Value
MCFold:
0.432
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs NanoFolder
Matthews Correlation Coefficient
RDfolder:
0.138
NanoFolder:
0.068
Sensitivity
RDfolder:
0.031
NanoFolder:
0.027
Positive Predictive Value
RDfolder:
0.615
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs NanoFolder
Matthews Correlation Coefficient
Murlet(seed):
0.110
NanoFolder:
0.068
Sensitivity
Murlet(seed):
0.015
NanoFolder:
0.027
Positive Predictive Value
Murlet(seed):
0.811
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs NanoFolder
Matthews Correlation Coefficient
CMfinder(seed):
0.111
NanoFolder:
0.068
Sensitivity
CMfinder(seed):
0.016
NanoFolder:
0.027
Positive Predictive Value
CMfinder(seed):
0.757
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(seed) vs NanoFolder
Matthews Correlation Coefficient
TurboFold(seed):
0.106
NanoFolder:
0.068
Sensitivity
TurboFold(seed):
0.014
NanoFolder:
0.027
Positive Predictive Value
TurboFold(seed):
0.822
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(seed) vs NanoFolder
Matthews Correlation Coefficient
PPfold(seed):
0.097
NanoFolder:
0.068
Sensitivity
PPfold(seed):
0.012
NanoFolder:
0.027
Positive Predictive Value
PPfold(seed):
0.794
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(seed) vs NanoFolder
Matthews Correlation Coefficient
Carnac(seed):
0.093
NanoFolder:
0.068
Sensitivity
Carnac(seed):
0.010
NanoFolder:
0.027
Positive Predictive Value
Carnac(seed):
0.867
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(seed) vs NanoFolder
Matthews Correlation Coefficient
RNASampler(seed):
0.088
NanoFolder:
0.068
Sensitivity
RNASampler(seed):
0.009
NanoFolder:
0.027
Positive Predictive Value
RNASampler(seed):
0.847
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(seed) vs NanoFolder
Matthews Correlation Coefficient
Mastr(seed):
0.072
NanoFolder:
0.068
Sensitivity
Mastr(seed):
0.006
NanoFolder:
0.027
Positive Predictive Value
Mastr(seed):
0.820
NanoFolder:
0.173
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 1.14023792096e-06
|
|
+
NanoFolder vs Multilign(seed)
Matthews Correlation Coefficient
NanoFolder:
0.068
Multilign(seed):
0.059
Sensitivity
NanoFolder:
0.027
Multilign(seed):
0.005
Positive Predictive Value
NanoFolder:
0.173
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
| Multilign(seed) |
1242
ContextFold vs Multilign(seed)
Matthews Correlation Coefficient
ContextFold:
0.754
Multilign(seed):
0.059
Sensitivity
ContextFold:
0.717
Multilign(seed):
0.005
Positive Predictive Value
ContextFold:
0.794
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(seed):
0.737
Multilign(seed):
0.059
Sensitivity
PETfold_pre2.0(seed):
0.667
Multilign(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(seed):
0.814
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(seed):
0.611
Multilign(seed):
0.059
Sensitivity
CentroidAlifold(seed):
0.414
Multilign(seed):
0.005
Positive Predictive Value
CentroidAlifold(seed):
0.901
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
IPknot vs Multilign(seed)
Matthews Correlation Coefficient
IPknot:
0.607
Multilign(seed):
0.059
Sensitivity
IPknot:
0.551
Multilign(seed):
0.005
Positive Predictive Value
IPknot:
0.670
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PETfold_pre2.0(20) vs Multilign(seed)
Matthews Correlation Coefficient
PETfold_pre2.0(20):
0.575
Multilign(seed):
0.059
Sensitivity
PETfold_pre2.0(20):
0.410
Multilign(seed):
0.005
Positive Predictive Value
PETfold_pre2.0(20):
0.807
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidFold vs Multilign(seed)
Matthews Correlation Coefficient
CentroidFold:
0.571
Multilign(seed):
0.059
Sensitivity
CentroidFold:
0.515
Multilign(seed):
0.005
Positive Predictive Value
CentroidFold:
0.633
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidAlifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
CentroidAlifold(20):
0.573
Multilign(seed):
0.059
Sensitivity
CentroidAlifold(20):
