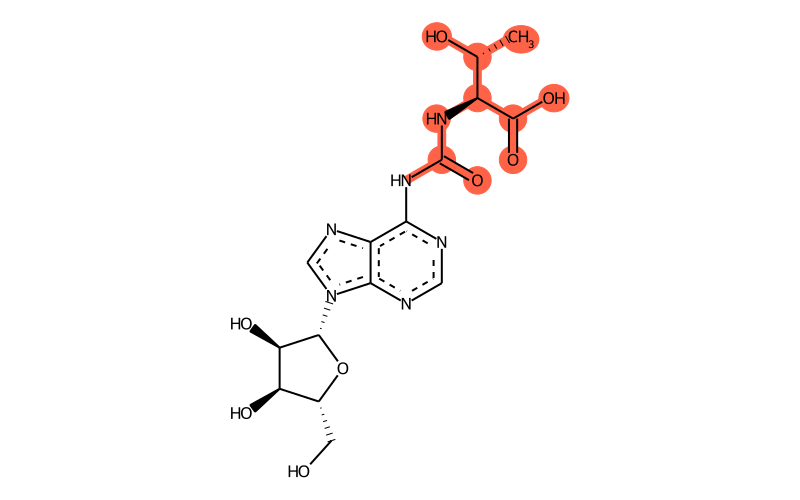
Summary
| Full name | N6-methyl-N6-threonylcarbamoyladenosine |
| IUPAC name | (2S,3R)-2-[[9-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]purin-6-yl]carbamoylamino]-3-hydroxybutanoic acid |
| Short name | m6t6A |
| MODOMICS code new | 2000000662A |
| MODOMICS code | 662A |
| Synonyms |
24719-82-2
28874-45-5 28882-41-9 (2S,3R)-2-(3-(9-((2R,3R,4S,5R)-3,4-Dihydroxy-5-(hydroxymethyl)tetrahydrofuran-2-yl)-9H-purin-6-yl)ureido)-3-hydroxybutanoic acid (2S,3R)-2-[({9-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-9H-purin-6-yl}carbamoyl)amino]-3-hydroxybutanoic acid (2S,3R)-2-[[9-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]purin-6-yl]carbamoylamino]-3-hydroxybutanoic acid (2s,3r)-2-[({9-[(2r,3r,4s,5r)-3,4-dihydroxy-5-(hydroxymethyl)tetrahydrofuran-2-yl]-9h-purin-6-yl}carbamoyl)amino]-3-hydroxybutanoic acid(non-preferred name) 3C55P4W4TB ((9-((2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)tetrahydrofuran-2-yl)-9H-purin-6-yl)carbamoyl)-L-threonine AC1L4PHC AC1Q5QSL Acaptma AR-1A3389 AR-1A3390 C15-H26-N3-O.Br.Br-H C15H26N3O.Br.BrH CHEBI:21440 CID161466 DTXSID70947655 KST-1A3490 KST-1A3491 L-Threonine, N-(((9-beta-D-ribofuranosyl-9H-purin-6-yl)amino)carbonyl)- N-{[(1S,2R)-1-carboxy-2-hydroxypropyl]carbamoyl}adenosine N6-[((1S,2R)-1-CARBOXY-2-HYDROXYBUTYLAMINO)CARBONYL]ADENOSINE SODIUM SALT N6-Carbamoyl-L-threonyladenosine N6-Carbamoylthreonyladenosine n6-(n-threonylcarbonyl)adenosine N(6)-(N-Threonylcarbonyl)adenosine N6-Threoninocarbonyladenosine N6-Threonylcarbamoyladenosine N-[(9- -D-ribofuranosylpurin-6-yl)carbamoyl]threonine N-[(9-beta-D-ribofuranosylpurin-6-yl)carbamoyl]-L-threonine N-[(9-beta-D-ribofuranosylpurin-6-yl)carbamoyl]threonine N-((9-beta-D-Ribofuranosylpurin-6-yl)-carbamoyl)threonine N(epsilon-Aminocaproyl)-p-aminophenyltrimethylammonium N-{Hydroxy[(9-pentofuranosyl-9H-purin-6-yl)amino]methylidene}threonine N-(Purin-6-ylcarbamoyl)-L-threonine riboside Q15632802 SCHEMBL63155 t(6)a T6a UNII-3C55P4W4TB ZINC31474771 |
| Nature of the modified residue | Natural |
| RNAMods code | E |
| Residue unique ID | 16 |
| Found in RNA | Yes |
| Related nucleotides | 231 |
| Enzymes |
TRMO (Homo sapiens) TrmO (Escherichia coli) |
| Found in phylogeny | Eubacteria |
| Found naturally in RNA types | tRNA |
Chemical information
| Sum formula | C15H20N6O8 |
| Type of moiety | nucleoside |
| Degeneracy | not applicable |
| ChEBI ID | 133071 |
| CAS Registry Number | 24719-82-2 |
| Reaxys Registry Number | 23417855 |
| SMILES | C[C@H]([C@@H](C(=O)O)NC(Nc1c2c([n]([C@H]3[C@H](O)[C@H](O)[C@@H](CO)O3)cn2)ncn1)=O)O |
| logP | -2.1428 |
| TPSA | 212.18 |
| Number of atoms | 29 |
| Number of Hydrogen Bond Acceptors 1 (HBA1) | 13 |
| Number of Hydrogen Bond Acceptors 2 (HBA2) | 14 |
| Number of Hydrogen Bond Donors (HBD) | 7 |
| InChI | InChI=1S/C15H20N6O8/c1-5(23)7(14(26)27)19-15(28)20-11-8-12(17-3-16-11)21(4-18-8)13-10(25)9(24)6(2-22)29-13/h3-7,9-10,13,22-25H,2H2,1H3,(H,26,27)(H2,16,17,19,20,28)/t5-,6-,7+,9-,10-,13-/m1/s1 |
| InChIKey | UNUYMBPXEFMLNW-DWVDDHQFSA-N |
