

| |
|||||||||||
|
|||||||||||
ModeRNA server
A server for RNA structure modeling and analysis
by Magdalena Rother, Kaja Milanowska, Tomasz Puton, Kristian Rother, and Janusz M. Bujnicki.
What is the ModeRNA server?

This service is based on the ModeRNA software for RNA 3D structure prediction, analysis, and manipulation.
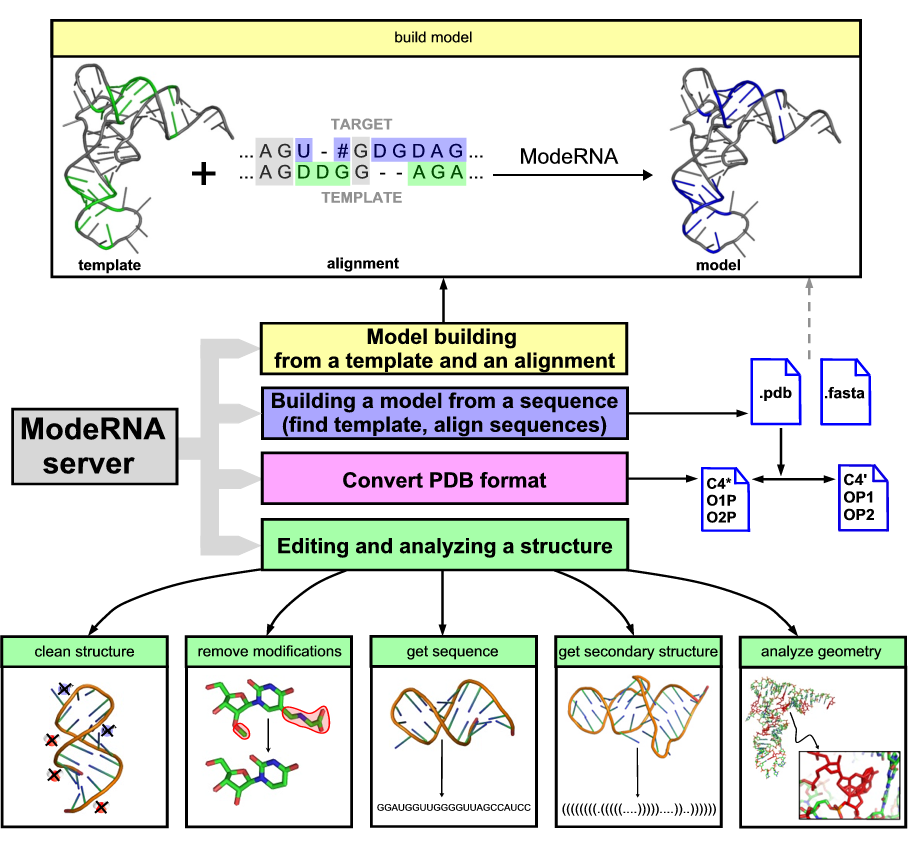
Currently, the server allows you to:
- build a model (requires a pairwise sequence alignment and a structural template).
- find template (looks for potential templates using ParAlign and Rfam).
- align sequences (prepares an alignment between two RNA sequences using R-coffee).
- convert PDB naming (changes nomenclature in a PDB file).
- clean up a structure (remove water, ions, and ligands, adds missing P atoms and changes '*' into ''' in ribose).
- remove modifications (replace modified nucleotides with standard ones).
- get a sequence (returns the sequence obtained from atomic coordinates in one-letter-code).
- get a secondary structure (extract the 2D structure from 3D and returns it in the Vienna format).
- analyze geometry (checks whether the residues of a structure have any unusual features like strange bond and angle values).
To execute your request go to the [submit] section.
The server was tested on the following browsers:
- Windows: Internet Explorer v. 8 or higher, Firefox v. 4, Google Chrome, Safari 5, Opera v. 10 or higher
- Linux: Firefox v. 4, Google Chrome, Opera v. 10 or higher
- Mac OS: Firefox v. 4, Google Chrome, Safari 5, Opera v. 10 or higher
|
|

|