0.369
Multilign(seed):
0.005
Positive Predictive Value
CentroidAlifold(20):
0.891
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CentroidHomfold‑LAST vs Multilign(seed)
Matthews Correlation Coefficient
CentroidHomfold‑LAST:
0.560
Multilign(seed):
0.059
Sensitivity
CentroidHomfold‑LAST:
0.359
Multilign(seed):
0.005
Positive Predictive Value
CentroidHomfold‑LAST:
0.874
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Contrafold vs Multilign(seed)
Matthews Correlation Coefficient
Contrafold:
0.556
Multilign(seed):
0.059
Sensitivity
Contrafold:
0.542
Multilign(seed):
0.005
Positive Predictive Value
Contrafold:
0.571
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(seed) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(seed):
0.536
Multilign(seed):
0.059
Sensitivity
MXScarna(seed):
0.382
Multilign(seed):
0.005
Positive Predictive Value
MXScarna(seed):
0.753
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(20):
0.532
Multilign(seed):
0.059
Sensitivity
RNAalifold(20):
0.359
Multilign(seed):
0.005
Positive Predictive Value
RNAalifold(20):
0.787
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Sfold vs Multilign(seed)
Matthews Correlation Coefficient
Sfold:
0.527
Multilign(seed):
0.059
Sensitivity
Sfold:
0.477
Multilign(seed):
0.005
Positive Predictive Value
Sfold:
0.582
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAalifold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNAalifold(seed):
0.523
Multilign(seed):
0.059
Sensitivity
RNAalifold(seed):
0.328
Multilign(seed):
0.005
Positive Predictive Value
RNAalifold(seed):
0.835
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MaxExpect vs Multilign(seed)
Matthews Correlation Coefficient
MaxExpect:
0.516
Multilign(seed):
0.059
Sensitivity
MaxExpect:
0.505
Multilign(seed):
0.005
Positive Predictive Value
MaxExpect:
0.528
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
ProbKnot vs Multilign(seed)
Matthews Correlation Coefficient
ProbKnot:
0.512
Multilign(seed):
0.059
Sensitivity
ProbKnot:
0.508
Multilign(seed):
0.005
Positive Predictive Value
ProbKnot:
0.517
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MXScarna(20) vs Multilign(seed)
Matthews Correlation Coefficient
MXScarna(20):
0.500
Multilign(seed):
0.059
Sensitivity
MXScarna(20):
0.353
Multilign(seed):
0.005
Positive Predictive Value
MXScarna(20):
0.710
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
UNAFold vs Multilign(seed)
Matthews Correlation Coefficient
UNAFold:
0.494
Multilign(seed):
0.059
Sensitivity
UNAFold:
0.491
Multilign(seed):
0.005
Positive Predictive Value
UNAFold:
0.498
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Fold vs Multilign(seed)
Matthews Correlation Coefficient
Fold:
0.493
Multilign(seed):
0.059
Sensitivity
Fold:
0.496
Multilign(seed):
0.005
Positive Predictive Value
Fold:
0.490
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAfold vs Multilign(seed)
Matthews Correlation Coefficient
RNAfold:
0.485
Multilign(seed):
0.059
Sensitivity
RNAfold:
0.486
Multilign(seed):
0.005
Positive Predictive Value
RNAfold:
0.485
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PknotsRG vs Multilign(seed)
Matthews Correlation Coefficient
PknotsRG:
0.474
Multilign(seed):
0.059
Sensitivity
PknotsRG:
0.440
Multilign(seed):
0.005
Positive Predictive Value
PknotsRG:
0.510
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CRWrnafold vs Multilign(seed)
Matthews Correlation Coefficient
CRWrnafold:
0.430
Multilign(seed):
0.059
Sensitivity
CRWrnafold:
0.379
Multilign(seed):
0.005
Positive Predictive Value
CRWrnafold:
0.490
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAsubopt vs Multilign(seed)
Matthews Correlation Coefficient
RNAsubopt:
0.422
Multilign(seed):
0.059
Sensitivity
RNAsubopt:
0.344
Multilign(seed):
0.005
Positive Predictive Value
RNAsubopt:
0.519
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(20) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(20):
