

| | ||||||||||||||
| ||||||||||||||
RNASLOpt 2D structure prediction for record 3ZND_W
REFERENCE VS PREDICTED RNA SECONDARY STRUCTURE
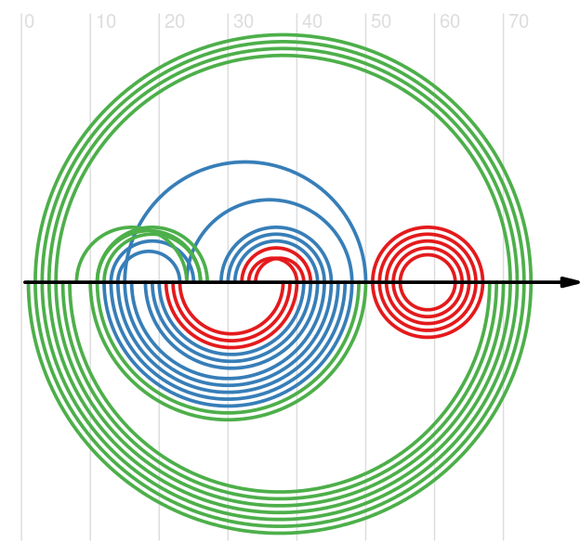
Figure 1. Graph showing comparison of reference (upper part) and predicted by RNASLOpt (lower part) secondary structures of RNA chain 3ZND_W.
Download RNASLOpt input in FASTA format
Table 1. Summary of base pair counts (true positives, true negatives, etc). Individual counts can be directly read on the graph above or can be extracted by comparing two bpseq files above.
| Base Pair Counts | |
|---|---|
| MCC | 0.21 |
| SENS | 0.22 |
| PPV | 0.22 |
| TP | 5 |
| TN | 2980 |
| FP | 20 |
| FP CONTRA | 1 |
| FP INCONS | 17 |
| FP COMP | 2 |
| FN | 18 |
RNA 2D structure plot generated with R-Chie.