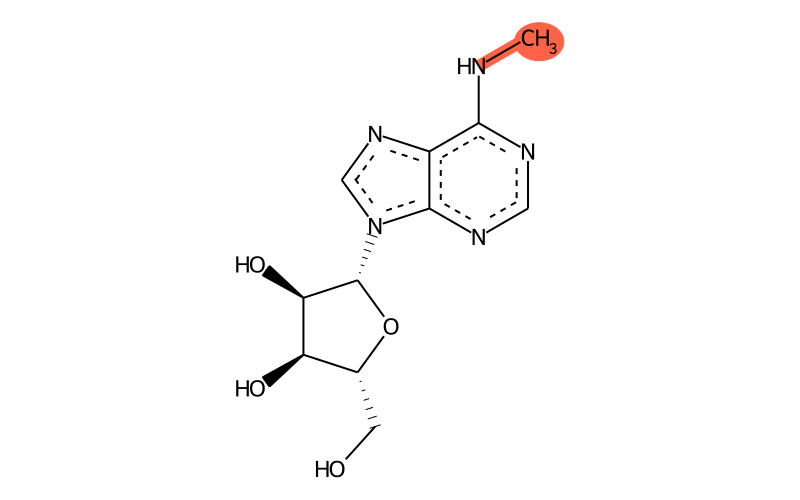
Summary
| Full name | N6-methyladenosine |
| IUPAC name | (2R,3S,4R,5R)-2-(hydroxymethyl)-5-[6-(methylamino)purin-9-yl]oxolane-3,4-diol |
| Short name | m6A |
| MODOMICS code new | 2000000006A |
| MODOMICS code | 6A |
| Synonyms |
1867-73-8
2575AH (2R,3S,4R,5R)-2-(hydroxymethyl)-5-[6-(methylamino)-9H-purin-9-yl]oxolane-3,4-diol (2R,3S,4R,5R)-2-(Hydroxymethyl)-5-(6-(methylamino)-9H-purin-9-yl)tetrahydrofuran-3,4-diol (2R,3S,4R,5R)-2-(hydroxymethyl)-5-[6-(methylamino)purin-9-yl]oxolane-3,4-diol (2R,3S,4R,5R)-2-Hydroxymethyl-5-(6-methylamino-purin-9-yl)-tetrahydro-furan-3,4-diol (2R,3S,4R,5R)-2-(hydroxymethyl)-5-[6-(methylamino)purin-9-yl]tetrahydrofuran-3,4-diol 4rdn (4S,2R,3R,5R)-5-(hydroxymethyl)-2-[6-(methylamino)purin-9-yl]oxolane-3,4-diol 56860-12-9 60209-41-8 6MD 6-Methyladenosine 6-methylamino-9-beta-delta-ribofuranosyl-Purine 6-methylamino-9-beta-D-ribofuranosyl-Purine 6-Methylaminopurine 9-ribofuranoside 6-Methylaminopurine arabinoside 6-Methylaminopurine D-riboside 6-Methylaminopurine ribonucleoside 6-Methylaminopurine riboside 6-Methylaminopurinosine 6MZ AC1L2SAS AC1Q4Y57 Adenosine, N-methyl- Adenosine, N-methyl- (8CI) Adenosine, N-methyl- (8CI)(9CI) AJ-49684 AKOS024260287 AR-1K7612 Ara-map BDBM50163023 BPBio1_000413 BRD-K81807412-001-03-1 BSPBio_000375 C010223 C11-H15-N5-O4 C11H15N5O4 CCG-220338 CHEBI:21891 CHEMBL383144 CID102175 CLE6G00625 CS-3285 CTK0H6388 DTXSID6020858 Epitope ID:140951 GS-6701 HMS1569C17 HMS2096C17 HMS2234N05 HMS3713C17 HY-N0086 J-700317 LS-7512 M6A MFCD00005739 MLS002153780 N6 -methyladenosine N(6)mAdo N6-Methyladenosine (N-6)-Methyladenosine N(6)-Methyladenosine N6-methyladenosine (m6A) N6-METHYLADENOSINE-5'-MONOPHOSPHATE N(6)-monomethyladenosine N/A NCGC00162241-01 NCGC00162241-02 NCGC00162241-04 N-methyl-9-alpha-L-ribofuranosyl-9H-purin-6-amine N-methyl-9-beta-L-ribofuranosyl-9H-purin-6-amine N-Methyladenosine NSC 29409 NSC-29409 NSC627045 PDSP1_001030 PDSP2_001014 Prestwick0_000338 Prestwick1_000338 Prestwick2_000338 Prestwick3_000338 Prestwick_343 Purine, 6-methylamino-9-beta-D-ribofuranosyl- Q6951996 s3190 SB34650 SCHEMBL40634 SMR001233164 SPBio_002296 ST24034024 ST50412061 SW196617-3 UNII-CLE6G00625 VQAYFKKCNSOZKM-IOSLPCCCSA-N ZINC4245632 |