| Search the molecule in external databases | ChEMBL PubChem Compound Database Ligand Expo WIPO |
| PubChem CID | |
| PubChem SIDs |
46236306
50980346 57349533 103067534 103067535 103080266 103080267 113447121 117541698 129395049 135117788 135668177 152344590 226444550 260050018 274077190 312234225 319175847 341299675 349525476 381382645 381832734 384504704 385078257 385642117 386478390 388608286 404760147 419579651 434311205 434369767 438490155 441225637 443280185 |
* Chemical properties calculated with Open Babel - O'Boyle et al. Open Babel: An open chemical toolbox. J Cheminform 3, 33 (2011) (link)
Download Structures
| 2D | .png .mol .mol2 .sdf .pdb .smi |
| 3D | .mol .mol2 .sdf .pdb |
Tautomers
| Tautomers SMILES |
CC(C(C(O)=O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #0
CC(C(C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)=O tautomer #1 CC(C(C(O)O)=NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #2 CC(C(C(O)=O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #3 CC(C(C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)=O tautomer #4 CC(C(C(O)O)=NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #5 C=C(C(C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #6 C=C(C(C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #7 CC(=C(C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #8 CC(=C(C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #9 CC(C(C(O)=O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)=O)O tautomer #10 CC(C(C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)=O)=O tautomer #11 CC(C(C(O)O)=NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)=O)O tautomer #12 CC(C(C(O)=O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)=O)O tautomer #13 CC(C(C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)=O)=O tautomer #14 CC(C(C(O)O)=NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)=O)O tautomer #15 CC(C(=C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #16 CC(C(=C(O)O)NC(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)=O)O tautomer #17 CC(C(C(O)=O)N=C(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #18 CC(C(C(O)=O)N=C(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #19 CC(C(C(O)=O)NC(=Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #20 CC(C(C(O)O)NC(=Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)=O tautomer #21 CC(C(C(O)O)=NC(=Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #22 C=C(C(C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)=O)O tautomer #23 C=C(C(C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)=O)O tautomer #24 CC(C(C(O)=O)NC(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)=O)O tautomer #25 CC(C(C(O)O)NC(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)=O)=O tautomer #26 CC(C(C(O)O)=NC(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)=O)O tautomer #27 CC(=C(C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)=O)O tautomer #28 CC(=C(C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)=O)O tautomer #29 C=C(C(C(O)O)NC(=Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #30 CC(=C(C(O)O)NC(=Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #31 