Table of contents:
- Overview
- Performance Plots
- Performance of PETfold_pre2.0(seed)
- scored higher in this pairwise comparison
- Performance of CentroidFold
- scored lower in this pairwise comparison
- Compile and download dataset for PETfold_pre2.0(seed) & CentroidFold [.zip] - may take several seconds...
Overview
| Metric |
PETfold_pre2.0(seed) |
|
CentroidFold |
|---|
| MCC |
0.828 |
>
|
0.634 |
| Average MCC ± 95% Confidence Intervals |
0.777
±
0.178
|
>
|
0.658
±
0.205
|
| Sensitivity |
0.772 |
>
|
0.602 |
| Positive Predictive Value |
0.892 |
>
|
0.678 |
| Total TP |
132 |
>
|
103 |
| Total TN |
12516 |
>
|
12512 |
| Total FP |
42 |
<
|
69 |
| Total FP CONTRA |
3 |
<
|
13 |
| Total FP INCONS |
13 |
<
|
36 |
| Total FP COMP |
26 |
>
|
20 |
| Total FN |
39 |
<
|
68 |
| P-value |
0.0 |
Performance plots
-
Comparison of performance of PETfold_pre2.0(seed) and CentroidFold. Positive Predictive Value (PPV) is plotted against sensitivity. Each dot represents a single test of each method. See tables below for raw data
(individual counts for PETfold_pre2.0(seed)
and CentroidFold).

-
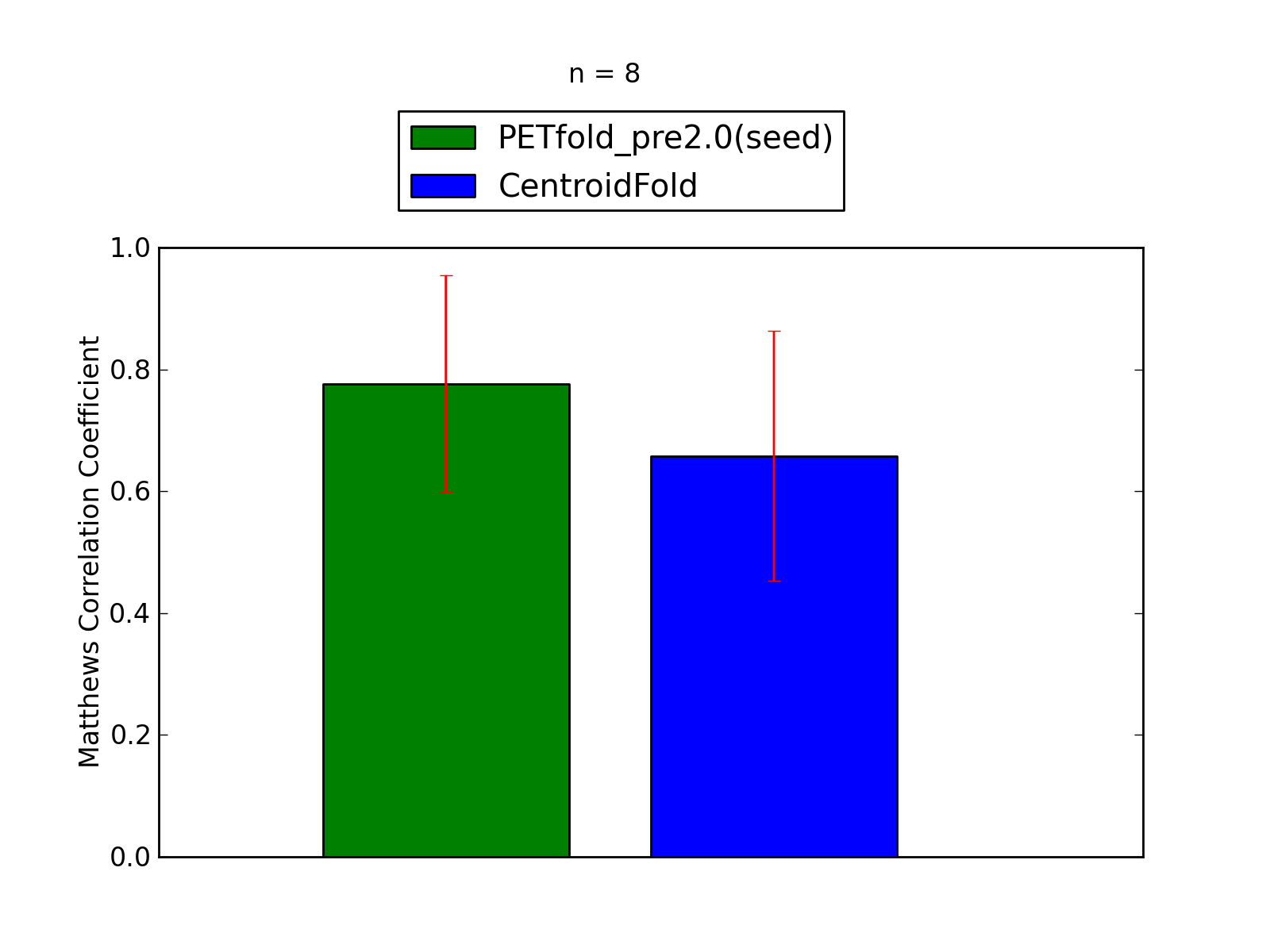
Comparison of average Matthews Correlation Coefficients (MCCs) for PETfold_pre2.0(seed) and CentroidFold. The whiskers correspond to 95% confidence intervals (CIs). 'n' denotes the number of MCCs used to calculate average MCCs and CIs. See tables below for raw data
(individual counts for PETfold_pre2.0(seed)
and CentroidFold).
Performance of PETfold_pre2.0(seed)
- scored higher in this pairwise comparison
1. Total counts & total scores for PETfold_pre2.0(seed)
| Total Base Pair Counts |
|---|
| Total TP |
132 |
| Total TN |
12516 |
| Total FP |
42 |
| Total FP CONTRA |
3 |
| Total FP INCONS |
13 |
| Total FP COMP |
26 |
| Total FN |
39 |
| Total Scores |
|---|
| MCC |
0.828 |
| Average MCC ± 95% Confidence Intervals |
0.777
±
0.178
|
| Sensitivity |
0.772 |
| Positive Predictive Value |
0.892 |
| Nr of predictions |
8 |
Performance of CentroidFold
- scored lower in this pairwise comparison
1. Total counts & total scores for CentroidFold
| Total Base Pair Counts |
|---|
| Total TP |
103 |
| Total TN |
12512 |
| Total FP |
69 |
| Total FP CONTRA |
13 |
| Total FP INCONS |
36 |
| Total FP COMP |
20 |
| Total FN |
68 |
| Total Scores |
|---|
| MCC |
0.634 |
| Average MCC ± 95% Confidence Intervals |
0.658
±
0.205
|
| Sensitivity |
0.602 |
| Positive Predictive Value |
0.678 |
| Nr of predictions |
8 |
Matthews Correlation Coeffient, Sensitivity and Positive Predictive Value have been calculated based
on the paper by Gardener & Giegerich, 2004.
| 