| Nature of the modified residue | Natural |
| RNAMods code | Ж |
| Residue unique ID | 96 |
| Found in RNA | Yes |
| Related nucleotides | 211 |
| Enzymes |
Bmc1 (Schizosaccharomyces pombe) DIM1B (Arabidopsis thaliana) DIMT1 (Caenorhabditis elegans) DIMT1L (Homo sapiens) ErmAM (Streptococcus pneumoniae) ErmBC (Escherichia coli) ErmC' (Bacillus subtilis) FIONA1 (Arabidopsis thaliana) HVO_1475 (Haloferax volcanii) Ime4 (Saccharomyces cerevisiae) METT-10 (Caenorhabditis elegans) METTL14 (Homo sapiens) METTL3 (Homo sapiens) Mettl16 (Schizosaccharomyces pombe) Mettl5 (Arabidopsis thaliana) RlmF (Escherichia coli) RlmJ (Escherichia coli) RsmA (Escherichia coli) RsmA (Saccharomyces cerevisiae) RsmA (Thermus thermophilus) RsmA (Pseudomonas aeruginosa) TFB1M (Mus musculus) TRM112 (Homo sapiens) TrmM (Escherichia coli) ksgA (Haloferax volcanii) mtTFBp (Saccharomyces cerevisiae) |
| Found in phylogeny | Eubacteria, Eukaryota |
| Found naturally in RNA types | mRNA, rRNA, snRNA, tRNA |
Chemical information
* Chemical properties calculated with Open Babel - O'Boyle et al. Open Babel: An open chemical toolbox. J Cheminform 3, 33 (2011) (link)
QM Data:
| Dipole Magnitude [D]: | 3.952752323 |
| Energy [Eh]: | -1002.51573901082 |
| HOMO [eV]: | -8.6444 |
| LUMO [eV]: | 1.0651 |
| Gap [eV]: | 9.7095 |
Download QM Data:
| Charges | charge.txt |
Download Structures
| 2D | .png .mol .mol2 .sdf .pdb .smi |
| 3D | .mol .mol2 .sdf .pdb |
Tautomers
| Tautomers SMILES |
CNc1ncnc2c1ncn2C3OC(CO)C(O)C3O tautomer #0
CNc1ncnc2c1ncn2C3OC(CO)C(O)C3O tautomer #1 CN=c1nc[nH]c2c1ncn2C3OC(CO)C(O)C3O tautomer #2 CN=c1[nH]cnc2c1ncn2C3OC(CO)C(O)C3O tautomer #3 CN=C1N=CN=C2C1N=CN2C3OC(CO)C(O)C3O tautomer #4 |
| Tautomer image | Show Image |
Predicted CYP Metabolic Sites
| CYP3A4 | CYP2D6 | CYP2C9 |
|---|---|---|
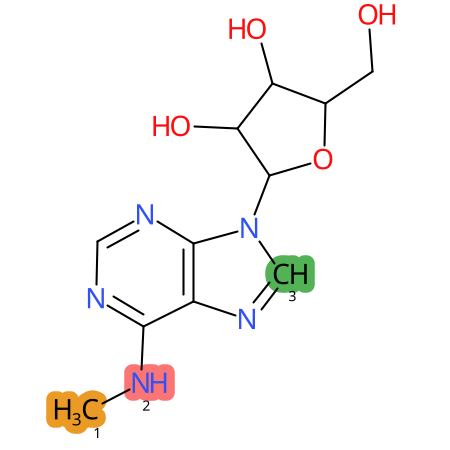
|
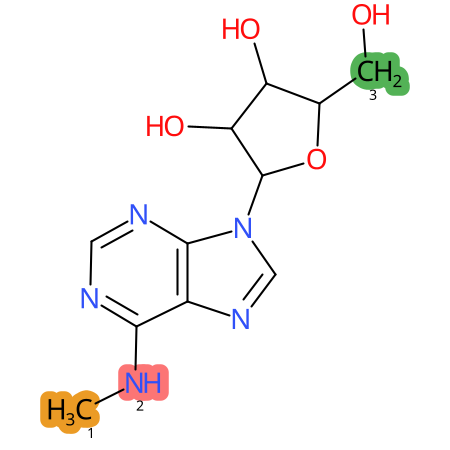
|