0.419
Multilign(seed):
0.059
Sensitivity
TurboFold(20):
0.210
Multilign(seed):
0.005
Positive Predictive Value
TurboFold(20):
0.837
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(20) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(20):
0.416
Multilign(seed):
0.059
Sensitivity
PPfold(20):
0.197
Multilign(seed):
0.005
Positive Predictive Value
PPfold(20):
0.879
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Afold vs Multilign(seed)
Matthews Correlation Coefficient
Afold:
0.408
Multilign(seed):
0.059
Sensitivity
Afold:
0.343
Multilign(seed):
0.005
Positive Predictive Value
Afold:
0.487
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(20) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(20):
0.401
Multilign(seed):
0.059
Sensitivity
Carnac(20):
0.180
Multilign(seed):
0.005
Positive Predictive Value
Carnac(20):
0.892
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
McQFold vs Multilign(seed)
Matthews Correlation Coefficient
McQFold:
0.389
Multilign(seed):
0.059
Sensitivity
McQFold:
0.308
Multilign(seed):
0.005
Positive Predictive Value
McQFold:
0.492
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(20) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(20):
0.377
Multilign(seed):
0.059
Sensitivity
RSpredict(20):
0.241
Multilign(seed):
0.005
Positive Predictive Value
RSpredict(20):
0.590
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAshapes vs Multilign(seed)
Matthews Correlation Coefficient
RNAshapes:
0.367
Multilign(seed):
0.059
Sensitivity
RNAshapes:
0.285
Multilign(seed):
0.005
Positive Predictive Value
RNAshapes:
0.474
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASLOpt vs Multilign(seed)
Matthews Correlation Coefficient
RNASLOpt:
0.364
Multilign(seed):
0.059
Sensitivity
RNASLOpt:
0.261
Multilign(seed):
0.005
Positive Predictive Value
RNASLOpt:
0.510
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Multilign(20) vs Multilign(seed)
Matthews Correlation Coefficient
Multilign(20):
0.350
Multilign(seed):
0.059
Sensitivity
Multilign(20):
0.161
Multilign(seed):
0.005
Positive Predictive Value
Multilign(20):
0.762
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(20) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(20):
0.321
Multilign(seed):
0.059
Sensitivity
Murlet(20):
0.131
Multilign(seed):
0.005
Positive Predictive Value
Murlet(20):
0.787
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNAwolf vs Multilign(seed)
Matthews Correlation Coefficient
RNAwolf:
0.310
Multilign(seed):
0.059
Sensitivity
RNAwolf:
0.261
Multilign(seed):
0.005
Positive Predictive Value
RNAwolf:
0.369
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RSpredict(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RSpredict(seed):
0.295
Multilign(seed):
0.059
Sensitivity
RSpredict(seed):
0.146
Multilign(seed):
0.005
Positive Predictive Value
RSpredict(seed):
0.597
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(20) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(20):
0.292
Multilign(seed):
0.059
Sensitivity
CMfinder(20):
0.107
Multilign(seed):
0.005
Positive Predictive Value
CMfinder(20):
0.794
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold4 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold4:
0.270
Multilign(seed):
0.059
Sensitivity
Vsfold4:
0.198
Multilign(seed):
0.005
Positive Predictive Value
Vsfold4:
0.371
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Vsfold5 vs Multilign(seed)
Matthews Correlation Coefficient
Vsfold5:
0.260
Multilign(seed):
0.059
Sensitivity
Vsfold5:
0.195
Multilign(seed):
0.005
Positive Predictive Value
Vsfold5:
0.347
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(20) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(20):
0.246
Multilign(seed):
0.059
Sensitivity
RNASampler(20):
0.069
Multilign(seed):
0.005
Positive Predictive Value
RNASampler(20):
0.877
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