C=C(C(C(O)O)NC(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)=O)O tautomer #32 CC(C(=C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)=O)O tautomer #33 CC(C(=C(O)O)NC(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)=O)O tautomer #34 CC(C(C(O)=O)N=C(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)O)O tautomer #35 CC(C(C(O)=O)N=C(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)O)O tautomer #36 CC(=C(C(O)O)NC(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)=O)O tautomer #37 CC(C(=C(O)O)N=C(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #38 CC(C(=C(O)O)N=C(Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #39 CC(C(=C(O)O)NC(=Nc1c2c(n(C3C(O)C(O)C(CO)O3)cn2)ncn1)O)O tautomer #40 CC(C(=C(O)O)NC(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)=O)O tautomer #41 CC(C(C(O)=O)N=C(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)O)O tautomer #42 CC(C(=C(O)O)N=C(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)[nH]cn1)O)O tautomer #43 CC(C(=C(O)O)N=C(N=c1c2c(n(C3C(O)C(O)C(CO)O3)cn2)nc[nH]1)O)O tautomer #44 CC(C(=C(O)O)N=C(N=C1C2C(N(C3C(O)C(O)C(CO)O3)C=N2)=NC=N1)O)O tautomer #45 |
| Tautomer image | Show Image |
Predicted CYP Metabolic Sites
| CYP3A4 | CYP2D6 | CYP2C9 |
|---|---|---|
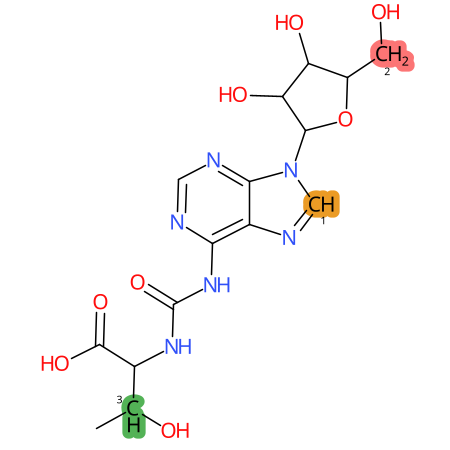
|
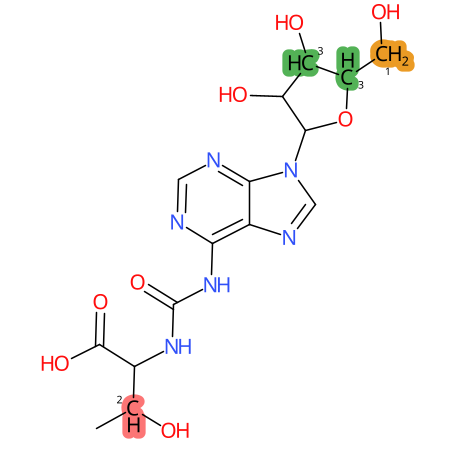
|
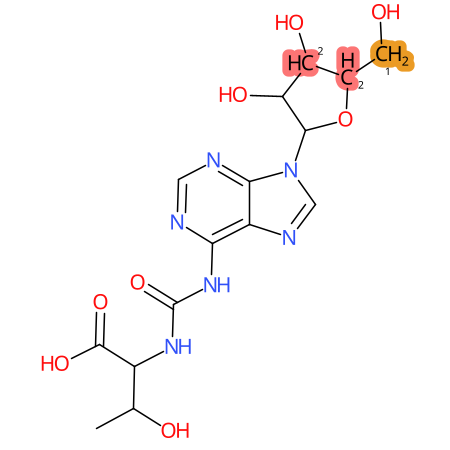
|
* CYP Metabolic sites predicted with SMARTCyp. SMARTCyp is a method for prediction of which sites in a molecule that are most liable to metabolism by Cytochrome P450. It has been shown to be applicable to metabolism by the isoforms 1A2, 2A6, 2B6, 2C8, 2C19, 2E1, and 3A4 (CYP3A4), and specific models for the isoform 2C9 (CYP2C9) and isoform 2D6 (CYP2D6). CYP3A4, CYP2D6, and CYP2C9 are the three of the most important enzymes in drug metabolism since they are involved in the metabolism of more than half of the drugs used today. The three top-ranked atoms are highlighted. See: SmartCYP and SmartCYP - background; Patrik Rydberg, David E. Gloriam, Lars Olsen, The SMARTCyp cytochrome P450 metabolism prediction server, Bioinformatics, Volume 26, Issue 23, 1 December 2010, Pages 2988–2989 (link)
LC-MS Information
| Monoisotopic mass | 426.1499 |
| Average mass | 412.355 |
| [M+H]+ | 427.1577 |
| Product ions | 295 |
| Normalized LC elution time * | 1,93 (Kellner 2014) |
| LC elution order/characteristics | between A and m6A (Kellner 2014) |
* normalized to guanosine (G), measured with a RP C-18 column with acetonitrile/ammonium acetate as mobile phase.
LC-MS Publications
| Title | Authors | Journal | Details | ||
|---|---|---|---|---|---|
| Profiling of RNA modifications by multiplexed stable isotope labelling. | Kellner S, Neumann J, Rosenkranz D, Lebedeva S, Ketting RF, Zischler H, Schneider D, Helm M. | Chem Commun (Camb). | [details] | 24567952 | - |
Chemical groups contained
| Type | Subtype |
|---|---|
| aminoacyl group | threonyl |
| methyl group | methyl at other N |
| other | carbamoyl |
Reactions producing N6-methyl-N6-threonylcarbamoyladenosine
| Name |
|---|
| t6A:m6t6A |
Last modification of this entry: July 16, 2025