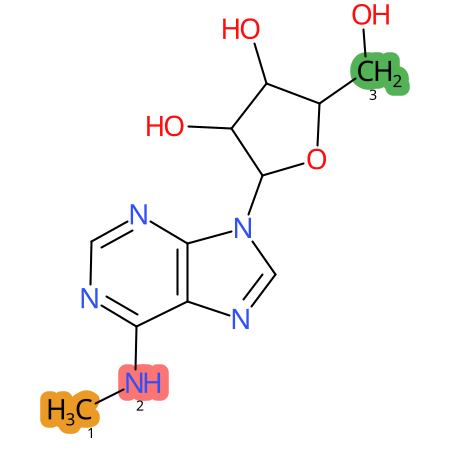
|
* CYP Metabolic sites predicted with SMARTCyp. SMARTCyp is a method for prediction of which sites in a molecule that are most liable to metabolism by Cytochrome P450. It has been shown to be applicable to metabolism by the isoforms 1A2, 2A6, 2B6, 2C8, 2C19, 2E1, and 3A4 (CYP3A4), and specific models for the isoform 2C9 (CYP2C9) and isoform 2D6 (CYP2D6). CYP3A4, CYP2D6, and CYP2C9 are the three of the most important enzymes in drug metabolism since they are involved in the metabolism of more than half of the drugs used today. The three top-ranked atoms are highlighted. See: SmartCYP and SmartCYP - background; Patrik Rydberg, David E. Gloriam, Lars Olsen, The SMARTCyp cytochrome P450 metabolism prediction server, Bioinformatics, Volume 26, Issue 23, 1 December 2010, Pages 2988–2989 (link)
LC-MS Information
| Monoisotopic mass | 281.1124 |
| Average mass | 281.268 |
| [M+H]+ | 282.1202 |
| Product ions | 150 |
| Normalized LC elution time * | 1,66 (Kellner 2014); 1,97 (Kellner 2014) |
| LC elution order/characteristics | after A (Kellner 2014, Kellner 2014) |
* normalized to guanosine (G), measured with a RP C-18 column with acetonitrile/ammonium acetate as mobile phase.
LC-MS Publications
| Title | Authors | Journal | Details | ||
|---|---|---|---|---|---|
| Profiling of RNA modifications by multiplexed stable isotope labelling. | Kellner S, Neumann J, Rosenkranz D, Lebedeva S, Ketting RF, Zischler H, Schneider D, Helm M. | Chem Commun (Camb). | [details] | 24567952 | - |
| Absolute and relative quantification of RNA modifications via biosynthetic isotopomers. | Kellner S, Ochel A, Thuring K, Spenkuch F, Neumann J, Sharma S, Entian KD, Schneider D, Helm M... | Nucleic Acids Res | [details] | 25129236 | - |
Chemical groups contained
| Type | Subtype |
|---|---|
| methyl group | methyl at other N |
Reactions producing N6-methyladenosine
| Name |
|---|
| A:m6A |
Reactions starting from N6-methyladenosine
| Name |
|---|
| m6A:ms2m6A |
| m6A:m6,6A |
| m6A:m6Am |
| m6A:A |
| m6A:hm6A |
Last modification of this entry: Sept. 15, 2025