HotKnots vs Multilign(seed)
Matthews Correlation Coefficient
HotKnots:
0.212
Multilign(seed):
0.059
Sensitivity
HotKnots:
0.069
Multilign(seed):
0.005
Positive Predictive Value
HotKnots:
0.659
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(20) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(20):
0.187
Multilign(seed):
0.059
Sensitivity
Mastr(20):
0.044
Multilign(seed):
0.005
Positive Predictive Value
Mastr(20):
0.790
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Cylofold vs Multilign(seed)
Matthews Correlation Coefficient
Cylofold:
0.178
Multilign(seed):
0.059
Sensitivity
Cylofold:
0.053
Multilign(seed):
0.005
Positive Predictive Value
Cylofold:
0.597
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Pknots vs Multilign(seed)
Matthews Correlation Coefficient
Pknots:
0.165
Multilign(seed):
0.059
Sensitivity
Pknots:
0.041
Multilign(seed):
0.005
Positive Predictive Value
Pknots:
0.662
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Alterna vs Multilign(seed)
Matthews Correlation Coefficient
Alterna:
0.145
Multilign(seed):
0.059
Sensitivity
Alterna:
0.036
Multilign(seed):
0.005
Positive Predictive Value
Alterna:
0.582
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
MCFold vs Multilign(seed)
Matthews Correlation Coefficient
MCFold:
0.138
Multilign(seed):
0.059
Sensitivity
MCFold:
0.044
Multilign(seed):
0.005
Positive Predictive Value
MCFold:
0.432
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RDfolder vs Multilign(seed)
Matthews Correlation Coefficient
RDfolder:
0.138
Multilign(seed):
0.059
Sensitivity
RDfolder:
0.031
Multilign(seed):
0.005
Positive Predictive Value
RDfolder:
0.615
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Murlet(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Murlet(seed):
0.110
Multilign(seed):
0.059
Sensitivity
Murlet(seed):
0.015
Multilign(seed):
0.005
Positive Predictive Value
Murlet(seed):
0.811
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
CMfinder(seed) vs Multilign(seed)
Matthews Correlation Coefficient
CMfinder(seed):
0.111
Multilign(seed):
0.059
Sensitivity
CMfinder(seed):
0.016
Multilign(seed):
0.005
Positive Predictive Value
CMfinder(seed):
0.757
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
TurboFold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
TurboFold(seed):
0.106
Multilign(seed):
0.059
Sensitivity
TurboFold(seed):
0.014
Multilign(seed):
0.005
Positive Predictive Value
TurboFold(seed):
0.822
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
PPfold(seed) vs Multilign(seed)
Matthews Correlation Coefficient
PPfold(seed):
0.097
Multilign(seed):
0.059
Sensitivity
PPfold(seed):
0.012
Multilign(seed):
0.005
Positive Predictive Value
PPfold(seed):
0.794
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Carnac(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Carnac(seed):
0.093
Multilign(seed):
0.059
Sensitivity
Carnac(seed):
0.010
Multilign(seed):
0.005
Positive Predictive Value
Carnac(seed):
0.867
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
RNASampler(seed) vs Multilign(seed)
Matthews Correlation Coefficient
RNASampler(seed):
0.088
Multilign(seed):
0.059
Sensitivity
RNASampler(seed):
0.009
Multilign(seed):
0.005
Positive Predictive Value
RNASampler(seed):
0.847
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
Mastr(seed) vs Multilign(seed)
Matthews Correlation Coefficient
Mastr(seed):
0.072
Multilign(seed):
0.059
Sensitivity
Mastr(seed):
0.006
Multilign(seed):
0.005
Positive Predictive Value
Mastr(seed):
0.820
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
1242
NanoFolder vs Multilign(seed)
Matthews Correlation Coefficient
NanoFolder:
0.068
Multilign(seed):
0.059
Sensitivity
NanoFolder:
0.027
Multilign(seed):
0.005
Positive Predictive Value
NanoFolder:
0.173
Multilign(seed):
0.741
Number of pairs reference - predicted secondary structure: 1242
Wilcoxon single-rank test P-value: 3.56938820447e-08
|
|